Abstract
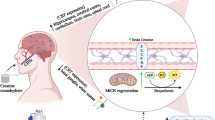
In the last decade, the study of Wnt and Notch signaling in cell biology has led to significant progress in understanding embryogenesis, bone development, muscle healing, neurogenesis, and tumorigenesis. It has been found that regular physical activity can counteract the decline of skeletal muscle caused by aging, which is linked to osteoporosis, regenerative neurogenesis, hippocampal function, cognitive ability, and the creation of neuromuscular junctions. Despite these discoveries, there is still uncertainty about how cell biology and exercise can impact the Wnt and Notch signaling pathways in the locomotor system. This review aims to summarize the potential influence of exercise on Wnt and Notch signaling.


Similar content being viewed by others
References
Caspersen CJ, Powell KE, Christenson GM. Physical activity, exercise, and physical fitness: definitions and distinctions for health-related research. Public Health Rep (Washington, DC: 1974). 1985;100(2):126–31.
Garber CE, Blissmer B, Deschenes MR, Franklin BA, Lamonte MJ, Lee IM, et al. American College of Sports Medicine position stand. Quantity and quality of exercise for developing and maintaining cardiorespiratory, musculoskeletal, and neuromotor fitness in apparently healthy adults: guidance for prescribing exercise. Med Sci Sports Exerc. 2011;43(7):1334–59.
Arem H, Moore SC, Patel A, Hartge P, de BerringtonGonzalez A, Visvanathan K, et al. Leisure time physical activity and mortality: a detailed pooled analysis of the dose−response relationship. JAMA Intern Med. 2015;175(6):959–67.
Bauman AE, Kamada M, Reis RS, Troiano RP, Ding D, Milton K, et al. An evidence-based assessment of the impact of the Olympic Games on population levels of physical activity. Lancet (London, England). 2021;398(10298):456–64.
Lieberman DE, Kistner TM, Richard D, Lee IM, Baggish AL. The active grandparent hypothesis: Physical activity and the evolution of extended human healthspans and lifespans. Proc Natl Acad Sci U S A. 2021. https://doi.org/10.1073/pnas.2107621118.
Ferguson B. ACSM’s guidelines for exercise testing and prescription 9th Ed. 2014: J Can Chiropr Assoc. 2014;58(3):328.
García-Pinillos F, Laredo-Aguilera JA, Muñoz-Jiménez M, Latorre-Román PA. Effects of 12-week concurrent high-intensity interval strength and endurance training program on physical performance in healthy older people. J Strength Cond Res. 2019;33(5):1445–52.
Lamberti N, Straudi S, Malagoni AM, Argirò M, Felisatti M, Nardini E, et al. Effects of low-intensity endurance and resistance training on mobility in chronic stroke survivors: a pilot randomized controlled study. Eur J Phys Rehabil Med. 2017;53(2):228–39.
Wehrle A, Kneis S, Dickhuth HH, Gollhofer A, Bertz H. Endurance and resistance training in patients with acute leukemia undergoing induction chemotherapy-a randomized pilot study. Supportive care in cancer : official journal of the Multinational Association of Supportive Care in Cancer. 2019;27(3):1071–9.
Wodarz A, Nusse R. Mechanisms of Wnt signaling in development. Annu Rev Cell Dev Biol. 1998;14:59–88.
Habas R, Dawid IB. Dishevelled and Wnt signaling: is the nucleus the final frontier? J Biol. 2005;4(1):2.
Yamaguchi TP. Heads or tails: Wnts and anterior−posterior patterning. Current biology : CB. 2001;11(17):R713–24.
Logan CY, Nusse R. The Wnt signaling pathway in development and disease. Annu Rev Cell Dev Biol. 2004;20:781–810.
Artavanis-Tsakonas S, Rand MD, Lake RJ. Notch signaling: cell fate control and signal integration in development. Science (New York, NY). 1999;284(5415):770–6.
Baron M. An overview of the Notch signalling pathway. Semin Cell Dev Biol. 2003;14(2):113–9.
Greenwald I. LIN-12/Notch signaling: lessons from worms and flies. Genes Dev. 1998;12(12):1751–62.
Kadesch T. Notch signaling: a dance of proteins changing partners. Exp Cell Res. 2000;260(1):1–8.
Kadesch T. Notch signaling: the demise of elegant simplicity. Curr Opin Genet Dev. 2004;14(5):506–12.
Mumm JS, Kopan R. Notch signaling: from the outside in. Dev Biol. 2000;228(2):151–65.
Schweisguth F. Regulation of notch signaling activity. Current biology : CB. 2004;14(3):R129–38.
Ehebauer M, Hayward P, Arias AM. Notch, a universal arbiter of cell fate decisions. Science (New York, NY). 2006;314(5804):1414–5.
Ehebauer M, Hayward P, Martinez-Arias A. Notch signaling pathway. Science's STKE : signal transduction knowledge environment. 2006;2006(364):cm7.
Nusse R, Varmus HE. Many tumors induced by the mouse mammary tumor virus contain a provirus integrated in the same region of the host genome. Cell. 1982;31(1):99–109.
Niehrs C. The complex world of WNT receptor signalling. Nat Rev Mol Cell Biol. 2012;13(12):767–79.
Clevers H. Wnt/beta-catenin signaling in development and disease. Cell. 2006;127(3):469–80.
Perugorria MJ, Olaizola P, Labiano I, Esparza-Baquer A, Marzioni M, Marin JJG, et al. Wnt-β-catenin signalling in liver development, health and disease. Nat Rev Gastroenterol Hepatol. 2019;16(2):121–36.
Skronska-Wasek W, Mutze K, Baarsma HA, Bracke KR, Alsafadi HN, Lehmann M, et al. Reduced Frizzled Receptor 4 Expression Prevents WNT/β-Catenin-driven Alveolar Lung Repair in Chronic Obstructive Pulmonary Disease. Am J Respir Crit Care Med. 2017;196(2):172–85.
Nusse R, Clevers H. Wnt/β-Catenin Signaling, Disease, and Emerging Therapeutic Modalities. Cell. 2017;169(6):985–99.
Cruciat CM, Niehrs C. Secreted and transmembrane wnt inhibitors and activators. Cold Spring Harb Perspect Biol. 2013;5(3): a015081.
Reyes M, Flores T, Betancur D, Peña-Oyarzún D, Torres VA. Wnt/β-Catenin Signaling in Oral Carcinogenesis. Int J Mol Sci. 2020;21(13).
Muñoz-Castañeda JR, Rodelo-Haad C, Pendon-Ruiz de Mier MV, Martin-Malo A, Santamaria R, Rodriguez M. Klotho/FGF23 and Wnt Signaling as Important Players in the Comorbidities Associated with Chronic Kidney Disease. Toxins (Basel). 2020;12(3).
Liu J, Xiao Q, Xiao J, Niu C, Li Y, Zhang X, et al. Wnt/β-catenin signalling: function, biological mechanisms, and therapeutic opportunities. Signal Transduct Target Ther. 2022;7(1):3.
Binkley JM, Harris SR, Levangie PK, Pearl M, Guglielmino J, Kraus V, et al. Patient perspectives on breast cancer treatment side effects and the prospective surveillance model for physical rehabilitation for women with breast cancer. Cancer. 2012;118(8 Suppl):2207–16.
Fong DY, Ho JW, Hui BP, Lee AM, Macfarlane DJ, Leung SS, et al. Physical activity for cancer survivors: meta-analysis of randomised controlled trials. BMJ (Clinical research ed). 2012;344: e70.
Juvet LK, Thune I, Elvsaas I, Fors EA, Lundgren S, Bertheussen G, et al. The effect of exercise on fatigue and physical functioning in breast cancer patients during and after treatment and at 6 months follow-up: a meta-analysis. Breast (Edinburgh, Scotland). 2017;33:166–77.
Mijwel S, Jervaeus A, Bolam KA, Norrbom J, Bergh J, Rundqvist H, et al. High-intensity exercise during chemotherapy induces beneficial effects 12 months into breast cancer survivorship. J Cancer Surviv Res pract. 2019;13(2):244–56.
Zeng Y, Huang M, Cheng AS, Zhou Y, So WK. Meta-analysis of the effects of exercise intervention on quality of life in breast cancer survivors. Breast cancer (Tokyo, Japan). 2014;21(3):262–74.
McNeely ML, Campbell KL, Rowe BH, Klassen TP, Mackey JR, Courneya KS. Effects of exercise on breast cancer patients and survivors: a systematic review and meta-analysis. CMAJ Can Medical Assoc J. 2006;175(1):34–41.
Clevers H, Nusse R. Wnt/β-catenin signaling and disease. Cell. 2012;149(6):1192–205.
MacDonald BT, Tamai K, He X. Wnt/beta-catenin signaling: components, mechanisms, and diseases. Dev Cell. 2009;17(1):9–26.
Nelson WJ, Nusse R. Convergence of Wnt, beta-catenin, and cadherin pathways. Science (New York, NY). 2004;303(5663):1483–7.
Berschneider B, Königshoff M. WNT1 inducible signaling pathway protein 1 (WISP1): a novel mediator linking development and disease. Int J Biochem Cell Biol. 2011;43(3):306–9.
Gurbuz I, Chiquet-Ehrismann R. CCN4/WISP1 (WNT1 inducible signaling pathway protein 1): a focus on its role in cancer. Int J Biochem Cell Biol. 2015;62:142–6.
Chiang KC, Yeh CN, Chung LC, Feng TH, Sun CC, Chen MF, et al. WNT-1 inducible signaling pathway protein-1 enhances growth and tumorigenesis in human breast cancer. Sci Rep. 2015;5:8686.
Hörbelt T, Tacke C, Markova M, de Herzfeld Wiza D, Van de Velde F, Bekaert M, et al. The novel adipokine WISP1 associates with insulin resistance and impairs insulin action in human myotubes and mouse hepatocytes. Diabetologia. 2018;61(9):2054–65.
Murahovschi V, Pivovarova O, Ilkavets I, Dmitrieva RM, Döcke S, Keyhani-Nejad F, et al. WISP1 is a novel adipokine linked to inflammation in obesity. Diabetes. 2015;64(3):856–66.
Tacke C, Aleksandrova K, Rehfeldt M, Murahovschi V, Markova M, Kemper M, et al. Assessment of circulating Wnt1 inducible signalling pathway protein 1 (WISP-1)/CCN4 as a novel biomarker of obesity. J Cell Commun Signal. 2018;12(3):539–48.
Carmichael AR. Obesity and prognosis of breast cancer. Obesity Rev. 2006;7(4):333–40.
Cust AE, Stocks T, Lukanova A, Lundin E, Hallmans G, Kaaks R, et al. The influence of overweight and insulin resistance on breast cancer risk and tumour stage at diagnosis: a prospective study. Breast Cancer Res Treat. 2009;113(3):567–76.
Mendonça FM, de Sousa FR, Barbosa AL, Martins SC, Araújo RL, Soares R, et al. Metabolic syndrome and risk of cancer: which link? Metabolism. 2015;64(2):182–9.
Chang JS, Kim TH, Kong ID. Exercise intervention lowers aberrant serum WISP-1 levels with insulin resistance in breast cancer survivors: a randomized controlled trial. Sci Rep. 2020;10(1):10898.
Chen D, Zhang Y, Zhang M, Chang J, Zeng Z, Kou X, et al. Exercise attenuates brain aging by rescuing down-regulated Wnt/β-catenin signaling in aged rats. Front Aging Neurosci. 2020;12:105.
Bayod S, Mennella I, Sanchez-Roige S, Lalanza JF, Escorihuela RM, Camins A, et al. Wnt pathway regulation by long-term moderate exercise in rat hippocampus. Brain Res. 2014;1543:38–48.
Bashiri J, NourAzar A, Purrazi H. Effect of three months aerobic training on Wnt-signaling pathway in skeletal muscle of male rats. Razi J Med Sci. 2017;24(160):7–16.
Gardinier JD, Daly-Seiler C, Rostami N, Kundal S, Zhang C. Loss of the PTH/PTHrP receptor along the osteoblast lineage limits the anabolic response to exercise. PLoS ONE. 2019;14(1): e0211076.
Iura A, McNerny EG, Zhang Y, Kamiya N, Tantillo M, Lynch M, et al. Mechanical loading synergistically increases trabecular bone volume and improves mechanical properties in the mouse when BMP signaling is specifically ablated in osteoblasts. PLoS ONE. 2015;10(10): e0141345.
Herbst E, Gale T, Nagai K, Tashiro Y, Irrgang JJ, Anderst W, et al. Posterior tibial subchondral bone and meniscal slope correlate with in vivo internal tibial rotation. Orthop J Sports Med. 2017;5(7_suppl6):2325967117S00307.
Akpinar B, Thorhauer E, Tashman S, Irrgang JJ, Fu FH, Anderst WJ. Tibiofemoral cartilage contact differences between level walking and downhill running. Orthop J Sports Med. 2019;7(4):2325967119836164.
Bello M, Sousa MC, Neto G, Oliveira L, Guerras I, Mendes R, et al. The effect of a long-term, community-based exercise program on bone mineral density in postmenopausal women with pre-diabetes and type 2 diabetes. J Hum Kinet. 2014;43:43–8.
Gushiken M, Komiya I, Ueda S, Kobayashi J. Heel bone strength is related to lifestyle factors in Okinawan men with type 2 diabetes mellitus. J Diabetes Investig. 2015;6(2):150–7.
Chen X, Yang K, Sun P, Zhao R, Liu B, Lu P. Exercise improves bone formation by upregulating the Wnt3a/β-catenin signalling pathway in type 2 diabetic mice. Diabetol Metab Syndr. 2021;13(1):116.
Kumar A, Kaur H, Singh A. Neuropathic pain models caused by damage to central or peripheral nervous system. Pharmacol Rep PR. 2018;70(2):206–16.
Mukai M, Uchida K, Hirosawa N, Murakami K, Kuniyoshi K, Inoue G, et al. Wrapping with basic fibroblast growth factor-impregnated collagen sheet reduces rat sciatic nerve allodynia. J Orthop Res. 2019;37(10):2258–63.
Bernetti A, Agostini F, de Sire A, Mangone M, Tognolo L, Di Cesare A, et al. Neuropathic pain and rehabilitation: a systematic review of international guidelines. Diagnostics (Basel, Switzerland). 2021;11(1):74.
Finnerup NB, Kuner R, Jensen TS. Neuropathic pain: from mechanisms to treatment. Physiol Rev. 2021;101(1):259–301.
Itokazu T, Hayano Y, Takahashi R, Yamashita T. Involvement of Wnt/β-catenin signaling in the development of neuropathic pain. Neurosci Res. 2014;79:34–40.
Zhao Y, Yang Z. Effect of Wnt signaling pathway on pathogenesis and intervention of neuropathic pain. Exp Ther Med. 2018;16(4):3082–8.
Komiya Y, Habas R. Wnt signal transduction pathways. Organogenesis. 2008;4(2):68–75.
Cho YH, Kim JE, Seo TB. Effect of treadmill exercise on pain-related Wnt/β-catenin signaling pathway in dorsal root ganglion neurons at the early phase regeneration of the injured sciatic nerve. J Exerc Rehabil. 2021;17(2):96–102.
Morgan TH. The theory of the gene. Am Nat. 1917;51(609):513–44.
Kidd S, Kelley MR, Young MW. Sequence of the notch locus of Drosophila melanogaster: relationship of the encoded protein to mammalian clotting and growth factors. Mol Cell Biol. 1986;6(9):3094–108.
Wharton KA, Johansen KM, Xu T, Artavanis-Tsakonas S. Nucleotide sequence from the neurogenic locus notch implies a gene product that shares homology with proteins containing EGF-like repeats. Cell. 1985;43(3 Pt 2):567–81.
Blaumueller CM, Qi H, Zagouras P, Artavanis-Tsakonas S. Intracellular cleavage of Notch leads to a heterodimeric receptor on the plasma membrane. Cell. 1997;90(2):281–91.
Radtke F, Raj K. The role of Notch in tumorigenesis: oncogene or tumour suppressor? Nat Rev Cancer. 2003;3(10):756–67.
Okajima T, Irvine KD. Regulation of notch signaling by o-linked fucose. Cell. 2002;111(6):893–904.
Okajima T, Xu A, Irvine KD. Modulation of notch-ligand binding by protein O-fucosyltransferase 1 and fringe. J Biol Chem. 2003;278(43):42340–5.
Rebay I, Fleming RJ, Fehon RG, Cherbas L, Cherbas P, Artavanis-Tsakonas S. Specific EGF repeats of Notch mediate interactions with Delta and Serrate: implications for Notch as a multifunctional receptor. Cell. 1991;67(4):687–99.
Moloney DJ, Panin VM, Johnston SH, Chen J, Shao L, Wilson R, et al. Fringe is a glycosyltransferase that modifies Notch. Nature. 2000;406(6794):369–75.
Brückner K, Perez L, Clausen H, Cohen S. Glycosyltransferase activity of Fringe modulates Notch-Delta interactions. Nature. 2000;406(6794):411–5.
Haines N, Irvine KD. Glycosylation regulates Notch signalling. Nat Rev Mol Cell Biol. 2003;4(10):786–97.
Nam Y, Sliz P, Song L, Aster JC, Blacklow SC. Structural basis for cooperativity in recruitment of MAML coactivators to Notch transcription complexes. Cell. 2006;124(5):973–83.
Radtke F, Wilson A, Mancini SJ, MacDonald HR. Notch regulation of lymphocyte development and function. Nat Immunol. 2004;5(3):247–53.
Wilson JJ, Kovall RA. Crystal structure of the CSL-Notch-Mastermind ternary complex bound to DNA. Cell. 2006;124(5):985–96.
Ong CT, Cheng HT, Chang LW, Ohtsuka T, Kageyama R, Stormo GD, et al. Target selectivity of vertebrate notch proteins. Collaboration between discrete domains and CSL-binding site architecture determines activation probability. J Biol Chem. 2006;281(8):5106–19.
Bray SJ. Notch signalling: a simple pathway becomes complex. Nat Rev Mol Cell Biol. 2006;7(9):678–89.
Shah S, Lee SF, Tabuchi K, Hao YH, Yu C, LaPlant Q, et al. Nicastrin functions as a gamma-secretase-substrate receptor. Cell. 2005;122(3):435–47.
Kao HY, Ordentlich P, Koyano-Nakagawa N, Tang Z, Downes M, Kintner CR, et al. A histone deacetylase corepressor complex regulates the Notch signal transduction pathway. Genes Dev. 1998;12(15):2269–77.
Mulligan P, Yang F, Di Stefano L, Ji JY, Ouyang J, Nishikawa JL, et al. A SIRT1-LSD1 corepressor complex regulates Notch target gene expression and development. Mol Cell. 2011;42(5):689–99.
Nagel AC, Krejci A, Tenin G, Bravo-Patiño A, Bray S, Maier D, et al. Hairless-mediated repression of notch target genes requires the combined activity of Groucho and CtBP corepressors. Mol Cell Biol. 2005;25(23):10433–41.
Yatim A, Benne C, Sobhian B, Laurent-Chabalier S, Deas O, Judde JG, et al. NOTCH1 nuclear interactome reveals key regulators of its transcriptional activity and oncogenic function. Mol Cell. 2012;48(3):445–58.
Hamidi H, Gustafason D, Pellegrini M, Gasson J. Identification of novel targets of CSL-dependent Notch signaling in hematopoiesis. PLoS ONE. 2011;6(5): e20022.
Ntziachristos P, Tsirigos A, Van Vlierberghe P, Nedjic J, Trimarchi T, Flaherty MS, et al. Genetic inactivation of the polycomb repressive complex 2 in T cell acute lymphoblastic leukemia. Nat Med. 2012;18(2):298–301.
Palomero T, Lim WK, Odom DT, Sulis ML, Real PJ, Margolin A, et al. NOTCH1 directly regulates c-MYC and activates a feed-forward-loop transcriptional network promoting leukemic cell growth. Proc Natl Acad Sci USA. 2006;103(48):18261–6.
Weng AP, Millholland JM, Yashiro-Ohtani Y, Arcangeli ML, Lau A, Wai C, et al. c-Myc is an important direct target of Notch1 in T-cell acute lymphoblastic leukemia/lymphoma. Genes Dev. 2006;20(15):2096–109.
Pitsouli C, Delidakis C. The interplay between DSL proteins and ubiquitin ligases in Notch signaling. Development (Cambridge, England). 2005;132(18):4041–50.
Gupta-Rossi N, Le Bail O, Gonen H, Brou C, Logeat F, Six E, et al. Functional interaction between SEL-10, an F-box protein, and the nuclear form of activated Notch1 receptor. J Biol Chem. 2001;276(37):34371–8.
Gupta-Rossi N, Six E, LeBail O, Logeat F, Chastagner P, Olry A, et al. Monoubiquitination and endocytosis direct gamma-secretase cleavage of activated Notch receptor. J Cell Biol. 2004;166(1):73–83.
Oberg C, Li J, Pauley A, Wolf E, Gurney M, Lendahl U. The Notch intracellular domain is ubiquitinated and negatively regulated by the mammalian Sel-10 homolog. J Biol Chem. 2001;276(38):35847–53.
Wu G, Lyapina S, Das I, Li J, Gurney M, Pauley A, et al. SEL-10 is an inhibitor of notch signaling that targets notch for ubiquitin-mediated protein degradation. Mol Cell Biol. 2001;21(21):7403–15.
Nakayama KI, Nakayama K. Ubiquitin ligases: cell-cycle control and cancer. Nat Rev Cancer. 2006;6(5):369–81.
Wei W, Jin J, Schlisio S, Harper JW, Kaelin WG Jr. The v-Jun point mutation allows c-Jun to escape GSK3-dependent recognition and destruction by the Fbw7 ubiquitin ligase. Cancer Cell. 2005;8(1):25–33.
Welcker M, Orian A, Jin J, Grim JE, Harper JW, Eisenman RN, et al. The Fbw7 tumor suppressor regulates glycogen synthase kinase 3 phosphorylation-dependent c-Myc protein degradation. Proc Natl Acad Sci USA. 2004;101(24):9085–90.
Welcker M, Singer J, Loeb KR, Grim J, Bloecher A, Gurien-West M, et al. Multisite phosphorylation by Cdk2 and GSK3 controls cyclin E degradation. Mol Cell. 2003;12(2):381–92.
Kemp Z, Rowan A, Chambers W, Wortham N, Halford S, Sieber O, et al. CDC4 mutations occur in a subset of colorectal cancers but are not predicted to cause loss of function and are not associated with chromosomal instability. Can Res. 2005;65(24):11361–6.
Kwak EL, Moberg KH, Wahrer DC, Quinn JE, Gilmore PM, Graham CA, et al. Infrequent mutations of Archipelago (hAGO, hCDC4, Fbw7) in primary ovarian cancer. Gynecol Oncol. 2005;98(1):124–8.
Strohmaier H, Spruck CH, Kaiser P, Won KA, Sangfelt O, Reed SI. Human F-box protein hCdc4 targets cyclin E for proteolysis and is mutated in a breast cancer cell line. Nature. 2001;413(6853):316–22.
O’Neil J, Grim J, Strack P, Rao S, Tibbitts D, Winter C, et al. FBW7 mutations in leukemic cells mediate NOTCH pathway activation and resistance to gamma-secretase inhibitors. J Exp Med. 2007;204(8):1813–24.
Thompson BJ, Buonamici S, Sulis ML, Palomero T, Vilimas T, Basso G, et al. The SCFFBW7 ubiquitin ligase complex as a tumor suppressor in T cell leukemia. J Exp Med. 2007;204(8):1825–35.
Ye X, Nalepa G, Welcker M, Kessler BM, Spooner E, Qin J, et al. Recognition of phosphodegron motifs in human cyclin E by the SCF(Fbw7) ubiquitin ligase. J Biol Chem. 2004;279(48):50110–9.
Fryer CJ, White JB, Jones KA. Mastermind recruits CycC:CDK8 to phosphorylate the Notch ICD and coordinate activation with turnover. Mol Cell. 2004;16(4):509–20.
Mo JS, Kim MY, Han SO, Kim IS, Ann EJ, Lee KS, et al. Integrin-linked kinase controls Notch1 signaling by down-regulation of protein stability through Fbw7 ubiquitin ligase. Mol Cell Biol. 2007;27(15):5565–74.
Saint Just Ribeiro M, Hansson ML, Lindberg MJ, Popko-Scibor AE, Wallberg AE. GSK3beta is a negative regulator of the transcriptional coactivator MAML1. Nucl Acids Res. 2009;37(20):6691–700.
Lobry C, Oh P, Mansour MR, Look AT, Aifantis I. Notch signaling: switching an oncogene to a tumor suppressor. Blood. 2014;123(16):2451–9.
Englund DA, Figueiredo VC, Dungan CM, Murach KA, Peck BD, Petrosino JM, et al. Satellite cell depletion disrupts transcriptional coordination and muscle adaptation to exercise. Function (Oxford, England). 2021;2(1):033.
Yin H, Price F, Rudnicki MA. Satellite cells and the muscle stem cell niche. Physiol Rev. 2013;93(1):23–67.
Clarkson PM, Hubal MJ. Exercise-induced muscle damage in humans. Am J Phys Med Rehabil. 2002;81(11 Suppl):S52-69.
Darr KC, Schultz E. Exercise-induced satellite cell activation in growing and mature skeletal muscle. J Appl Physiol (Bethesda, Md: 1985). 1987;63(5):1816–21.
Zhong W, Jiang MM, Weinmaster G, Jan LY, Jan YN. Differential expression of mammalian Numb, Numblike and Notch1 suggests distinct roles during mouse cortical neurogenesis. Development (Cambridge, England). 1997;124(10):1887–97.
Bubak MP, Stout K, Tomtschik J, Peterson E, Cardozo CP, Graham ZA, et al. Notch, Numb and Numb-like responses to exercise-induced muscle damage in human skeletal muscle. Exp Physiol. 2022;107(8):800–6.
Brandt MD, Maass A, Kempermann G, Storch A. Physical exercise increases Notch activity, proliferation and cell cycle exit of type-3 progenitor cells in adult hippocampal neurogenesis. Eur J Neurosci. 2010;32(8):1256–64.
Lee SJ, McPherron AC. Regulation of myostatin activity and muscle growth. Proc Natl Acad Sci USA. 2001;98(16):9306–11.
McPherron AC, Lawler AM, Lee SJ. Regulation of skeletal muscle mass in mice by a new TGF-beta superfamily member. Nature. 1997;387(6628):83–90.
Welle S, Burgess K, Mehta S. Stimulation of skeletal muscle myofibrillar protein synthesis, p70 S6 kinase phosphorylation, and ribosomal protein S6 phosphorylation by inhibition of myostatin in mature mice. Am J Physiol Endocrinol Metab. 2009;296(3):E567–72.
Hulmi JJ, Oliveira BM, Silvennoinen M, Hoogaars WM, Ma H, Pierre P, et al. Muscle protein synthesis, mTORC1/MAPK/Hippo signaling, and capillary density are altered by blocking of myostatin and activins. Am J Physiol Endocrinol Metab. 2013;304(1):E41-50.
Sutrave P, Kelly AM, Hughes SH. ski can cause selective growth of skeletal muscle in transgenic mice. Genes Dev. 1990;4(9):1462–72.
Sutrave P, Leferovich JM, Kelly AM, Hughes SH. The induction of skeletal muscle hypertrophy by a ski transgene is promoter-dependent. Gene. 2000;241(1):107–16.
Hulmi JJ, Ahtiainen JP, Kaasalainen T, Pöllänen E, Häkkinen K, Alen M, et al. Postexercise myostatin and activin IIb mRNA levels: effects of strength training. Med Sci Sports Exerc. 2007;39(2):289–97.
Louis E, Raue U, Yang Y, Jemiolo B, Trappe S. Time course of proteolytic, cytokine, and myostatin gene expression after acute exercise in human skeletal muscle. J Appl Physiol (Bethesda, Md: 1985). 2007;103(5):1744–51.
Kim JS, Petrella JK, Cross JM, Bamman MM. Load-mediated downregulation of myostatin mRNA is not sufficient to promote myofiber hypertrophy in humans: a cluster analysis. J Appl Physiol. 2007;103(5):1488–95.
Baar K, Esser K. Phosphorylation of p70(S6k) correlates with increased skeletal muscle mass following resistance exercise. Am J Physiol. 1999;276(1):C120–7.
Terzis G, Georgiadis G, Stratakos G, Vogiatzis I, Kavouras S, Manta P, et al. Resistance exercise-induced increase in muscle mass correlates with p70S6 kinase phosphorylation in human subjects. Eur J Appl Physiol. 2008;102(2):145–52.
Dalbo VJ, Roberts MD, Sunderland KL, Poole CN, Stout JR, Beck TW, et al. Acute loading and aging effects on myostatin pathway biomarkers in human skeletal muscle after three sequential bouts of resistance exercise. J Gerontol A Biol Sci Med Sci. 2011;66(8):855–65.
Marshall A, Salerno MS, Thomas M, Davies T, Berry C, Dyer K, et al. Mighty is a novel promyogenic factor in skeletal myogenesis. Exp Cell Res. 2008;314(5):1013–29.
Salerno MS, Dyer K, Bracegirdle J, Platt L, Thomas M, Siriett V, et al. Akirin1 (Mighty), a novel promyogenic factor regulates muscle regeneration and cell chemotaxis. Exp Cell Res. 2009;315(12):2012–21.
Conboy IM, Conboy MJ, Smythe GM, Rando TA. Notch-mediated restoration of regenerative potential to aged muscle. Science (New York, NY). 2003;302(5650):1575–7.
Carlson ME, Hsu M, Conboy IM. Imbalance between pSmad3 and Notch induces CDK inhibitors in old muscle stem cells. Nature. 2008;454(7203):528–32.
Akiho M, Nakashima H, Sakata M, Yamasa Y, Yamaguchi A, Sakuma K. Expression profile of Notch-1 in mechanically overloaded plantaris muscle of mice. Life Sci. 2010;86(1–2):59–65.
Davis E, Jensen CH, Schroder HD, Farnir F, Shay-Hadfield T, Kliem A, et al. Ectopic expression of DLK1 protein in skeletal muscle of padumnal heterozygotes causes the callipyge phenotype. Current biology : CB. 2004;14(20):1858–62.
Guo S, Liu M, Gonzalez-Perez RR. Role of Notch and its oncogenic signaling crosstalk in breast cancer. Biochem Biophys Acta. 2011;1815(2):197–213.
Chu J, Jeffries S, Norton JE, Capobianco AJ, Bresnick EH. Repression of activator protein-1-mediated transcriptional activation by the Notch-1 intracellular domain. J Biol Chem. 2002;277(9):7587–97.
MacKenzie MG, Hamilton DL, Pepin M, Patton A, Baar K. Inhibition of myostatin signaling through Notch activation following acute resistance exercise. PLoS ONE. 2013;8(7): e68743.
Acknowledgements
Research Fund Project of Hebei Provincial Health Commission. Project title: A Study on the Promotion of Health for University Teachers through Exercise Prescription Management in the Context of Body-Medicine Integration. Approval number: 20231552.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, Y., Wang, G., Wei, Z. et al. Wnt, notch signaling and exercise: what are their functions?. Human Cell (2024). https://doi.org/10.1007/s13577-024-01036-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13577-024-01036-3