Abstract
Purpose
The Chromobox (CBX) family proteins are crucial elements of the epigenetic regulatory machinery and play a significant role in the development and advancement of cancer. Nevertheless, there is limited understanding regarding the role of CBXs in development or progression of prostate cancer (PCa). Our objective is to develop a unique prognostic model associated with CBXs to improve the accuracy of predicting outcomes of patients with PCa.
Methods
Data from TCGA and GEO databases were analyzed to assess differential expression, prognostic value, gene pathway enrichment, and immune cell infiltration. COX regression analysis was utilized to identify the independent prognostic factors that impact disease-free survival (DFS). The expression of CBX2 and FOXP3+ cells infiltration was verified by immunohistochemical staining of clinical tissue sections. In vitro proliferation, migration and invasion assay were conducted to examine the function of CBX2. RNA-seq was employed to examine the CBX2 related pathway enrichment.
Results
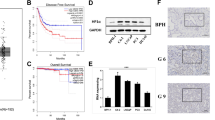
CBX2, CBX3, CBX4, and CBX8 were upregulated, while CBX6 and CBX7 were downregulated in PCa tissues. CBXs expression varied by stage and grade. Elevated expression of CBX1, CBX2, CBX3, CBX4 and CBX8 is correlated with poor outcome. CBX2 expression, T stage, and Gleason score were independent prognostic factors. The expression level of CBX2 in PCa tissues was significantly higher than that in adjacent normal tissues. More Treg infiltration was observed in the group with high CBX2 expression. CBX2 expression affected PCa cell growth, migration, and invasion.
Conclusions
CBX2 is involved in the development and advancement of PCa, suggesting its potential as a reliable prognostic indicator for PCa patients.














Similar content being viewed by others
Data availability
The data for bioinformatics analysis are publicly accessible, and the specific details can be found in the materials and methods section of the paper.
References
H. Sung, J. Ferlay, R.L. Siegel, M. Laversanne, I. Soerjomataram, A. Jemal, F. Bray, CA Cancer J. Clin. 71, 209–249 (2021). https://doi.org/10.3322/caac.21660
R.L. Siegel, K.D. Miller, H.E. Fuchs, A. Jemal, CA Cancer J. Clin. 72, 7–33 (2022). https://doi.org/10.3322/caac.21708
S. Gillessen, A. Bossi, I.D. Davis, J. de Bono, K. Fizazi, N.D. James, N. Mottet, N. Shore, E. Small, M. Smith, C. Sweeney, B. Tombal, E.S. Antonarakis, A.M. Aparicio, A.J. Armstrong, G. Attard, T.M. Beer, H. Beltran, A. Bjartell, P. Blanchard, A. Briganti, R.G. Bristow, M. Bulbul, O. Caffo, D. Castellano, E. Castro, H.H. Cheng, K.N. Chi, S. Chowdhury, C.S. Clarke, N. Clarke, G. Daugaard, M. De Santis, I. Duran, R. Eeles, E. Efstathiou, J. Efstathiou, O. Ngozi Ekeke, C.P. Evans, S. Fanti, F.Y. Feng, V. Fonteyne, N. Fossati, M. Frydenberg, D. George, M. Gleave, G. Gravis, S. Halabi, D. Heinrich, K. Herrmann, C. Higano, M.S. Hofman, L.G. Horvath, M. Hussain, B.A. Jereczek-Fossa, R. Jones, R. Kanesvaran, P.L. Kellokumpu-Lehtinen, R.B. Khauli, L. Klotz, G. Kramer, R. Leibowitz, C.J. Logothetis, B.A. Mahal, F. Maluf, J. Mateo, D. Matheson, N. Mehra, A. Merseburger, A.K. Morgans, M.J. Morris, H. Mrabti, D. Mukherji, D.G. Murphy, V. Murthy, P.L. Nguyen, W.K. Oh, P. Ost, J.M. O’Sullivan, A.R. Padhani, C. Pezaro, D.M.C. Poon, C.C. Pritchard, D.M. Rabah, D. Rathkopf, R.E. Reiter, M.A. Rubin, C.J. Ryan, F. Saad, J. Pablo Sade, O.A. Sartor, H.I. Scher, N. Sharifi, I. Skoneczna, H. Soule, D.E. Spratt, S. Srinivas, C.N. Sternberg, T. Steuber, H. Suzuki, M.R. Sydes, M.E. Taplin, D. Tilki, L. Turkeri, F. Turco, H. Uemura, H. Uemura, Y. Urun, C.L. Vale, I. van Oort, N. Vapiwala, J. Walz, K. Yamoah, D. Ye, E.Y. Yu, A. Zapatero, T. Zilli, A. Omlin, Eur. Urol. 83, 267–293 (2023). https://doi.org/10.1016/j.eururo.2022.11.002
N.I. Simon, C. Parker, T.A. Hope, C.J. Paller, Am. Soc. Clin. Oncol. Educ. Book 42, 1–8 (2022). https://doi.org/10.1200/EDBK_351033
S. Goel, V. Bhatia, T. Biswas, B. Ateeq, Semin. Cancer Biol. 83, 136–151 (2022). https://doi.org/10.1016/j.semcancer.2021.01.009
J. Kim, R.E. Kingston, J. Biosci. 45 (2020)
R.G. Ma, Y. Zhang, T.T. Sun, B. Cheng, J. Zhejiang Univ. Sci. B 15, 412–428 (2014). https://doi.org/10.1631/jzus.B1400077
H. Zhou, Y. Xiong, Z. Liu, S. Hou, T. Zhou, Cancer Cell Int. 21, 402 (2021). https://doi.org/10.1186/s12935-021-02106-4
C. Zhang, L. Chang, Y. Yao, C. Chao, Z. Ge, C. Fan, H. Yu, B. Wang, J. Yang, Front. Genet. 12, 771062 (2021). https://doi.org/10.3389/fgene.2021.771062
K. Hu, L. Yao, Z. Xu, Y. Yan, J. Li, Front. Cell Dev. Biol. 10, 832354 (2022). https://doi.org/10.3389/fcell.2022.832354
J. Liu, H. Shen, X. Chen, Y. Ding, H. Wang, N. Xu, L. Teng, Genes (Basel) 13 (2022). https://doi.org/10.3390/genes13091582
A.A.T. Naqvi, S.A.M. Rizvi, M.I. Hassan, Biochim. Biophys. Acta Mol. Basis Dis. 1869, 166561 (2023). https://doi.org/10.1016/j.bbadis.2022.166561
L. Cheng, R. Montironi, D.G. Bostwick, A. Lopez-Beltran, D.M. Berney, Histopathology 60, 87–117 (2012). https://doi.org/10.1111/j.1365-2559.2011.04025.x
F. Zhou, L. Chen, P. Lu, Y. Cao, C. Deng, G. Liu, BMC Cancer 23, 641 (2023). https://doi.org/10.1186/s12885-023-11108-6
X. Li, L. Li, X. Xiong, Q. Kuang, M. Peng, K. Zhu, P. Luo, Diagnostics (Basel) 13 (2023). https://doi.org/10.3390/diagnostics13081393
D. Liao, X. Liu, L. He, Y. Yao, X. Yuan, P. Feng, C. Li, Y. Liu, Iran. J. Basic Med. Sci. 26, 468–477 (2023). https://doi.org/10.22038/IJBMS.2023.64845.14281
L. Wang, L. Zhao, Y. Zhang, S. Shao, Q. Ning, X. Zhao, M. Luo, Clin. Breast Cancer 23, e206–e218 (2023). https://doi.org/10.1016/j.clbc.2023.02.007
A.M. Grimaldi, O. Affinito, M. Salvatore, M. Franzese, Diagnostics (Basel) 12 (2022). https://doi.org/10.3390/diagnostics12102452
Z.Q. Zheng, G.Q. Yuan, N.L. Kang, Q.Q. Nie, G.G. Zhang, Z. Wang, Front. Neurol. 13, 912039 (2022). https://doi.org/10.3389/fneur.2022.912039
X. Meng, R. Cao, X. Liu, B. Fu, L. Luo, M. Jiang, K. Wang, Y. Liu, Q. Zhu, C. Yang, L. Zhou, Oncol. Lett. 25, 172 (2023). https://doi.org/10.3892/ol.2023.13758
J. Wang, B. Yang, X. Zhang, S. Liu, X. Pan, C. Ma, S. Ma, D. Yu, W. Wu, Int. J. Oncol. 62 (2023). https://doi.org/10.3892/ijo.2023.5484
L. Li, W. Zhang, J. Qiu, W. Zhang, M. Lu, J. Wang, Y. Jin, Q. Xi, Stem. Cells Int. 2023, 4500561 (2023). https://doi.org/10.1155/2023/4500561
Y. Xu, S. Pan, Y. Song, C. Pan, C. Chen, X. Zhu, J. Cancer 11, 5198–5209 (2020). https://doi.org/10.7150/jca.44475
Z. Yang, Q. Zi, K. Xu, C. Wang, Q. Chi, Int. Immunopharmacol. 90, 107238 (2021). https://doi.org/10.1016/j.intimp.2020.107238
S. Zheng, P. Lv, J. Su, K. Miao, H. Xu, M. Li, Am. J. Transl. Res. 11, 1668–1682 (2019)
M. Zeng, B. Li, L. Yang, Q. Guan, Mol. Med. Rep. 23 (2021). https://doi.org/10.3892/mmr.2020.11776
Y. Xing, J. Ba-Tu, C. Dong, X. Cao, B. Li, X. Jia, Y. Juan, X. Lv, H. Zhang, N. Qin, W. Han, D. Wang, X. Qi, Y. Wang, X. Hao, S. Zhang, X. Du, H. Wang, M. Wang, Cell Death Dis. 14, 782 (2023). https://doi.org/10.1038/s41419-023-06304-y
Y.J. Zhang, L.Y. Zhao, X. He, R.F. Yao, F. Lu, B.N. Lu, Z.R. Pang, Aging (Albany NY) 14, 6227–6254 (2022). https://doi.org/10.18632/aging.204214
F.F. Hu, H. Chen, Y. Duan, B. Lan, C.J. Liu, H. Hu, X. Dong, Q. Zhang, Y.M. Cheng, M. Liu, A.Y. Guo, C. Xuan, Mol. Ther. Nucleic Acids 27, 670–684 (2022). https://doi.org/10.1016/j.omtn.2021.12.032
L.W. Brubaker, D.S. Backos, V.T. Nguyen, P. Reigan, T.M. Yamamoto, E.R. Woodruff, R. Iwanaga, M.F. Wempe, V. Kumar, C. Persenaire, Z.L. Watson, B.G. Bitler, Expert Opin. Ther. Targets 27, 361–371 (2023). https://doi.org/10.1080/14728222.2023.2218614
A. Di Costanzo, N. Del Gaudio, L. Conte, C. Dell’Aversana, M. Vermeulen, H. de The, A. Migliaccio, A. Nebbioso, L. Altucci, Oncogene 37, 2559–2572 (2018). https://doi.org/10.1038/s41388-018-0143-1
Funding
Guangdong Province key areas research and development plan (2023B1111030006), National Natural Science Foundation of China (82072841 and 82372766), Natural Science Foundation of Guangdong Province (2021A1515010199), Key Areas Research and Development Program of Guangdong (2020B111114002), Guangdong Provincial Clinical Research Center for Urological Diseases (2020B1111170006) and Guangdong Science and Technology Department (2020B1212060018). Fundamental Research Funds for the Central Universities, Sun Yat-sen University (1320223001), Medical Science and Technology Research Fund of Guangdong Province (A2022541).
Author information
Authors and Affiliations
Contributions
K.W.X. and J.G.H. designed and directed the research. X.T.X. and C.L. analyzed the data and drafted the manuscript. J.W.L. performed the in vitro experiments and revised the manuscript. J.Y.S., K.X.G. and J.T.H. collected references and revised the manuscript. Y.M., Y.F.X. and D.G.K. were responsible for cell culture and data collection. C.L. revised the manuscript. The final manuscript was approved by all authors.
Corresponding authors
Ethics declarations
Ethical approval
The study was approved by the Sun Yat-sen Memorial Hospital ethical committee (SYSKY-2023-1156-01).
Competing interests
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, X., Lai, C., Luo, J. et al. The predictive significance of chromobox family members in prostate cancer in humans. Cell Oncol. (2024). https://doi.org/10.1007/s13402-024-00929-7
Accepted:
Published:
DOI: https://doi.org/10.1007/s13402-024-00929-7