Abstract
Purpose
Serine metabolism is frequently dysregulated in many types of cancers and the tumor suppressor p53 is recently emerging as a key regulator of serine metabolism. However, the detailed mechanism remains unknown. Here, we investigate the role and underlying mechanisms of how p53 regulates the serine synthesis pathway (SSP) in bladder cancer (BLCA).
Methods
Two BLCA cell lines RT-4 (WT p53) and RT-112 (p53 R248Q) were manipulated by applying CRISPR/Cas9 to examine metabolic differences under WT and mutant p53 status. Liquid chromatography-tandem mass spectrometry (LC-MS/MS) and non-targeted metabolomics analysis were adopted to identify metabolomes changes between WT and p53 mutant BLCA cells. Bioinformatics analysis using the cancer genome atlas and Gene Expression Omnibus datasets and immunohistochemistry (IHC) staining was used to investigate PHGDH expression. Loss-of-function of PHGDH and subcutaneous xenograft model was adopted to investigate the function of PHGDH in mice BLCA. Chromatin immunoprecipitation (Ch-IP) assay was performed to analyze the relationships between YY1, p53, SIRT1 and PHGDH expression.
Results
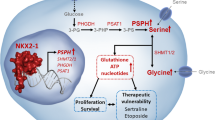
SSP is one of the most prominent dysregulated metabolic pathways by comparing the metabolomes changes between wild-type (WT) p53 and mutant p53 of BLCA cells. TP53 gene mutation shows a positive correlation with PHGDH expression in TCGA-BLCA database. PHGDH depletion disturbs the reactive oxygen species homeostasis and attenuates the xenograft growth in the mouse model. Further, we demonstrate WT p53 inhibits PHGDH expression by recruiting SIRT1 to the PHGDH promoter. Interestingly, the DNA binding motifs of YY1 and p53 in the PHGDH promoter are partially overlapped which causes competition between the two transcription factors. This competitive regulation of PHGDH is functionally linked to the xenograft growth in mice.
Conclusion
YY1 drives PHGDH expression in the context of mutant p53 and promotes bladder tumorigenesis, which preliminarily explains the relationship between high-frequency mutations of p53 and dysfunctional serine metabolism in bladder cancer.







Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
F. Massari, C. Ciccarese, M. Santoni, R. Iacovelli, R. Mazzucchelli, F. Piva, M. Scarpelli, R. Berardi, G. Tortora, A. Lopez-Beltran, L. Cheng, R. Montironi, Metabolic phenotype of bladder cancer. Cancer Treat. Rev. 45, 46–57 (2016). https://doi.org/10.1016/j.ctrv.2016.03.005
B.L. Woolbright, M. Ayres, J.A. Taylor III, Metabolic changes in bladder cancer. Urol. Oncol. 36, 327–337 (2018). https://doi.org/10.1016/j.urolonc.2018.04.010
E.R. Kastenhuber, S.W. Lowe, Putting p53 in Context. Cell 170, 1062–1078 (2017). https://doi.org/10.1016/j.cell.2017.08.028
P. Berggren, G. Steineck, J. Adolfsson, J. Hansson, O. Jansson, P. Larsson, B. Sandstedt, H. Wijkstrom, K. Hemminki, p53 mutations in urinary bladder cancer. Br. J. Cancer 84, 1505–1511 (2001). https://doi.org/10.1054/bjoc.2001.1823
W.C. Kusser, X. Miao, B.W. Glickman, J.M. Friedland, N. Rothman, G.P. Hemstreet, J. Mellot, D.C. Swan, P.A. Schulte, R.B. Hayes, p53 mutations in human bladder cancer. Environ. Mol. Mutagen. 24, 156–160 (1994). https://doi.org/10.1002/em.2850240303
J.C. Schroeder, K. Conway, Y. Li, K. Mistry, D.A. Bell, J.A. Taylor, p53 mutations in bladder cancer: evidence for exogenous versus endogenous risk factors. Cancer Res. 63, 7530–7538 (2003)
J. Liu, C. Zhang, W. Hu, Z. Feng, Tumor suppressor p53 and its mutants in cancer metabolism. Cancer Lett. 356, 197–203 (2015). https://doi.org/10.1016/j.canlet.2013.12.025
P.A. Muller, K.H. Vousden, Mutant p53 in cancer: new functions and therapeutic opportunities. Cancer Cell 25, 304–317 (2014). https://doi.org/10.1016/j.ccr.2014.01.021
L. Sun, L. Song, Q. Wan, G. Wu, X. Li, Y. Wang, J. Wang, Z. Liu, X. Zhong, X. He, S. Shen, X. Pan, A. Li, Y. Wang, P. Gao, H. Tang, H. Zhang, Cmyc-mediated activation of serine biosynthesis pathway is critical for cancer progression under nutrient deprivation conditions. Cell. Res. 25, 429–444 (2015). https://doi.org/10.1038/cr.2015.33
Q. Li, J. Qiu, H. Yang, G. Sun, Y. Hu, D. Zhu, Z. Deng, X. Wang, J. Tang, R. Jiang, Kinesin family member 15 promotes cancer stem cell phenotype and malignancy via reactive oxygen species imbalance in hepatocellular carcinoma. Cancer Lett (2019). https://doi.org/10.1016/j.canlet.2019.11.008
L. Wei, D. Lee, C.T. Law, M.S. Zhang, J. Shen, D.W. Chin, A. Zhang, F.H. Tsang, C.L. Wong, I.O. Ng, C.C. Wong, C.M. Wong, Genome-wide CRISPR/Cas9 library screening identified PHGDH as a critical driver for Sorafenib resistance in HCC. Nat. Commun. 10, 4681 (2019). https://doi.org/10.1038/s41467-019-12606-7
B. Zhang, A. Zheng, P. Hydbring, G. Ambroise, A.T. Ouchida, M. Goiny, H. Vakifahmetoglu-Norberg, E. Norberg, PHGDH defines a metabolic subtype in lung adenocarcinomas with poor prognosis. Cell. Rep. 19, 2289–2303 (2017). https://doi.org/10.1016/j.celrep.2017.05.067
D. Samanta, Y. Park, S.A. Andrabi, L.M. Shelton, D.M. Gilkes, G.L. Semenza, PHGDH expression is required for mitochondrial redox homeostasis, breast Cancer stem cell maintenance, and lung metastasis. Cancer Res. 76, 4430–4442 (2016). https://doi.org/10.1158/0008-5472.CAN-16-0530
E. Gronroos, A.A. Terentiev, T. Punga, J. Ericsson, YY1 inhibits the activation of the p53 tumor suppressor in response to genotoxic stress. Proc. Natl. Acad. Sci. U.S.A 101, 12165–12170 (2004). https://doi.org/10.1073/pnas.0402283101
C. Zhang, X. Zhang, W. Zhao, C. Zeng, W. Li, B. Li, X. Luo, J. Li, J. Jiang, B. Deng, D.W. McComb, Y. Dong, Chemotherapy drugs derived nanoparticles encapsulating mRNA encoding tumor suppressor proteins to treat triple-negative breast cancer. Nano Res. 12, 855–861 (2019). https://doi.org/10.1007/s12274-019-2308-9
F.A. Ran, P.D. Hsu, J. Wright, V. Agarwala, D.A. Scott, F. Zhang, Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013). https://doi.org/10.1038/nprot.2013.143
J. Shao, J. Lu, W. Zhu, H. Yu, X. Jing, Y.L. Wang, X. Wang, X.J. Wang, Derepression of LOXL4 inhibits liver cancer growth by reactivating compromised p53. Cell. Death Differ. 26, 2237–2252 (2019). https://doi.org/10.1038/s41418-019-0293-x
Y. Wu, D. Liang, Y. Wang, M. Bai, W. Tang, S. Bao, Z. Yan, D. Li, J. Li, Correction of a genetic disease in mouse via use of CRISPR-Cas9. Cell. Stem Cell 13, 659–662 (2013). https://doi.org/10.1016/j.stem.2013.10.016
J.P. Zhang, X.L. Li, G.H. Li, W. Chen, C. Arakaki, G.D. Botimer, D. Baylink, L. Zhang, W. Wen, Y.W. Fu, J. Xu, N. Chun, W. Yuan, T. Cheng, X.B. Zhang (2017) Efficient precise knockin with a double cut HDR donor after CRISPR/Cas9-mediated double-stranded DNA cleavage. Genome Biol. 18, 35. https://doi.org/10.1186/s13059-017-1164-8
X. Wang, R. Liu, X. Qu, H. Yu, H. Chu, Y. Zhang, W. Zhu, X. Wu, H. Gao, B. Tao, W. Li, J. Liang, G. Li, W. Yang, Alpha-ketoglutarate-activated NF-kappaB signaling promotes compensatory glucose uptake and brain Tumor Development. Mol. Cell 76(e147), 148–162 (2019). https://doi.org/10.1016/j.molcel.2019.07.007
O.D. Maddocks, C.R. Berkers, S.M. Mason, L. Zheng, K. Blyth, E. Gottlieb, K.H. Vousden, Serine starvation induces stress and p53-dependent metabolic remodelling in cancer cells. Nature 493, 542–546 (2013). https://doi.org/10.1038/nature11743
Y. Ou, S.J. Wang, L. Jiang, B. Zheng, W. Gu, p53 protein-mediated regulation of phosphoglycerate dehydrogenase (PHGDH) is crucial for the apoptotic response upon serine starvation. J. Biol. Chem. 290, 457–466 (2015). https://doi.org/10.1074/jbc.M114.616359
T. Terzian, Y.A. Suh, T. Iwakuma, S.M. Post, M. Neumann, G.A. Lang, C.S. Van Pelt, G. Lozano, The inherent instability of mutant p53 is alleviated by Mdm2 or p16INK4a loss. Genes Dev. 22, 1337–1344 (2008). https://doi.org/10.1101/gad.1662908
M. Halasi, M. Wang, T.S. Chavan, V. Gaponenko, N. Hay, A.L. Gartel, ROS inhibitor N-acetyl-L-cysteine antagonizes the activity of proteasome inhibitors. Biochem. J. 454, 201–208 (2013). https://doi.org/10.1042/BJ20130282
A.M. Florea, D. Busselberg, Cisplatin as an anti-tumor drug: cellular mechanisms of activity, drug resistance and induced side effects. Cancers 3, 1351–1371 (2011). https://doi.org/10.3390/cancers3011351
R. Marullo, E. Werner, N. Degtyareva, B. Moore, G. Altavilla, S.S. Ramalingam, P.W. Doetsch, Cisplatin induces a mitochondrial-ROS response that contributes to cytotoxicity depending on mitochondrial redox status and bioenergetic functions. PLoS ONE 8, e81162 (2013). https://doi.org/10.1371/journal.pone.0081162
A.M. Deaton, A. Bird, CpG islands and the regulation of transcription. Genes Dev. 25, 1010–1022 (2011). https://doi.org/10.1101/gad.2037511
L. Verdone, E. Agricola, M. Caserta, E. Di Mauro, Histone acetylation in gene regulation. Brief. Funct. Genomic. Proteomic 5, 209–221 (2006). https://doi.org/10.1093/bfgp/ell028
C.L. Brooks, W. Gu, How does SIRT1 affect metabolism, senescence and cancer? Nat. Rev. Cancer 9, 123–128 (2009). https://doi.org/10.1038/nrc2562
J.T. Zilfou, S.W. Lowe (2009) Tumor suppressive functions of p53. Cold Spring Harb. Perspect. Biol. 1, a001883. https://doi.org/10.1101/cshperspect.a001883
M. Zuker, Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003). https://doi.org/10.1093/nar/gkg595
Y. Han, B. Kim, U. Cho, I.S. Park, S.I. Kim, D.N. Dhanasekaran, B.K. Tsang, Y.S. Song, Mitochondrial fission causes cisplatin resistance under hypoxic conditions via ROS in ovarian cancer cells. Oncogene 38, 7089–7105 (2019). https://doi.org/10.1038/s41388-019-0949-5
D. Trachootham, W. Lu, M.A. Ogasawara, R.D. Nilsa, P. Huang, Redox regulation of cell survival. Antioxid. Redox. Signal 10, 1343–1374 (2008). https://doi.org/10.1089/ars.2007.1957
S.H. Moon, C.H. Huang, S.L. Houlihan, K. Regunath, W.A. Freed-Pastor, D.F. Morris JPt, Tschaharganeh, E.R. Kastenhuber, A.M. Barsotti, R. Culp-Hill, W. Xue, Y.J. Ho, T. Baslan, X. Li, A. Mayle, E. de Stanchina, L. Zender, D.R. Tong, A. D’Alessandro, S.W. Lowe, C. Prives, p53 represses the Mevalonate pathway to Mediate Tumor suppression. Cell 176(e519), 564–580 (2019). https://doi.org/10.1016/j.cell.2018.11.011
L. Li, Y. Mao, L. Zhao, L. Li, J. Wu, M. Zhao, W. Du, L. Yu, P. Jiang, p53 regulation of ammonia metabolism through urea cycle controls polyamine biosynthesis. Nature 567, 253–256 (2019). https://doi.org/10.1038/s41586-019-0996-7
L.M. Khachigian, The Yin and Yang of YY1 in tumor growth and suppression. Int. J. Cancer 143, 460–465 (2018). https://doi.org/10.1002/ijc.31255
Y. Shi, E. Seto, L.S. Chang, T. Shenk, Transcriptional repression by YY1, a human GLI-Kruppel-related protein, and relief of repression by adenovirus E1A protein. Cell 67, 377–388 (1991). https://doi.org/10.1016/0092-8674(91)90189-6
K.H. Lee, S. Evans, T.Y. Ruan, A.B. Lassar, SMAD-mediated modulation of YY1 activity regulates the BMP response and cardiac-specific expression of a GATA4/5/6-dependent chick Nkx2.5 enhancer. Development 131, 4709–4723 (2004). https://doi.org/10.1242/dev.01344
S. Wu, H. Wang, Y. Li, Y. Xie, C. Huang, H. Zhao, M. Miyagishi, V. Kasim, Transcription factor YY1 promotes cell proliferation by directly activating the pentose phosphate pathway. Cancer Res. 78, 4549–4562 (2018). https://doi.org/10.1158/0008-5472.CAN-17-4047
Acknowledgements
We thank Feng Zhang for supplying lenti-CRISPR v2 plasmid from addgene and the useful suggestion for plasmid extraction and lentivirus packaging.
Funding
This study was supported by funds from the National Natural Science Foundation of China (No. 82172920), National Natural Science Foundation for Young Scholars of China (No. 81902566) and Shanghai Jiaotong University Medical-Engineering Cross Research Fund (No. YG2019QNA53).
Author information
Authors and Affiliations
Contributions
Conception and design: J. Shao, L. Yao and X. Wang. Development of methodology: Z. Yuan, Yan. He and T. Shi. Acquisition of data: T. Shi and Z. Yuan. Analysis and interpretation of data: J. Shao, T. Shi, Z. Yuan, D. Zhang, X. Wang, and S. Chen. Writing, review, and/or revision of the manuscript: L. Yao, XJ. Wang and T. Shi. Administrative, technical, or material support: J. Shao, Z. Yuan, D. Zhang, and Y. He. Study supervision: X. Wang.
Corresponding authors
Ethics declarations
Ethical approval
All procedures related to patients were carried out in accordance with International Ethical Guidelines for Biomedical Research Involving Human Subjects (CIOMS). The study was approved by the Research Ethics Committee of Shanghai General Hospital, Shanghai Jiao Tong University. All the animal studies were approved by the Institutional Committee for Animal Care and Use of Laboratory Animals prepared by Shanghai Jiao Tong University.
Consent for publication
All authors had agreed to publish this manuscript.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 1.40 MB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Shi, T., Yuan, Z., He, Y. et al. Competition between p53 and YY1 determines PHGDH expression and malignancy in bladder cancer. Cell Oncol. 46, 1457–1472 (2023). https://doi.org/10.1007/s13402-023-00823-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-023-00823-8