Abstract
Purpose
Metastatic bladder cancer (BC) has the highest somatic mutation frequency and recurrence rate of all tumors. However, the cellular and molecular characteristics of BC remain unclear.
Methods
We performed single-cell RNA sequencing (scRNA-seq) on the samples of paracancerous normal tissue (PNT), primary tumor (PT) and lymph node metastasis (LNM). The proportions and gene expression profiles of different cell types in the tumor microenvironment (TME) were investigated.
Results
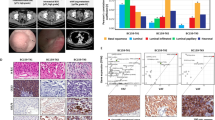
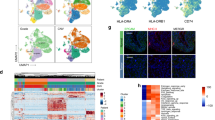
In total, 50,158 cells were classified into six populations. Malignant cells of PT and LNM exhibited large mutant DNA fragments, while the cell phenotypes and gene expression profiles differed during differentiation. Metastasis was associated with a poorer prognosis than PT. Tumor-associated stromal cells and inhibitory immune cells were the main cell populations in PT and LNM. Cell-cell communication analysis revealed the roles of signaling pathways of inflammatory cancer-associated fibroblast (iCAF) and tumor-associated macrophage (TAM) in exhaustion of T cells. In addition, iCAF may recruit TAM to promote formation of the TME earlier than the differentiation of tumor cells.
Conclusion
This study through scRNA-seq enhanced our understanding of new features about the cellular and molecular similarities and differences of high-grade and metastatic bladder cancer, which might provide potential therapeutic targets in future treatment.







Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article.
References
H. Sung, J. Ferlay, R.L. Siegel, M. Laversanne, I. Soerjomataram, A. Jemal, F. Bray, Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and Mortality Worldwide for 36 cancers in 185 countries. CA Cancer J Clin 71(3), 209–249 (2021)
Comprehensive molecular characterization, Of urothelial bladder carcinoma. Nature 507(7492), 315–322 (2014)
E.J. Pietzak, A. Bagrodia, E.K. Cha, E.N. Drill, G. Iyer, S. Isharwal, I. Ostrovnaya, P. Baez, Q. Li, M.F. Berger, A. Zehir, N. Schultz, J.E. Rosenberg, D.F. Bajorin, G. Dalbagni, H. Al-Ahmadie, D.B. Solit, B.H. Bochner, Next-generation sequencing of nonmuscle invasive bladder Cancer reveals potential biomarkers and rational therapeutic targets. Eur. Urol. 72(6), 952–959 (2017)
L.B. Alexandrov, S. Nik-Zainal, D.C. Wedge, S.A. Aparicio, S. Behjati, A.V. Biankin, G.R. Bignell, N. Bolli, A. Borg, A.L. Børresen-Dale, S. Boyault, B. Burkhardt, A.P. Butler, C. Caldas, H.R. Davies, C. Desmedt, R. Eils, J.E. Eyfjörd, J.A. Foekens, M. Greaves, F. Hosoda, B. Hutter, T. Ilicic, S. Imbeaud, M. Imielinski, N. Jäger, D.T. Jones, D. Jones, S. Knappskog, M. Kool, S.R. Lakhani, C. López-Otín, S. Martin, N.C. Munshi, H. Nakamura, P.A. Northcott, M. Pajic, E. Papaemmanuil, A. Paradiso, J.V. Pearson, X.S. Puente, K. Raine, M. Ramakrishna, A.L. Richardson, J. Richter, P. Rosenstiel, M. Schlesner, T.N. Schumacher, P.N. Span, J.W. Teague, Y. Totoki, A.N. Tutt, R. Valdés-Mas, M.M. van Buuren, L. van ‘t Veer, A. Vincent-Salomon, N. Waddell, L.R. Yates, J. Zucman-Rossi, P.A. Futreal, U. McDermott, P. Lichter, M. Meyerson, S.M. Grimmond, R. Siebert, E. Campo, T. Shibata, S.M. Pfister, P.J. Campbell, M.R. Stratton, Signatures of mutational processes in human cancer, Nature 500(7463), 415–421 (2013)
L.M. Merlo, J.W. Pepper, B.J. Reid, C.C. Maley, Cancer as an evolutionary and ecological process. Nat. Rev. Cancer 6(12), 924–935 (2006)
T. Okamoto, D. duVerle, K. Yaginuma, Y. Natsume, H. Yamanaka, D. Kusama, M. Fukuda, M. Yamamoto, F. Perraudeau, U. Srivastava, Y. Kashima, A. Suzuki, Y. Kuze, Y. Takahashi, M. Ueno, Y. Sakai, T. Noda, K. Tsuda, Y. Suzuki, S. Nagayama, R. Yao, Comparative analysis of patient-matched PDOs revealed a reduction in OLFM4-Associated clusters in metastatic lesions in Colorectal Cancer. Stem Cell. Reports 16(4), 954–967 (2021)
J.A. Witjes, H.M. Bruins, R. Cathomas, E.M. Compérat, N.C. Cowan, G. Gakis, V. Hernández, E. Linares Espinós, A. Lorch, Y. Neuzillet, M. Rouanne, G.N. Thalmann, E. Veskimäe, M.J. Ribal, A.G. van der Heijden, European Association of Urology Guidelines on muscle-invasive and metastatic bladder Cancer: Summary of the 2020 guidelines. Eur. Urol. 79(1), 82–104 (2021)
V.G. Patel, W.K. Oh, M.D. Galsky, Treatment of muscle-invasive and advanced bladder cancer in 2020. CA Cancer J Clin 70(5), 404–423 (2020)
Y. Zheng, X. Yang, Spatial RNA sequencing methods show high resolution of single cell in cancer metastasis and the formation of tumor microenvironment, Biosci Rep 43(2) (2023)
J.J. Quinn, M.G. Jones, R.A. Okimoto, S. Nanjo, M.M. Chan, N. Yosef, T.G. Bivona, J.S. Weissman, Single-cell lineages reveal the rates, routes, and drivers of metastasis in cancer xenografts, Science 371(6532) (2021)
Y. Chi, J. Remsik, V. Kiseliovas, C. Derderian, U. Sener, M. Alghader, F. Saadeh, K. Nikishina, T. Bale, C. Iacobuzio-Donahue, T. Thomas, D. Pe’er, L. Mazutis, A. Boire, Cancer cells deploy lipocalin-2 to collect limiting iron in leptomeningeal metastasis. Science 369(6501), 276–282 (2020)
A. Dobin, C.A. Davis, F. Schlesinger, J. Drenkow, C. Zaleski, S. Jha, P. Batut, M. Chaisson, T.R. Gingeras, STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29(1), 15–21 (2013)
A. Butler, P. Hoffman, P. Smibert, E. Papalexi, R. Satija, Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36(5), 411–420 (2018)
C.S. McGinnis, L.M. Murrow, Z.J. Gartner, DoubletFinder: Doublet Detection in single-cell RNA sequencing data using Artificial Nearest neighbors. Cell. Syst. 8(4), 329–337.e4 (2019)
E.Z. Macosko, A. Basu, R. Satija, J. Nemesh, K. Shekhar, M. Goldman, I. Tirosh, A.R. Bialas, N. Kamitaki, E.M. Martersteck, J.J. Trombetta, D.A. Weitz, J.R. Sanes, A.K. Shalek, A. Regev, S.A. McCarroll, Highly parallel genome-wide expression profiling of individual cells using Nanoliter Droplets. Cell 161(5), 1202–1214 (2015)
D. Aran, A.P. Looney, L. Liu, E. Wu, V. Fong, A. Hsu, S. Chak, R.P. Naikawadi, P.J. Wolters, A.R. Abate, A.J. Butte, M. Bhattacharya, Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20(2), 163–172 (2019)
S.V. Puram, I. Tirosh, A.S. Parikh, A.P. Patel, K. Yizhak, S. Gillespie, C. Rodman, C.L. Luo, E.A. Mroz, K.S. Emerick, D.G. Deschler, M.A. Varvares, R. Mylvaganam, O. Rozenblatt-Rosen, J.W. Rocco, W.C. Faquin, D.T. Lin, A. Regev, B.E. Bernstein, single-cell transcriptomic analysis of primary and metastatic Tumor Ecosystems in Head and Neck Cancer. Cell 171(7), 1611–1624.e24 (2017)
C. Trapnell, D. Cacchiarelli, J. Grimsby, P. Pokharel, S. Li, M. Morse, N.J. Lennon, K.J. Livak, T.S. Mikkelsen, J.L. Rinn, The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32(4), 381–386 (2014)
S. Hänzelmann, R. Castelo, J. Guinney, GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform 14, 7 (2013)
S. Jin, C.F. Guerrero-Juarez, L. Zhang, I. Chang, R. Ramos, C.H. Kuan, P. Myung, M.V. Plikus, Q. Nie, Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12(1), 1088 (2021)
P.M. Moonen, G.F. Merkx, P. Peelen, H.F. Karthaus, D.F. Smeets, J.A. Witjes, UroVysion compared with cytology and quantitative cytology in the surveillance of non-muscle-invasive bladder cancer. Eur. Urol. 51(5), 1275–1280 (2007); discussion 1280
X. Diao, J. Cai, J. Zheng, J. Kong, S. Wu, H. Yu, H. Huang, W. Xie, X. Chen, C. Huang, L. Huang, H. Qin, J. Huang, T. Lin, Association of chromosome 7 aneuploidy measured by fluorescence in situ hybridization assay with muscular invasion in bladder cancer. Cancer Commun. (Lond) 40(4), 167–180 (2020)
J. Bellmunt, Stem-Like Signature Predicting Disease Progression in Early Stage Bladder Cancer. The Role of E2F3 and SOX4, Biomedicines 6(3) (2018)
M. Riester, L. Werner, J. Bellmunt, S. Selvarajah, E.A. Guancial, B.A. Weir, E.C. Stack, R.S. Park, R. O’Brien, F.A. Schutz, T.K. Choueiri, S. Signoretti, J. Lloreta, L. Marchionni, E. Gallardo, F. Rojo, D.I. Garcia, Y. Chekaluk, D.J. Kwiatkowski, B.H. Bochner, W.C. Hahn, A.H. Ligon, J.A. Barletta, M. Loda, D.M. Berman, P.W. Kantoff, F. Michor, J.E. Rosenberg, Integrative analysis of 1q23.3 copy-number gain in metastatic urothelial carcinoma. Clin. Cancer Res. 20(7), 1873–1883 (2014)
A. Veerakumarasivam, H.E. Scott, S.F. Chin, A. Warren, M.J. Wallard, D. Grimmer, K. Ichimura, C. Caldas, V.P. Collins, D.E. Neal, J.D. Kelly, High-resolution array-based comparative genomic hybridization of bladder cancers identifies mouse double minute 4 (MDM4) as an amplification target exclusive of MDM2 and TP53. Clin. Cancer Res. 14(9), 2527–2534 (2008)
X. Li, M. Xu, L. Ding, J. Tang, MiR-27a: a novel biomarker and potential therapeutic target in tumors. J. Cancer 10(12), 2836–2848 (2019)
L. Haghverdi, M. Büttner, F.A. Wolf, F. Buettner, F.J. Theis, Diffusion pseudotime robustly reconstructs lineage branching. Nat. Methods 13(10), 845–848 (2016)
E. Sahai, I. Astsaturov, E. Cukierman, D.G. DeNardo, M. Egeblad, R.M. Evans, D. Fearon, F.R. Greten, S.R. Hingorani, T. Hunter, R.O. Hynes, R.K. Jain, T. Janowitz, C. Jorgensen, A.C. Kimmelman, M.G. Kolonin, R.G. Maki, R.S. Powers, E. Puré, D.C. Ramirez, R. Scherz-Shouval, M.H. Sherman, S. Stewart, T.D. Tlsty, D.A. Tuveson, F.M. Watt, V. Weaver, A.T. Weeraratna, Z. Werb, A framework for advancing our understanding of cancer-associated fibroblasts. Nat. Rev. Cancer 20(3), 174–186 (2020)
T. Zhan, N. Rindtorff, M. Boutros, Wnt signaling in cancer. Oncogene 36(11), 1461–1473 (2017)
K.D. Howarth, T. Mirza, S.L. Cooke, S.F. Chin, J.C. Pole, E. Turro, M.D. Eldridge, R.M. Garcia, O.M. Rueda, C. Boursnell, J.E. Abraham, C. Caldas, P.A.W. Edwards, NRG1 fusions in breast cancer. Breast Cancer Res 23(1), 3 (2021)
D.H. Shin, J.Y. Jo, J.Y. Han, Dual targeting of ERBB2/ERBB3 for the treatment of SLC3A2-NRG1-Mediated Lung Cancer. Mol. Cancer Ther. 17(9), 2024–2033 (2018)
C.D. Mills, K. Kincaid, J.M. Alt, M.J. Heilman, A.M. Hill, M-1/M-2 macrophages and the Th1/Th2 paradigm. J. Immunol. 164(12), 6166–6173 (2000)
Y. Pan, Y. Yu, X. Wang, T. Zhang, Tumor-Associated Macrophages in Tumor Immunity. Front. Immunol. 11, 583084 (2020)
C. Yunna, H. Mengru, W. Lei, C. Weidong, Macrophage M1/M2 polarization. Eur. J. Pharmacol. 877, 173090 (2020)
N. Zhang, S.H. Kim, A. Gainullina, E.C. Erlich, E.J. Onufer, J. Kim, R.S. Czepielewski, B.A. Helmink, J.R. Dominguez, B.T. Saunders, J. Ding, J.W. Williams, J.X. Jiang, B.H. Segal, B.H. Zinselmeyer, G.J. Randolph, K.W. Kim, LYVE1 + macrophages of murine peritoneal mesothelium promote omentum-independent ovarian tumor growth, J Exp Med 218(12) (2021)
M.B. Buechler, W. Fu, S.J. Turley, Fibroblast-macrophage reciprocal interactions in health, fibrosis, and cancer. Immunity 54(5), 903–915 (2021)
J.E. Cheong, L. Sun, Targeting the IDO1/TDO2-KYN-AhR pathway for Cancer Immunotherapy - Challenges and Opportunities. Trends Pharmacol. Sci. 39(3), 307–325 (2018)
L.M. McLane, M.S. Abdel-Hakeem, E.J. Wherry, CD8 T cell exhaustion during chronic viral infection and Cancer. Annu. Rev. Immunol. 37, 457–495 (2019)
D. Masopust, V. Vezys, A.L. Marzo, L. Lefrançois, Preferential localization of effector memory cells in nonlymphoid tissue. Science 291(5512), 2413–2417 (2001)
S. Olalekan, B. Xie, R. Back, H. Eckart, A. Basu, Characterizing the tumor microenvironment of metastatic ovarian cancer by single-cell transcriptomics. Cell. Rep. 35(8), 109165 (2021)
S.K. Daniel, Y.D. Seo, V.G. Pillarisetty, The CXCL12-CXCR4/CXCR7 axis as a mechanism of immune resistance in gastrointestinal malignancies. Semin Cancer Biol 65, 176–188 (2020)
X. Zhang, Y. Dong, M. Zhao, L. Ding, X. Yang, Y. Jing, Y. Song, S. Chen, Q. Hu, Y. Ni, ITGB2-mediated metabolic switch in CAFs promotes OSCC proliferation by oxidation of NADH in mitochondrial oxidative phosphorylation system. Theranostics 10(26), 12044–12059 (2020)
M. Marchesi, E. Andersson, L. Villabona, B. Seliger, A. Lundqvist, R. Kiessling, G.V. Masucci, HLA-dependent tumour development: a role for tumour associate macrophages? J. Transl Med. 11, 247 (2013)
Acknowledgements
We acknowledge the OE Biotech Co. Ltd (Shanghai, China) for providing single-cell RNA sequencing technology. And the support Funding: including the Central Guidance on Local Science and Technology Development Fund of Shanxi Province (No. YDZJSX2021C010); Nature Science Foundation of Shanxi Province (No.202103021224412); Fund Program for the Scientific Activities of Selected Returned Overseas Professionals in Shanxi Province (No.20210005); Shanxi Provincial Basic Applied Research Project (No.20210302124611).
Funding
Nature Science Foundation of Shanxi Province (No.202,103,021,224,412). The Central Guidance on Local Science and Technology Development Fund of Shanxi Province (No.YDZJSX2021C010). Fund Program for the Scientific Activities of Selected Returned Overseas Professionals in Shanxi Province (No.20,210,005). Shanxi Provincial Basic Applied Research Project (No.20,210,302,124,611).
Author information
Authors and Affiliations
Contributions
Material preparation, data collection and analysis were performed by Yue Zheng. The first draft of the manuscript was written by Yue Zheng and Xin Wang. Yue Zheng was the major contributor in designing and writing this manuscript. Xin Wang was responsible for writing the experimental part. Funding acquisition, reviewing and editing were performed by Xiaofeng Yang. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Ethics approval and consent to participate
The research involving human samples have been performed in accordance with the Declaration of Helsinki. This study was approved by the Ethics Committee of the First Hospital of Shanxi Medical University (K093). Informed consent was obtained from the patients.
Consent for publication
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.






Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zheng, Y., Wang, X., Yang, X. et al. Single-cell RNA sequencing reveals the cellular and molecular characteristics of high-grade and metastatic bladder cancer. Cell Oncol. 46, 1415–1427 (2023). https://doi.org/10.1007/s13402-023-00820-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13402-023-00820-x