Abstract
Cleavable cross-linkers are gaining increasing importance for chemical cross-linking/mass spectrometry (MS) as they permit a reliable and automated data analysis in structural studies of proteins and protein assemblies. Here, we introduce 1,3-diallylurea (DAU) as the first CID-MS/MS-cleavable, photo-thiol-reactive cross-linker. DAU is a commercially available, inexpensive reagent that efficiently undergoes an anti-Markovnikov hydrothiolation with cysteine residues in the presence of a radical initiator upon UV-A irradiation. Radical cysteine cross-linking proceeds via an orthogonal “click reaction” and yields stable alkyl sulfide products. DAU reacts at physiological pH and cross-linking reactions with peptides, and proteins can be performed at temperatures as low as 4 °C. The central urea bond is efficiently cleaved upon collisional activation during tandem MS experiments generating characteristic product ions. This improves the reliability of automated cross-link identification. Different radical initiators have been screened for the cross-linking reaction of DAU using the thiol-containing compounds cysteine and glutathione. Our concept has also been exemplified for the biologically relevant proteins bMunc13-2 and retinal guanylyl cyclase-activating protein-2.

ᅟ
Similar content being viewed by others
Introduction
Chemical cross-linking in combination with mass spectrometry (MS) has evolved as a powerful method to elucidate the three-dimensional structures of proteins and protein complexes [1,2,3,4]. Protein cross-linkers with two reactive groups, separated by a spacer with a defined length, can connect functional groups of amino acid side chains. As such, the cross-linkers serve as “molecular rulers” that impose geometrical constraints and serve as basis for a subsequent computational modeling of 3D structures of proteins and their assemblies [5,6,7,8,9,10,11]. A wide variety of reagents and application protocols has been developed for targeting different amino acid residues in proteins in vitro as well as in vivo [12,13,14,15]. The length of the cross-linker can be adapted to a specific protein system. Diazirine-containing amino acids, such as 2-amino-5,5-azihexanoic acid (“photo-methionine”) [16, 17] and 2-amino-3-(3-(2-aminoethyl)-2H-diazirin-3-yl)propanoic acid (“photo-lysine”) [18] can be incorporated into proteins. Moreover, various genetically encoded, unnatural amino acids can be site-specifically incorporated into proteins [19,20,21,22]. The advantage of modifying proteins with cross-linkable, unnatural amino acids consists in the chance to conduct the photo-cross-linking reaction after UV-irradiation directly in the cell. An additional strength of the diazirine-containing amino acid photo-methionine is its short distance. Another class of cross-linkers that are able to connect amino acid residues in close proximity to each other are so-called zero-length cross-linkers, such as carbodiimides [23], 4-(4,6-dimethoxy-1,3,5-triazin-2-yl)-4-methylmorpholinium chloride (DMTMM) [24, 25], and 1,1′-carbonyldiimidazole (CDI) [26]. Short-distance constraints provide highly valuable geometrical information as input for computational modeling studies of the protein(s) under investigation [27]. On the other hand, longer cross-linkers connecting amino acids that are spatially apart from each other have proven useful for mapping large protein assemblies [28,29,30].
The vast majority of cross-linking/MS studies are conducted in a “bottom-up” approach, involving the enzymatic digestion of the connected protein(s), followed by LC/MS/MS analysis of the proteolytic peptides. These proteolytic peptide mixtures have a higher degree of complexity compared to those in conventional proteomics experiments, which is attributed to the presence of various types of cross-linked peptides [31]. LC/MS/MS analysis alone may not be sufficient for an unambiguous identification of cross-linked peptides as a complete sequencing of both cross-linked peptides cannot be achieved in all cases [32,33,34]. Therefore, different strategies have been developed to improve the unambiguous identification of cross-linked products [35,36,37,38,39].
Currently, one of the most promising cross-linking approaches seems to rely on cross-linkers that are cleaved under tandem MS conditions [26, 40,41,42,43,44,45,46]. The gas phase chemistry of such reagents can be fine-tuned by introducing specific, CID-MS/MS-labile moieties in the spacer of the cross-linker, such as urea [26, 44], sulfoxide [43], aspartic acid-proline [40, 45, 46], and azo [47] groups. As such, these cross-linkers generate characteristic, diagnostic product ions during CID-MS/MS analysis. The presence of these diagnostic product ions reduces the search space of cross-links from n2 to 2n [42]. CID-MS/MS-cleavable cross-linkers will therefore drastically reduce the number of false-positive cross-link identifications compared to non-cleavable reagents. We have recently shown that diazirine-based photo-cross-linkers also generate CID-MS/MS-cleavable cross-links [48]. This will pave the way towards a fully automated data analysis of cross-linked products, which allows conducting proteome-wide cross-linking studies [42, 49,50,51].
The sulfhydryl group of cysteine residues is an important target for protein conjugation, labeling, and cross-linking [52]. One major aspect of targeting cysteine side chains is that the respective cysteines should not be involved in disulfide bond formation as the reduction of disulfides might distort the protein’s 3D structure. Thiol groups are among the most reactive nucleophilic groups in biological macromolecules and can be targeted more selectively than primary amines [53]. In case a protein contains only a few or no cysteine residues at all, the number of sulfhydryl group might be increased by modifying lysine residues with, e.g., 2-iminothiolane (Traut’s reagent) [54].
Thiol-reactive groups can undergo nucleophilic substitution reactions under aqueous conditions. These include haloacetyls, alkyl halides, and aziridines as well as Michael acceptors, such as maleimides, acryloyls, and vinylsulfones [52]. Moreover, pyridyl disulfides, 5-thio-2-nitrobenzoic acid (TNB), and disulfide-reducing agents can be employed for a reversible functionalization of sulfhydryl groups [52].
Bioconjugation strategies based on the ability of thiols to participate in radical reactions have been mainly applied in the preparation of biomaterials [55,56,57]. These reactions can be directly initiated by UV light or by combining a radical initiator with heat or UV irradiation, and they proceed via an orthogonal and “click fashion” [58]. In particular, the radical-based thiol-ene “click reaction” is widely employed in polymer and material sciences but has also been successfully applied to bioconjugation [52, 59]. Under radical conditions, the sulfhydryl group reacts regioselectively with alkenes to form an anti-Markovnikov addition product (alkyl sulfide or thioether). The regioselectivity is more pronounced for non-activated alkenes that do not undergo a competing Michael addition.
The general mechanism (Scheme 1) involves the formation of a thiyl radical that adds to a C=C bond to generate a carbon-centered radical [58]. The latter is responsible for the radical transfer step by abstracting a hydrogen atom from another sulfhydryl group, thus regenerating the thiyl radical.
Inspired by the potential of radical-based thiol-ene photo click reaction, we investigated 1,3-diallylurea (DAU) as photochemical and MS-cleavable cross-linker for a covalent connection of cysteine residues in proteins. DAU is commercially available and inexpensive (~ 20 €/g). Alternatively, we present a one-step synthesis yielding DAU without additional purification steps (details of DAU synthesis and characterization are presented in the Supporting Information). The anti-Markovnikov regioselectivity of the thiol-ene reaction dictates the formation of two alkyl sulfide bonds at the sp2-hybridized carbons of DAU. Thus, the spacer length of DAU is estimated between 9 and 10 Å. The spacer of DAU contains a urea group enabling the cleavability of cross-linked products upon collisional activation [44]. As such, we expected DAU cross-links to proceed along the well-studied fragmentation pathway of the urea-based DSBU [41, 44] and CDI [26] cross-linkers. As for the latter two CID-MS/MS-cleavable cross-linkers, the two characteristic 26-u doublets should be visible in the fragment ion mass spectra of DAU cross-linked products. This allows distinguishing between different cross-link types, namely intra- and interpeptide cross-links and “dead-end” products [31]. In the case of DAU, the diagnostic fragment ions of the cross-linker possess a mass modification of 57 u (amine) and 83 u (isocyanate), respectively (Scheme 2).
In this work, we first tested three radical initiators for DAU-based cross-linking using the thiol-containing compounds cysteine and glutathione. We then applied our concept to two protein systems: (i) bMunc13-2 that plays an important role in synaptic vesicle priming [60] and (ii) retinal guanylyl cyclase-activating protein-2 that is relevant for photo-signal transduction [61].
Experimental
Chemicals and Peptides
All chemicals and solvents were used without further purification (Acros Organics, Aldrich, Fluka, Merck).
Cross-linker Synthesis
One-step synthesis of DAU is described in the Supplementary Material.
Protein Expression and Purification
Rat bMunc13-2 domain, comprising amino acids 367–780, was expressed in Escherichia coli cells (BL21(DE3)c+RIL) using a pGEX-4T-1 plasmid. The glutathione-S-transferase (GST)-tagged bMunc13-2(aa 367–780) domain was purified from E. coli cell lysates by GST affinity chromatography (2 × 1 ml GSTrapFF column) and strong anion exchange chromatography (1 ml HiTrap Q HP column). Cleavage of the GST tag was performed with thrombin (0.08 mg/ml protein solution) overnight at 6 °C. The protein was stored at − 20 °C in 20 mM HEPES, 300 mM NaCl, 0.5 mM TCEP, 10% glycerol (pH 7.2). Bovine GCAP-2 was expressed in E. coli with N-terminal myristoylation and purified as described previously [62].
Cross-linking Experiments
-
1.
Photo-Cross-linking of Cysteine and GSH. One microliter of a DMSO solution containing DAU (25 mM) and a photo-radical-initiator (2.5 mM), namely benzophenone (BP), 2,2-dimethoxy-2-phenylacetophenone (DMPA), or 2,2′-azobis(2-methylpropionamidine) dihydrochloride (AAPH), was added to 49 μL of an aqueous solution containing cysteine or GSH (1.1 mM) and HEPES (20 mM, pH 7.5). The resulting reaction mixture was kept on ice, placed in a home-built UV-irradiation device [63], and exposed to UV-A irradiation (8000 mJ/cm2; corresponding to an irradiation time of ca. 20 min). The photo-reaction was quenched by adding dithiothreitol (DTT) to a final concentration of 5 mM. The solution was desalted using C18-ZipTips (Millipore, Billerica, MA, USA) and analyzed by offline nano-ESI-MS/MS.
-
2.
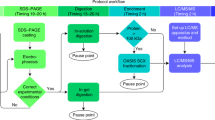
Photo-cross-linking of bMunc13-2 and GCAP-2. The protein stock solution was diluted to give a final concentration of 10 μM protein (bMunc13-2 in 20 mM HEPES, 0.5 mM TCEP, 7.5% (v/v) glycerol, 1 mM CaCl2, 230 mM NaCl, pH 7.2; GCAP-2 in 20 mM HEPES, 1 mM CaCl2, pH 7.5). A freshly prepared stock solution of DAU and BP in DMSO was added to the protein solution at 4 °C to give final concentrations of 1 mM DAU and 200 μM BP. The photo-reaction and quenching were conducted in the same fashion as described for cysteine and GSH. Afterwards, the protein solution was subjected to reduction and alkylation. In-solution digestion was performed according to an existing protocol [64, 65] using a trypsin/GluC mixture. The resulting peptide mixture was directly analyzed by HPLC/nano-ESI-MS/MS.
Offline Nano-ESI-MS/MS
Samples (4 μl) were filled into fire-polished borosilicate glass capillaries with filament (4″, 1.2 mm/0.68 mm OD/ID, World Precision Instruments) that were in-house pulled (Model P-1000, Flaming/Brown Micropipette Puller, Sutter Instruments) and gold-coated. MS and MS/MS analyses were performed on an Orbitrap Fusion Tribrid mass spectrometer (Thermo Fisher Scientific) equipped with a Nanospray Flex ion source (Thermo Fisher Scientific). High-resolution full scans (R = 120,000 at m/z 200) were recorded in the m/z range 50–1000. Fragmentation was performed by higher-energy collision-induced dissociation (HCD) at 20% normalized collision energy (NCE) using an isolation window of 2 Th. High-resolution HCD product ion scans were conducted at R = 60,000 at m/z 200 with the maximum accumulation time set to 200 ms.
Nano-HPLC/Nano-ESI-Orbitrap-MS/MS
Peptide mixtures were analyzed by LC/MS/MS on an UltiMate 3000 RSLC nano-HPLC system (Thermo Fisher Scientific) coupled to an Orbitrap Fusion Tribrid (Thermo Fisher Scientific) (for bMunc13-2) or an Orbitrap Q-Exactive Plus mass spectrometer (Thermo Fisher Scientific) (for GCAP-2), both equipped with Nanospray Flex ion source (Thermo Fisher Scientific). Fragmentation was performed by HCD (30 ± 3% NCE); data were acquired in data-dependent MS/MS mode. Each high-resolution full scan (R = 120,000 (Fusion) or R = 140,000 (Q-Exactive) at m/z 200) in the Orbitrap was followed by high-resolution HCD product ion scans (R = 15,000 (Fusion) or R = 17,500 (Q-Exactive) at m/z 200) within 5 s, starting with the most intense signal in the mass spectrum (isolation window of 2 Th). The target value was set to 50,000 (Fusion) or 200,000 (Q-Exactive) with maximum accumulation times of 200 ms (Fusion) or 250 ms (Q-Exactive). Dynamic exclusion (exclusion duration 60s) was enabled.
Identification of Cross-Linked Products
Cross-linked products of bMunc13-2 and GCAP-2 were automatically annotated with MeroX [65] and manually validated. Mass deviations of 3 and 10 ppm were applied for precursor and product ions. A 1% FDR cut-off and a signal-to-noise ratio of ≥ 2 were applied. Cysteines were considered as potential cross-linking sites for DAU. Oxidation of Met and carbamidomethylation of cysteines were set as variable modifications. Three missed cleavage sites were considered for each amino acid [Lys and Arg (for trypsin), Glu and Asp (for GluC)]. The representation of cross-links in bMunc13-2 was performed with the Circos software [66]. The amino acid sequences of bMunc13-2 and GCAP-2 as well as mgf files and MeroX settings files are provided in the Supplementary Material.
Results and Discussion
Cysteine and GSH Cross-linking
To optimize the thiol-ene click reaction conditions, DAU was initially evaluated for the amino acid cysteine and GSH as simple, thiol-containing compounds. GSH is a tripeptide (γ-Glu-Cys-Gly) with a γ-peptide linkage between the carboxyl group of the glutamate side chain and the α-amino group of cysteine. Photo-cross-linking reactions were performed in 20 mM HEPES buffer at pH 7.5 on ice. First, we screened three radical photo-initiators at UV-A irradiation, namely benzophenone (BP), 2,2-dimethoxy-2-phenylacetophenone (DMPA), and 2,2′-azobis(2-methylpropionamidine) dihydrochloride (AAPH), using a 11:5:0.5 M ratio of thiol compound/cross-linker/radical initiator. The three radical initiators were selected to compare the reaction efficiencies of unimolecular (type I) and bimolecular (type II) radical photo-initiators with different water solubility (Scheme 3).
First, the efficiency of the thiol-ene “click photo-reaction” was qualitatively tested on cysteine using BP as radical photo-initiator. ESI-MS and ESI-HCD-MS/MS data recorded in positive ionization mode from the reaction mixtures revealed the formation of the cysteine cross-linked product (Cys-DAU-Cys; Fig. 1). HCD-MS/MS experiments were conducted for the protonated, cross-linked cysteine species ([M + H]+ at m/z 383) to confirm the cleavability of the central urea group upon collisional activation. Indeed, the singly protonated ion of cross-linked cysteine undergoes the same unimolecular dissociation as observed for the established cross-linkers DSBU [41, 44] and CDI [26], with the relevant amine and isocyanate product ions exhibiting the diagnostic mass difference of 26 u (Fig. 1).
It should be noted that DAU cross-linked products with cysteine were not identified in cross-linking experiments without UV-A irradiation. This confirmed the absence of a side reaction between the thiol of cysteine residues and DAU due to a putative Michael addition. This illustrates that our DAU cross-linking protocol selectively generates anti-Markovnikov hydrothiolation products (see Schemes 1 and 2), defining the spacer length of DAU to be in the range between 9 and 10 Å.
As a next step, we compared the performance of the three radical photo-initiators for peptide cross-linking using GSH as model compound (Fig. 2). The cross-linked GSH product (GSH-DAU-GSH) is visible in the mass spectra as singly protonated ([M + H]+ at m/z 755) and doubly protonated ([M + 2H]2+ at m/z 378) species (Fig. 2). The relative signal intensities of the cross-linked species were found to be increased when the radical initiators BP or DMPA were employed. Reaction of DAU with a single GSH molecule ([M + H]+ at m/z 448) resulted in the formation of the thioether product when using AAPH as radical initiator (Fig. 2c). Also, a signal of unreacted DAU ([M + H]+ at m/z 141) was visible for AAPH. These results indicate that BP and DMPA are more effective than AAPH in initiating the photo-cross-linking reaction, resulting in an almost quantitative conversion of DAU. Therefore, we selected BP as radical photo-initiator for further testing DAU cross-linking on the two proteins bMunc13-2 and the retinal guanylate cyclase-activating protein GCAP-2.
(+)-ESI mass spectra of GSH cross-linking reaction mixtures with DAU using the radical initiators (a) BP, (b) DMPA, and (c) AAPH recorded in positive ion mode. The cross-linked product (GSH-DAU-GSH) appears as singly protonated ([M + H]+ at m/z 755) and doubly protonated ([M + 2H]2+ at m/z 378) species. The cross-linked product (shown in green) was found to be more abundant for BP or DMPA than for AAPH. Unreacted DAU ([M + H]+ at m/z 141; shown in red) and partially reacted DAU with one GSH molecule (GSH-DAU) ([M + H]+ at m/z 448; shown in blue) were still visible when AAPH was used. Unreacted GSH ([M + H]+ at m/z 308) is shown in gray
bMunc13-2 and GCAP-2 Protein Cross-Linking
To evaluate DAU for its efficiency on cross-linking cysteine-containing proteins, we applied our photo-cross-linking approach to two cytosolic proteins that are currently under investigation in our lab: (i) a bMunc13-2 domain, comprising amino acids 367–780 and (ii) GCAP-2. The bMunc13-2 domain is a ~ 46 kDa protein containing 11 free thiol groups in cysteine side chains, while GCAP-2 possesses a molecular weight of ~ 24 kDa and contains three free cysteines.
The two proteins were subjected to photo-cross-linking in the presence of DAU as cross-linker and BP as radical photo-initiator. After quenching, the cross-linked proteins were subjected to enzymatic proteolysis and LC/MS/MS analysis. Combined fragment ion mass spectra were annotated with the MeroX software [65], and all cross-links were manually validated. Exemplary product ion mass spectra assigned by MeroX are presented in Fig. 3. The diagnostic amine and isocyanate product ions of DAU cross-linked products are visible at the MS/MS level, together with b-, y-, and a-type ions of the connected peptides. The thorough sequencing of DAU cross-links already at the MS/MS level suggests that MS3 experiments might not be necessary for an unambiguous identification of cross-links, which is advantageous as it reduces the duty cycle.
Fragment ion mass spectra (HCD-MS/MS) of three bMunc13-2 cross-linked products with DAU, automatically annotated by MeroX [65]. (a) [M + 3H]3+ at m/z 803.083, Cys-92 cross-linked to Cys-101. (b) [M + 3H]3+ at m/z 963.809, [M + H]+, Cys-84 cross-linked to Cys-44. (c) [M + 3H]3+ at m/z 715.703, Cys-84 cross-linked to Cys-92
We identified 29 unique cross-links for bMunc13-2 (Fig. 4) and one cross-link for GCAP-2.
Circular plot representing the unique DAU cross-links identified for bMunc13-2. The plot was prepared with the Circos software [66]
Two thirds of bMunc13-2 cross-links were identified in both replicates, while the GCAP-2 cross-link was found in both replicates (Fig. 5). The distance of the intramolecular GCAP-2 cross-link was determined to be 25.2 Å (pdb entry 1JBA), while for bMunc13-2, there is no high-resolution structure available to map the cross-links. The complete list of unique cross-links is reported in the Supplementary Material, together with the mgf and MeroX settings files. The number of unique cross-links identified in bMunc13-2 is remarkable, testifying the high yield and the specificity of thiol-ene “click” cross-linking.
To date, the most commonly used principles for protein cross-linking are N-hydroxysuccinimide (NHS) esters [4]. One drawback of NHS esters is that they mainly target lysine residues, which might not be optimally suited for protein cross-linking for the following reasons: First, the lysine side chain is highly flexible and distance information obtained from cross-linking of lysines is not very stringent. Second, the positive charge of the ε-amine group in lysines is absent after modification by the cross-linking reagent. Another important aspect for data analysis is that cross-links composed of consecutive amino acid sequences are isobaric to peptides that are modified with a partially hydrolyzed cross-linker (type 0 or “dead-end” cross-links) involving identical amino acid sequences [31]. This complicates data analysis for non-MS-cleavable reagents and is currently the most frequent source of a misassignment of cross-links [34]. Notably, DAU cross-links are not affected by this issue as “dead-end” cross-links are in this case not isobaric to interpeptide cross-links. In fact, the thiol-ene photochemistry of DAU presents an addition reaction rather than a nucleophilic acyl substitution in the case of NHS esters. This greatly simplifies the correct assignment of DAU cross-linked products and eliminates the need to obtain complete fragmentation patterns of cross-linked peptides.
Conclusions
We introduce DAU as the first photo-thiol-reactive and CID-MS/MS-cleavable cross-linker to target cysteine residues in proteins. It relies on orthogonal thiol-ene “click chemistry,” is thiol-selective, and highly efficient. The reaction is induced by UV-A irradiation under physiological conditions. The cross-linking reaction of DAU was optimized for cysteine and GSH using three different radical initiators, yielding optimum cross-linking efficiencies for BP. The urea moiety of DAU ensures CID-MS/MS cleavability to facilitate an unambiguous cross-link assignment and allows conducting peptide sequencing at the MS/MS level. The applicability of DAU to thiol-containing proteins was demonstrated for bMunc13-2 and GCAP-2. We envision a broad applicability of DAU in structural proteomics studies as it specifically targets free cysteines, giving a complementary reactivity to the commonly used amine-reactive NHS esters. On top of that, the assignment of DAU cross-links is not hampered by the presence of isobaric species, i.e., dead-end cross-links versus interpeptide cross-links that are composed of consecutive sequences, as is the case for non-cleavable NHS ester cross-linkers.
Change history
11 February 2019
In this issue, the citation information on the opening page of each article PDF is incorrect. It should read ���Journal of the American Society of Mass Spectrometry (2019)���,��� not ���Journal of the American Society of Mass Spectrometry (2018)...���
11 February 2019
In this issue, the citation information on the opening page of each article PDF is incorrect. It should read ���Journal of the American Society of Mass Spectrometry (2019)���,��� not ���Journal of the American Society of Mass Spectrometry (2018)...���
Abbreviations
- aa :
-
Amino acid
- AAPH :
-
2,2′-Azobis(2-methylpropionamidine)dihydrochloride
- BP :
-
Benzophenone
- DMPA :
-
2,2-Dimethoxy-2-phenylacetophenone
- CDI :
-
1,1′-Carbonylimidazole
- CID :
-
Collision-induced dissociation
- DAU :
-
1,3-Diallylurea
- DTT :
-
Dithiothreitol
- ESI :
-
Electrospray ionization
- GCAP :
-
Guanylyl cyclase-activating protein
- GSH :
-
Glutathione
- GST :
-
Glutathione-S-transferase
- HCD :
-
Higher energy collision-induced dissociation
- LC :
-
Liquid chromatography
- MS :
-
Mass spectrometry
- MS/MS :
-
Tandem mass spectrometry
- NCE :
-
Normalized collision energy
- NHS :
-
N-Hydroxysuccinimide ester
- TNB :
-
5-Thio-2-nitrobenzoic acid
References
Yu, C., Huang, L.: Cross-linking mass spectrometry: an emerging technology for interactomics and structural biology. Anal. Chem. 90, 144–165 (2018)
Leitner, A., Faini, M., Stengel, F., Aebersold, R.: Crosslinking and mass spectrometry: an integrated technology to understand the structure and function of molecular machines. Trends Biochem. Sci. 41, 20–32 (2016)
Rappsilber, J.: The beginning of a beautiful friendship: cross-linking/mass spectrometry and modelling of proteins and multi-protein complexes. J. Struct. Biol. 173, 530–540 (2011)
Sinz, A.: The advancement of chemical cross-linking and mass spectrometry for structural proteomics: from single proteins to protein interaction networks. Exp. Rev. Proteomics. 11, 733–743 (2014)
Wang, X., Cimermancic, P., Yu, C., Schweitzer, A., Chopra, N., Engel, J.L., Greenberg, C., Huszagh, A.S., Beck, F., Sakata, E.: Molecular details underlying dynamic structures and regulation of the human 26S proteasome. Mol. Cell. Proteomics. 16, 840–854 (2017)
Teimer, R., Kosinski, J., von Appen, A., Beck, M., Hurt, E.: A short linear motif in scaffold Nup145C connects Y-complex with pre-assembled outer ring Nup82 complex. Nat. Commun. 8, 1107 (2017)
Chen, J., Wassarman, K.M., Feng, S., Leon, K., Feklistov, A., Winkelman, J.T., Li, Z., Walz, T., Campbell, E.A.: Darst, S.A.: 6S RNA mimics B-Form DNA to regulate Escherichia coli RNA polymerase. Mol. Cell. 68, 388–397. e386 (2017)
Fernandez-Martinez, J., Kim, S.J., Shi, Y., Upla, P., Pellarin, R., Gagnon, M., Chemmama, I.E., Wang, J., Nudelman, I., Zhang, W.: Structure and function of the nuclear pore complex cytoplasmic mRNA export platform. Cell. 167, 1215–1228. e1225 (2016)
Xu, Y., Bernecky, C., Lee, C.-T., Maier, K.C., Schwalb, B., Tegunov, D., Plitzko, J.M., Urlaub, H., Cramer, P.: Architecture of the RNA polymerase II-Paf1C-TFIIS transcription elongation complex. Nat. Commun. 8, 15741 (2017)
Schneider, M., Belsom, A., Rappsilber, J., Brock, O.: Blind testing of cross-linking/mass spectrometry hybrid methods in CASP11. Proteins. 84, 152–163 (2016)
Sheppard, C., Blombach, F., Belsom, A., Schulz, S., Daviter, T., Smollett, K., Mahieu, E., Erdmann, S., Tinnefeld, P., Garrett, R.: Repression of RNA polymerase by the archaeo-viral regulator ORF145/RIP. Nat. Commun. 7, 13595 (2016)
Yang, L., Zheng, C., Weisbrod, C.R., Tang, X., Munske, G.R., Hoopmann, M.R., Eng, J.K., Bruce, J.E.: In vivo application of photocleavable protein interaction reporter technology. J. Proteome Res. 11, 1027–1041 (2012)
Kaake, R.M., Wang, X., Burke, A., Yu, C., Kandur, W., Yang, Y., Novtisky, E.J., Second, T., Duan, J., Kao, A., Guan, S., Vellucci, D., Rychnovsky, S.D., Huang, L.: A new in vivo cross-linking mass spectrometry platform to define protein-protein interactions in living cells. Mol. Cell. Proteomics. 13, 3533–3543 (2014)
Agou, F., Véron, M.: In vivo protein cross-linking. In: Meyerkord, L.C., Fu, H. (eds.) Springer New York. NY, New York (2015)
Iacobucci, C., Reale, S., De Angelis, F.: Photoactivable amino acid bioisosteres and mass spectrometry: snapshots of in vivo 3D protein structures. Chembiochem. 14, 181–183 (2013)
Suchanek, M., Radzikowska, A., Thiele, C.: Photo-leucine and photo-methionine allow identification of protein-protein interactions in living cells. Nat. Methods. 2, 261–267 (2005)
Piotrowski, C., Ihling, C.H., Sinz, A.: Extending the cross-linking/mass spectrometry strategy: Facile incorporation of photo-activatable amino acids into the model protein calmodulin in Escherichia coli cells. Methods. 89, 121–127 (2015)
Yang, T., Li, X.-M., Bao, X., Fung, Y.M.E., Li, X.D.: Photo-lysine captures proteins that bind lysine post-translational modifications. Nat. Chem. Biol. 12, 70–72 (2016)
Tian, Y., Jacinto, M.P., Zeng, Y., Yu, Z., Qu, J., Liu, W.R., Lin, Q.: Genetically encoded 2-aryl-5-carboxytetrazoles for site-selective protein photo-cross-linking. J. Am. Chem. Soc. 139, 6078–6081 (2017)
Zhang, S., He, D., Lin, Z., Yang, Y., Song, H., Chen, P.R.: Conditional chaperone–client interactions revealed by genetically encoded photo-cross-linkers. Acc. Chem. Res. (2017)
Koole, C., Reynolds, C.A., Mobarec, J.C., Hick, C., Sexton, P.M., Sakmar, T.P.: Genetically encoded photocross-linkers determine the biological binding site of exendin-4 peptide in the N-terminal domain of the intact human glucagon-like peptide-1 receptor (GLP-1R). J. Biol. Chem. 292, 7131–7144 (2017)
Wang, W., Li, T., Felsovalyi, K., Chen, C., Cardozo, T., Krogsgaard, M.: Quantitative analysis of T cell receptor complex interaction sites using genetically encoded photo-cross-linkers. ACS Chem. Biol. 9, 2165–2172 (2014)
Novak, P., Kruppa, G.H.: Intra-molecular cross-linking of acidic residues for protein structure studies. Eur. J. Mass Spectrom. 14, 355–365 (2008)
Schwarz, R., Tänzler, D., Ihling, C.H., Sinz, A.: Monitoring solution structures of peroxisome proliferator-activated receptor β/δ upon ligand binding. PLoS One. 11, e0151412 (2016)
Leitner, A., Joachimiak, L.A., Unverdorben, P., Walzthoeni, T., Frydman, J., Förster, F., Aebersold, R.: Chemical cross-linking/mass spectrometry targeting acidic residues in proteins and protein complexes. Proc. Nat. Acad. Sci. 111, 9455–9460 (2014)
Hage, C., Iacobucci, C., Rehkamp, A., Arlt, C., Sinz, A.: The first zero-length mass spectrometry-cleavable cross-linker for protein structure analysis. Angew. Chem. Int. Ed. 56, 14551–14555 (2017)
Brodie, N.I., Popov, K.I., Petrotchenko, E.V., Dokholyan, N.V., Borchers, C.H.: Solving protein structures using short-distance cross-linking constraints as a guide for discrete molecular dynamics simulations. Sci. Adv. 3, e1700479 (2017)
Schweppe, D.K., Chavez, J.D., Lee, C.F., Caudal, A., Kruse, S.E., Stuppard, R., Marcinek, D.J., Shadel, G.S., Tian, R., Bruce, J.E.: Mitochondrial protein interactome elucidated by chemical cross-linking mass spectrometry. Proc. Natl. Acad. Sci. 114, 1732–1737 (2017)
Schmidt, C., Urlaub, H.: Combining cryo-electron microscopy (cryo-EM) and cross-linking mass spectrometry (CX-MS) for structural elucidation of large protein assemblies. Curr. Opin. Struct. Biol. 46, 157–168 (2017)
Weisz, D.A., Liu, H., Zhang, H., Thangapandian, S., Tajkhorshid, E., Gross, M.L., Pakrasi, H.B.: Mass spectrometry-based cross-linking study shows that the Psb28 protein binds to cytochrome b559 in Photosystem II. Proc. Natl. Acad. Sci. 114, 2224–2229 (2017)
Schilling, B., Row, R.H., Gibson, B.W., Guo, X., Young, M.M.: MS2Assign, automated assignment and nomenclature of tandem mass spectra of chemically crosslinked peptides. J. Am. Soc. Mass Spectrom. 14, 834–850 (2003)
Trnka, M.J., Baker, P.R., Robinson, P.J., Burlingame, A.L., Chalkley, R.J.: Matching cross-linked peptide spectra: only as good as the worse identification. Mol. Cell. Proteomics. 13, 420–434 (2014)
Rasmussen, M.I., Refsgaard, J.C., Peng, L., Houen, G., Hojrup, P.: CrossWork: software-assisted identification of cross-linked peptides. J. Proteome. 74, 1871–1883 (2011)
Iacobucci, C., Sinz, A.: To be or not to be? Five guidelines to avoid misassignments in cross-linking/mass spectrometry. Anal. Chem. 89, 7832–7835 (2017)
Petrotchenko, E.V., Olkhovik, V.K., Borchers, C.H.: Isotopically coded cleavable cross-linker for studying protein-protein interaction and protein complexes. Mol. Cell. Proteomics. 4, 1167–1179 (2005)
Ihling, C., Schmidt, A., Kalkhof, S., Schulz, D.M., Stingl, C., Mechtler, K., Haack, M., Beck-Sickinger, A.G., Cooper, D.M., Sinz, A.: Isotope-labeled cross-linkers and Fourier transform ion cyclotron resonance mass spectrometry for structural analysis of a protein/peptide complex. J. Am. Soc. Mass Spectrom. 17, 1100–1113 (2006)
Petrotchenko, E.V., Serpa, J.J., Borchers, C.H.: Use of a combination of isotopically coded cross-linkers and isotopically coded N-terminal modification reagents for selective identification of inter-peptide crosslinks. Anal. Chem. 82, 817–823 (2010)
Brodie, N.I., Makepeace, K.A., Petrotchenko, E.V., Borchers, C.H.: Isotopically-coded short-range hetero-bifunctional photo-reactive crosslinkers for studying protein structure. J. Proteome. 118, 12–20 (2015)
Hage, C., Falvo, F., Schäfer, M., Sinz, A.: Novel concepts of MS-cleavable cross-linkers for improved peptide structure analysis. J. Am. Soc. Mass Spectrom. 28, 2022–2038 (2017)
Soderblom, E.J., Goshe, M.B.: Collision-induced dissociative chemical cross-linking reagents and methodology: applications to protein structural characterization using tandem mass spectrometry analysis. Anal. Chem. 78, 8059–8068 (2006)
Arlt, C., Götze, M., Ihling, C.H., Hage, C., Schäfer, M., Sinz, A.: Integrated workflow for structural proteomics studies based on cross-linking/mass spectrometry with an MS/MS cleavable cross-linker. Anal. Chem. 88, 7930–7937 (2016)
Liu, F., Rijkers, D.T.S., Post, H., Heck, A.J.R.: Proteome-wide profiling of protein assemblies by cross-linking mass spectrometry. Nat. Methods. 12, 1179 (2015)
Kao, A.H., Chiu, C.L., Vellucci, D., Yang, Y.Y., Patel, V.R., Guan, S.H., Randall, A., Baldi, P., Rychnovsky, S.D., Huang, L.: Development of a novel cross-linking strategy for fast and accurate identification of cross-linked peptides of protein complexes. Mol. Cell. Proteomics. 10, M110.002212 (2011)
Müller, M.Q., Dreiocker, F., Ihling, C.H., Schäfer, M., Sinz, A.: Cleavable cross-linker for protein structure analysis: reliable identification of cross-linking products by tandem MS. Anal. Chem. 82, 6958–6968 (2010)
Tang, X., Munske, G.R., Siems, W.F., Bruce, J.E.: Mass spectrometry identifiable cross-linking strategy for studying protein−protein interactions. Anal. Chem. 77, 311–318 (2005)
Tang, X., Bruce, J.E.: A new cross-linking strategy: protein interaction reporter (PIR) technology for protein–protein interaction studies. Mol. BioSyst. 6, 939–947 (2010)
Iacobucci, C., Hage, C., Schäfer, M., Sinz, A.: A novel MS-cleavable azo cross-linker for peptide structure analysis by free radical initiated peptide sequencing (FRIPS). J. Am. Soc. Mass Spectrom. 28, 2039–2053 (2017)
Iacobucci, C., Götze, M., Piotrowski, C., Arlt, C., Rehkamp, A., Ihling, C.H., Hage, C., Sinz, A.: Carboxyl-photo-reactive ms-cleavable cross-linkers: unveiling a hidden aspect of diazirine-based reagents. Anal. Chem. 90, 2805–2809 (2018)
Liu, F., Lössl, P., Scheltema, R., Viner, R., Heck, A.J.R.: Optimized fragmentation schemes and data analysis strategies for proteome-wide cross-link identification. Nat. Commun. 8, 15473 (2017)
Anderson, G.A., Tolic, N., Tang, X., Zheng, C., Bruce, J.E.: Informatics strategies for large-scale novel cross-linking analysis. J. Prot. Res. 6, 3412–3421 (2007)
Zheng, C., Yang, L., Hoopmann, M.R., Eng, J.K., Tang, X., Weisbrod, C.R., Bruce, J.E.: Cross-linking measurements of in vivo protein complex topologies. Mol. Cell. Proteomics. 10, M110. 006841 (2011)
Stenzel, M.H.: Bioconjugation using thiols: old chemistry rediscovered to connect polymers with nature’s building blocks. ACS Macro Lett. 2, 14–18 (2013)
Yang, B., Tang, S., Ma, C., Li, S.-T., Shao, G.-C., Dang, B., DeGrado, W.F., Dong, M.-Q., Wang, P.G., Ding, S.: Spontaneous and specific chemical cross-linking in live cells to capture and identify protein interactions. Nat. Commun. 8, 2240 (2017)
Sinz, A., Wang, K.: Mapping protein interfaces with a fluorogenic cross-linker and mass spectrometry: application to nebulin-calmodulin complexes. Biochemistry. 40, 7903–7913 (2001)
Dondoni, A., Marra, A.: Recent applications of thiol–ene coupling as a click process for glycoconjugation. Chem. Soc. Rev. 41, 573–586 (2012)
Torres-Kolbus, J., Chou, C., Liu, J., Deiters, A.: Synthesis of non-linear protein dimers through a genetically encoded thiol-ene reaction. PLoS One. 9, e105467 (2014)
Colak, B., Da Silva, J.C., Soares, T.A., Gautrot, J.E.: Impact of the molecular environment on thiol–ene coupling for biofunctionalization and conjugation. Bioconjug. Chem. 27, 2111–2123 (2016)
Hoyle, C.E., Bowman, C.N.: Thiol-ene click chemistry. Angew. Chem. Int. Ed. 49, 1540–1573 (2010)
McKay, C.S., Finn, M.G.: Click chemistry in complex mixtures: bioorthogonal bioconjugation. Chem. Biol. 21, 1075–1101 (2014)
Brose, N., Rosenmund, C., Rettig, J.: Regulation of transmitter release by Unc-13 and its homologues. Curr. Opin. Neurobiol. 10, 303–311 (2000)
Gorczyca, W.A., Graykeller, M.P., Detwiler, P.B., Palczewski, K.: Purification and physiological evaluation of a guanylate-cyclase activating protein from retinal rods. Proc. Natl. Acad. Sci. 91, 4014–4018 (1994)
Schröder, T., Lilie, H., Lange, C.: The myristoylation of guanylate cyclase-activating protein-2 causes an increase in thermodynamic stability in the presence but not in the absence of Ca2+. Protein Sci. 20, 1155–1165 (2011)
Schaks, S., Maucher, D., Ihling, C.H., Sinz, A.: Investigation of a calmodulin/peptide complex by chemical cross-linking and high-resolution mass spectrometry. Biomacromol. Mass Spectrom. 2, 249–260 (2011)
Lössl, P., Sinz, A.: Combining amine-reactive cross-linkers and photo-reactive amino acids for 3D-structure analysis of proteins and protein complexes. Meth. Mol. Biol. 109–127 (2016)
Götze, M., Pettelkau, J., Fritzsche, R., Ihling, C.H., Schäfer, M., Sinz, A.: Automated assignment of MS/MS cleavable cross-links in protein 3D-structure analysis. J. Am. Soc. Mass Spectrom. 26, 83–97 (2015)
Krzywinski, M., Schein, J., Birol, I., Connors, J., Gascoyne, R., Horsman, D., Jones, S.J., Marra, M.A.: Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009)
Acknowledgments
The authors would like to thank Dirk Tänzler for GCAP-2 expression and purification, Christoph Hage for valuable discussions, and Dr. Michael Götze for continuous improvements of the MeroX software.
Funding
A.S. gratefully acknowledges financial support by the DFG (project Si 867/15-2). C.I. is funded by the Alexander von Humboldt Foundation.
Author information
Authors and Affiliations
Corresponding authors
Electronic Supplementary Material
ESM 1
(DOCX 34 kb)
Rights and permissions
About this article
Cite this article
Iacobucci, C., Piotrowski, C., Rehkamp, A. et al. The First MS-Cleavable, Photo-Thiol-Reactive Cross-Linker for Protein Structural Studies. J. Am. Soc. Mass Spectrom. 30, 139–148 (2019). https://doi.org/10.1007/s13361-018-1952-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-018-1952-8