Abstract
Thymidine (dThd) is a fundamental building block of DNA nucleic acids, whereas 5-methyluridine (Thd) is a common modified nucleoside found in tRNA. In order to determine the conformations of the sodium cationized thymine nucleosides [dThd+Na]+ and [Thd+Na]+ produced by electrospray ionization, their infrared multiple photon dissociation (IRMPD) action spectra are measured. Complementary electronic structure calculations are performed to determine the stable low-energy conformations of these complexes. Geometry optimizations and frequency analyses are performed at the B3LYP/6-311+G(d,p) level of theory, whereas energies are calculated at the B3LYP/6-311+G(2d,2p) level of theory. As protonation preferentially stabilizes minor tautomers of dThd and Thd, tautomerization facilitated by Na+ binding is also considered. Comparisons of the measured IRMPD and computed IR spectra find that [dThd+Na]+ prefers tridentate (O2,O4',O5') coordination to the canonical 2,4-diketo form of dThd with thymine in a syn orientation. In contrast, [Thd+Na]+ prefers bidentate (O2,O2') coordination to the canonical 2,4-diketo tautomer of Thd with thymine in an anti orientation. Although 2,4-dihydroxy tautomers and O2 protonated thymine nucleosides coexist in the gas phase, no evidence for minor tautomers is observed for the sodium cationized species. Consistent with experimental observations, the computational results confirm that the sodium cationized thymine nucleosides exhibit a strong preference for the canonical form of the thymine nucleobase. Survival yield analyses based on energy-resolved collision-induced dissociation (ER-CID) experiments suggest that the relative stabilities of protonated and sodium cationized dThd and Thd follow the order [dThd+H]+ < [Thd+H]+ < [dThd+Na]+ < [Thd+Na]+.

ᅟ
Similar content being viewed by others
Introduction
The overall structures of DNA and RNA are highly impacted by the conformations of their nucleoside building blocks (i.e., the nucleobase orientation and sugar puckering of the 2'-deoxyribose and ribose moieties). Anti orientations of the nucleobase residues and C2'-endo sugar puckering of the sugar moieties are found in the most common B-form of double-stranded (ds) DNA. A change in sugar puckering to C3'-endo results in the A-forms of DNA and RNA, which are generally found in RNA-DNA and RNA-RNA complexes [1]. The guanine residues take on syn orientations with C3'-endo sugar puckering of the sugars and left-handed coiling of the strands in the less common Z-form of ds DNA [2]. Nucleobase flipping has been observed as an important step during base excision repair to enable the enzyme access to the target base [3,4,5]. Thus, the determination of nucleoside conformation is of fundamental importance to fully understand both static and dynamic nucleic acid structures. The local environment also influences the structures of DNA and RNA. DNA triplex structures and the i-motif are stabilized by protons [6, 7], whereas B-form DNA and G-quadruplex structures are stabilized by alkali metal cations [8,9,10,11,12]. Sodium cations prefer to bind to the negatively charged phosphate backbone at low concentrations [13], whereas sodium cations may interact with the nucleobase and/or sugar moieties at higher concentrations. Metal ion binding to the nucleobase residues can lead to more profound influences on DNA conformation than those binding to the phosphate backbone [14].
The N-glycosidic bond, the covalent bond linking the nucleobase residue and sugar moiety, is important to the overall stability of DNA and RNA. Mutations in the DNA and RNA sequence via nucleobase excision or mismatch are influenced by the N-glycosidic bond stabilities [15]. N-glycosidic bond stability/cleavage is investigated here because of its role in biochemical processes that occur in vivo, such as nucleobase salvage and DNA repair [15,16,17]. For example, base excision repair (BER) is one of the cellular mechanisms available to repair damaged or inappropriate nucleobases in DNA, and explicitly involves cleavage of the N-glycosidic bond [18, 19]. Modifications on the nucleobase and sugar moiety have been found to affect the stabilities of N-glycosidic bonds [20,21,22]. Additionally, both theoretical and experimental studies demonstrate that the local environment of a nucleic acid also impacts N-glycosidic bond stabilities [23,24,25].
Thymine (Thy) is one of the naturally occurring pyrimidine nucleobases found in DNA. It is similar to uracil, but possesses a methyl substituent at the 5-position. Thy has been well studied because of its slow dissipation of electronic energy upon optical excitation [26, 27]. UV-induced carcinogenesis and mutagenesis may be caused by ultraviolet light absorption by Thy [28, 29]. Another interesting characteristic of Thy is tautomerization that may occur to the canonical 2,4-diketo structure, which may lead to point mutations and molecular based diseases resulting from nucleobase mismatches of tautomeric Thy [30]. The tautomerization of Thy has been examined both experimentally and theoretically [31,32,33,34]. For the canonical form of Thy, sodium cations show a preference for binding to the O4 position over the O2 position [35, 36]. However, a theoretical study found that tautomeric Thy with the O4 hydrogen atom pointed toward the 5-methyl substituent and sodium cation binding to the O2 and N3 atoms exhibits the largest total stabilization energy [36]. Thymidine (dThd) comprised of a Thy nucleobase and 2'-deoxyribose sugar is a naturally occurring nucleoside of DNA. 5-Methyluridine (Thd) is the ribonucleoside counterpart of dThd, but is not a common RNA nucleoside. Thd is found as a natural modified nucleoside in tRNA [37, 38]. In addition, Thd is an important reagent for the synthesis of the anti-AIDS drugs AZT and D4T [39, 40]. Thus, it is interesting to investigate the effects of the sugar moiety and in particular the 2'-hydroxyl substituent on sodium cation binding preferences of the Thy nucleosides.
Due to the fundamental importance of understanding the effects of protonation versus sodium cationization on the gas-phase conformations and N-glycosidic bond stabilities of dThd and Thd, infrared multiple photon dissociation (IRMPD) action spectroscopy studies of the sodium cationized Thy nucleosides, [dThd+Na]+ and [Thd+Na]+, and energy-resolved collision-induced dissociation (ER-CID) experiments of the protonated and sodium cationized Thy nucleosides are performed. IRMPD action spectroscopy studies of the protonated Thy nucleosides, [dThd+H]+ and [Thd+H]+, have previously been reported [41, 42]. The conformations and vibrational frequencies of [dThd+Na]+ and [Thd+Na]+ are calculated using density functional theory (DFT) at the B3LYP/6-311+G(d,p) level of theory, whereas energetics are determined at the B3LYP/6-311+G(2d,2p) level of theory. Comparisons between the experimental IRMPD and calculated IR spectra enable the determination of the stable low-energy conformations populated in the experiments. This approach has proven to be a robust technique for structural characterization of deprotonated, protonated, and metal cationized nucleobases [34,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,, 43–63], nucleosides [41,65,66,67,68,69,70,71,72,, 42, 64–73], and nucleotides [74,75,76,77,78,79,80,81,82]. The relative stabilities of the protonated and sodium cationized Thy nucleosides are determined via survival yield analyses based on the ER-CID experiments of these species. In particular, the relative N-glycosidic bond stabilities are explicitly determined when the major fragmentation pathways observed for these species involve solely N-glycosidic bond cleavage. By comparing the present results with previous IRMPD action spectroscopy and theoretical studies of protonated dThd and Thd [41], and sodium cationized uracil (Ura) nucleosides, uridine (Urd), and 2'-deoxyuridine (dUrd) [73], the effects of protonation versus sodium cationization and the 5-methyl substituent on the gas-phase conformations and energetics are elucidated.
Experimental and Computational
Fourier Transform Ion Cyclotron Resonance Mass Spectrometry and Photodissociation
IRMPD action spectroscopy experiments of [dThd+Na]+ and [Thd+Na]+ were performed using a custom-built 4.7 T Fourier transform ion cyclotron resonance mass spectrometer (FT-ICR MS) coupled to the free-electron laser for infrared experiments (FELIX) or an optical parametric oscillator/amplifier (OPO/OPA) laser system [83,84,85,86]. One mM dThd or Thd (purchased from Sigma-Aldrich, St. Louis, MO, USA) was dissolved in a 50%:50% (v/v) methanol:water mixture with 1 mM NaCl (the solvents and NaCl were purchased from Sigma-Aldrich, Zwijndrecht, The Netherlands). [dThd+Na]+ or [Thd+Na]+ were formed from the analyte solutions via introduction into a Micromass “Z-spray” electrospray ionization (ESI) source at a flow rate of ~10 μL/min. The ions were accumulated in an rf hexapole ion trap for several seconds to enhance ion signal and provide efficient thermalization, pulsed extracted via a quadrupole deflector, and then injected into the ICR cell through a 1 m long rf octopole ion guide with a pulsed positive DC bias applied to reduce collisional heating of the ions [84]. All ions were stored in the ICR cell for at least 300 ms for further thermalization by radiative emission. The ions of interest, [dThd+Na]+ or [Thd+Na]+, were isolated using stored waveform inverse Fourier transform (SWIFT) techniques. The isolated sodium cationized Thy nucleosides were irradiated for ~3 s by the free-electron laser over the frequency range between ~550 and 1850 cm–1 or for ~8 s by the OPO laser over the frequency range between ~3300 and 3800 cm–1 to induce IR photodissociation. Very low IRMPD yields for [Thd+Na]+ were observed when IR dissociation was induced by the OPO laser. Therefore, a CO2 laser was also used for the OPO experiments to facilitate excitation and dissociation of [Thd+Na]+ to improve the IRMPD efficiency. The IRMPD yield was calculated at each frequency using Equation 1,
where \( \sum_i{I}_{f_i} \) is the total fragment ion intensity, I p is the precursor ion intensity, and (\( {I}_p+\sum_i{I}_{f_i}\Big) \) is the total ion intensity. The experimental IRMPD spectra were generated by plotting the IRMPD yields as a function of IR frequency.
Quadrupole Ion Trap Mass Spectrometry and Energy-Resolved Collision-Induced Dissociation
ER-CID experiments of [dThd+H]+, [Thd+H]+, [dThd+Na]+, and [Thd+Na]+ were performed using a Bruker amaZon ETD quadrupole ion trap mass spectrometer (QIT MS) (Bruker Daltonics, Bremen, Germany). Ten μM dThd or Thd was dissolved in a 50%:50% (v/v) methanol:water mixture. One percent (v/v) acetic acid (purchased from Mallinckrodt Chemicals, St. Louis, MO, USA) was added to facilitate formation of [dThd+H]+ and [Thd+H]+, whereas 10 μM NaCl was added to facilitate formation of [dThd+Na]+ and [Thd+Na]+. The protonated and sodium cationized Thy nucleosides were formed using an Apollo ESI source operated at a flow rate of 3 μL/min. Helium was introduced into the ion trap at a stagnation pressure of ~1 mTorr for efficient trapping and cooling of the ions, and also served as the collision gas for the ER-CID experiments. The low mass cut-off was set to 27% of the precursor ion m/z, corresponding to a qz value of 0.25. The rf excitation amplitude (rfEA) was varied from 0.00 V to the rfEA required to completely dissociate the precursor ion at a step size of 0.01 V. The ER-CID experiments were performed in triplicate to assess reproducibility of the measurements. Data analysis was performed using Compass Data Analysis Software ver. 4.0 (Bruker Daltonics, Bremen, Germany).
Computational Details
The chemical structures of the canonical forms of neutral dThd and Thd including the numbering of the nucleobase and sugar atoms are shown in Figure 1. All candidate conformers of [dThd+Na]+ and [Thd+Na]+ subjected to DFT calculation were generated via simulated annealing processes using HyperChem software [87] with the Amber 3 force field. All possible favorable binding sites of Na+ (i.e., the O2, N3, O4, O2', O3', O4', and O5' atoms) to canonical 2,4-diketo and tautomeric 2-keto-4-hydroxy (t4) and 2-hydroxy-4-keto (t2) forms of the Thy nucleosides were considered when constructing the initial structures employed for simulated annealing. Each initial structure was subjected to 300 cycles of annealing. Each cycle began with 0.3 ps of thermal heating from 0 to 1000 K, followed by sampling of conformational space at 1000 K for 0.2 ps, and then 0.3 ps of thermal cooling from 1000 to 0 K. The resulting structure was optimized to a local minimum using the Amber 3 force field and saved as a snapshot, which was then used to initiate the next annealing cycle. Based on the relative stabilities of the resulting snapshots found in the simulated annealing process, more than 600 candidate conformers were chosen for higher level DFT calculation. Geometry optimizations and harmonic vibrational frequency analyses were performed at the B3LYP/6-311+G(d,p) level of theory, whereas single point energies were calculated at the B3LYP/6-311+G(2d,2p) level of theory using the Gaussian 09 suite of programs [88]. The relative Gibbs free energies at 298 K determined from the DFT calculations were used to assess the relative stabilities of the stable conformers found for [dThd+Na]+ and [Thd+Na]+. The vibrational frequencies were scaled by a factor of 0.978 in the FELIX region, whereas the vibrational frequencies were scaled by a factor of 0.954 in the OPO region. In order to better reproduce the measured IRMPD spectra, the computed vibrational frequencies were convoluted with 20 and 15 cm–1 FWHM Gaussian line shapes over the FELIX and OPO regions, respectively. To further assess the relative stabilities of the two lowest energy conformers computed for [Thd+Na]+ in aqueous environments, the mono- and dihydrated complexes of these species were also examined.
Chemical structures of thymidine (dThd) and 5-methyluridine (Thd). Numbering of the Thy nucleobase and sugar moieties is shown. The ground T1(O2O4′O5′) conformers of [dThd+Na]+ and [Thd+Na]+ as well as the low-energy B1(O2O2') conformer of [Thd+Na]+ predicted at the B3LYP/6-311+G(2d,2p)//B3LYP/6-311+G(d,p) level of theory (relative 298 K Gibbs free energies) are shown with the sodium cation binding mode, Thy orientation, and sugar puckering. The ground conformers of [dThd+H]+ and [Thd+H]+ are also shown for comparison and taken from reference [41]
Survival Yield Analyses
Survival yield analysis has proven to be a robust method to elucidate the relative stabilities of protonated and sodium cationized nucleosides [70,71,72,73] as well as those of a variety of other ions [89,90,91,92,93]. Survival yields based on the ER-CID experiments of the protonated and sodium cationized Thy nucleosides at each rfEA were calculated using Equation 2 [93],
where I p and (\( {I}_p+\sum_i{I}_{f_i}\Big) \) are as defined in Equation 1. Survival yields were calculated using custom software developed in our laboratory. The survival yield was plotted as a function of rfEA to generate the survival yield curves. Each survival yield curve was least-squares fit to the four parameter logistic curve summarized in Equation 3,
where the min and max are the minimum (0) and maximum (1) values of the survival yield, and Hillslope is the slope of the declining region of the survival yield curve. The relative stabilities of the protonated and sodium cationized dThd and Thd nucleosides are determined by comparing the rf excitation amplitudes required to produce 50% dissociation of the precursor ions, and designated as CID50%. Data analyses were performed using SigmaPlot 10.0 (Systat Software, Inc., San Jose, CA, USA).
Results
IRMPD Action Spectroscopy
The IR photodissociation pathways observed for [dThd+Na]+ and [Thd+Na]+ are summarized in Reactions 4 and 5,
where Nuo is either dThd or Thd. IR photodissociation of [dThd+Na]+ induced by either FELIX or the OPO laser produced both simple cleavage of the noncovalent interaction between Na+ and the nucleoside, Reaction 4, and cleavage of the N-glycosidic bond with Na+ retained by the Thy nucleobase, Reaction 5. In contrast, IR photodissociation of [Thd+Na]+ induced by the FEL led solely to simple loss of the neutral nucleoside, Reaction 4, whereas in the OPO region, both Reactions 4 and 5 were observed. The measured IRMPD spectra of [dThd+Na]+ and [Thd+Na]+ are compared in Figure 2. The most significant difference observed in the experimental IRMPD action spectra of [dThd+Na]+ and [Thd+Na]+ is the IRMPD yield, where the IRMPD yield of [dThd+Na]+ exceeds that of [Thd+Na]+ in the FELIX region, particularly at lower frequencies. Note that the IRMPD efficiency of [Thd+Na]+ for the OPO experiments was improved by also using a CO2 laser to facilitate excitation and dissociation. Therefore, although the plotted yields look very similar in this region, it is clear that the IRMPD yield of [dThd+Na]+ must exceed that of [Thd+Na]+ when equivalent excitation conditions are employed. Absence of the N-glycosidic bond cleavage pathway for [Thd+Na]+ in the FELIX region and the need for additional excitation via the CO2 laser for [Thd+Na]+ in the OPO region suggest that the 2'-hydroxyl substituent of Thd stabilizes the N-glycosidic bond relative to that of dThd. Similar behavior was observed for sodium cationized adenine and uracil nucleosides [69, 72]. This result is also consistent with the ER-CID experimental results discussed below. In the FELIX region, three additional IR features at 772, 862, and 944 cm–1 are observed for [dThd+Na]+ that are not seen for [Thd+Na]+. In addition, the two intense IR features at 1680 and 1750 cm–1 in the spectrum of [dThd+Na]+ are slightly red shifted in the spectrum of [Thd+Na]+. In contrast, the experimental IRMPD spectra of these two species are parallel in the OPO region. These minor differences in the experimental IRMPD spectra suggest that both [dThd+Na]+ and [Thd+Na]+ involve the same tautomeric form of the Thy nucleobase, but may exhibit different modes of Na+ binding or conformations due to the presence of the 2'-hydroxyl substituent of Thd. Parallel behaviors have been observed for the other pyrimidine nucleosides (i.e., both the cytosine (Cyt) and Ura nucleosides [72, 73].
IRMPD action spectra of [dThd+Na]+ and [Thd+Na]+ in the fingerprint (FELIX) and hydrogen-stretching (OPO) regions. IRMPD action spectra of [dThd+H]+ and [Thd+H]+ previously reported are overlaid in grey for comparison and taken from reference [41]
ER-CID Experiments
CID mass spectra of [dThd+H]+, [Thd+H]+, [dThd+Na]+, and [Thd+Na]+ acquired at an rf excitation amplitude that results in ~50% dissociation are shown in Figure 3. Survival/fragment yields for these systems determined from the ER-CID experiments over the entire range of rf excitation amplitudes examined are shown in Supplementary Figure S1. N-glycosidic bond cleavage processes are observed as the dominant fragmentation pathways for [Thd+H]+, [dThd+Na]+, and [Thd+Na]+, where the cation, H+ or Na+, is retained either by the Thy nucleobase or the sugar as summarized in Reactions 6 and 7.
In all cases, Reaction 6 is favored over Reaction 7. Although Reaction 6 remains the dominant CID pathway for [dThd+H]+, additional fragmentation pathways involving elimination of ketene (K, H2C=C=O), or both ketene and a water molecule (W) are observed, as summarized in Reactions 8 and 9, and become more competitive with glycosidic bond cleavage.
Four additional minor fragmentation pathways are also observed for [dThd+H]+ that involve elimination of one or two W molecules, or elimination of Thy and one or two W molecules, as summarized in Reactions 10, 11, 12, and 13.
Reaction 10 is also observed as a minor fragmentation pathway for [Thd+H]+. Reactions 9 and 11–13 obviously involve sequential fragmentation pathways. To determine further details regarding the order of dissociation of these sequential reaction pathways observed in the CID of [dThd+H]+, MS3 experiments were performed for the [dThd–W+H]+, [dThd–2W+H]+, [dThd–K+H]+, and [dThd-K-W+H]+ ionic products (see Supplementary Figure S2). MS3 experiments could not be performed for [dThd–Thy+H]+, [dThd–Thy–W+H]+, and [dThd–Thy–2W+H]+ due to the very low intensities of these species observed over the entire range of rf excitation amplitudes examined in the CID experiments. [dThd–K–W+H]+ (Reaction 9) can be formed via sequential loss of K from [dThd–W+H]+ or W from [dThd–K+H]+ (Reactions 10 and 8, see top and third panels of Supplementary Figure S2), respectively. [dThd–2W+H]+ is obviously formed via sequential loss of W from [dThd–W+H]+ (Reaction 11, top panel of Supplementary Figure S2). [dThd–Thy–W+H]+ is formed sequentially from [dThd–W+H]+ (Reaction 12, top panel of Supplementary Figure S2). However, the reverse order of dissociation could not be examined or confirmed as the intensity of the [dThd–Thy+H]+ was too low. [dThd–Thy–2W+H]+ is formed sequentially from [dThd–2W+H]+ via glycosidic bond cleavage (Reaction 13, second panel of Supplementary Figure S2). It is clear that when glycosidic bond cleavage occurs after elimination of water, the proton is preferentially retained by the dehydrated sugar moiety. Successive losses of K, W, and formaldehyde as well as glycosidic bond cleavage dominate the dissociation behavior of these fragment ions.
Theoretical Results
The nomenclature employed to describe the stable conformers of [dThd+Na]+ and [Thd+Na]+ uses an uppercase letter to indicate the number of chelation interactions between the sodium cation and Thy nucleoside (i.e., T for tridentate, B for bidentate, and M for monodentate). A number following the uppercase letter indicates the order of relative Gibbs free energies at 298 K of the stable conformers found that exhibit the same mode of Na+ binding. Structures that involve binding to the 2-keto-4-hydroxy and 2-hydroxy-4-keto tautomeric conformations of Thy are appended by t4 and t2, respectively. Supplementary Figure S3 introduces two designations for describing the Thy orientation and sugar puckering, which have been described previously [70,71,72,73]. Table 1 lists representative low-energy conformers computed for [dThd+Na]+ and [Thd+Na]+ along with their relative enthalpies at 0 and 298 K, relative Gibbs free energies at 298 K, pseudorotation phase angles (P), orientations of the Thy nucleobase, and sugar puckerings. Supplementary Figures S4 and S5 show all stable conformers found for [dThd+Na]+ and [Thd+Na]+ along with the orientations of the Thy nucleobase, sugar puckerings, and their relative Gibbs free energies at 298 K.
Ground Conformers of [dThd+Na]+ and [Thd+Na]+
The ground conformers of [dThd+Na]+ and [Thd+Na]+ computed at B3LYP/6-311+G(2d,2p)//B3LYP/6-311+G(d,p) level of theory are shown in Figure 1, while geometric information for these species is given in Table 2. The ground conformers calculated for [dThd+Na]+ and [Thd+Na]+ are highly parallel. They both exhibit a tridentate mode of Na+ binding to the O2, O4', and O5' atoms, forming 5- and 6-membered chelation rings, with a syn orientation of Thy, and similar O4'-endo sugar puckering that shifts from OT1 for [dThd+Na]+ to OT4 for [Thd+Na]+ because of the influence of the hydrogen bonding interaction between the 2'- and 3'-hydroxyl substituents. The main difference between the two ground conformers is an additional hydrogen-bonding interaction between the 2'- and 3'-hydroxyl substituents found for [Thd+Na]+. The N-glycosidic bond length calculated for the T1(O2O4'O5') conformer of [dThd+Na]+ is 1.458 Å, which slightly decreases to 1.453 Å in the T1(O2O4'O5') conformer of [Thd+Na]+.
Sodium Cation Binding to O2
For [dThd+Na]+, tridentate binding to the O2, O4', and O5' atoms is the most favorable mode of Na+ binding. Elimination of one (or both) chelation interactions between Na+ and the O5' (and O4') atoms of the sugar moiety significantly increases the relative Gibbs free energy by at least 30.0 and 59.2 kJ/mol, respectively (see the B1(O2O4') and M1(O2) conformers in Supplementary Figure S4). All anti oriented bidentate O2 and O3' binding conformers of [dThd+Na]+ are much less favorable, and are at least 51.4 kJ/mol less stable than the ground conformer (see the B1(O2O3') conformer in Supplementary Figure S4). Similarly, the most favorable mode of Na+ binding for [Thd+Na]+ involves Na+ binding to the O2, O4', and O5' atoms. The bidentate O2 binding conformers of [Thd+Na]+ that exhibit an anti orientation of Thy are at least 6.6 kJ/mol less favorable than the ground conformer (see the B1(O2O2') conformer in Figure 1 and Supplementary Figure S5). Lack of one chelation interaction between Na+ and the O5' atom of the sugar moiety increases the relative Gibbs free energy by at least 31.9 kJ/mol (see the B1(O2O4') conformer in Supplementary Figure S5). No stable monodentate O2 binding conformer is found for [Thd+Na]+. Unique tridentate O2, O2', and O3' binding conformers are only found for [Thd+Na]+, and are at least 19.2 kJ/mol less stable than the ground conformer (see the T1(O2O2'O3') conformer in Supplementary Figure S5).
Sodium Cation Binding to O4
For both [dThd+Na]+ and [Thd+Na]+, Na+ binding to the O4 atom is much less favorable than binding to the O2 atom. The most stable O4 binding conformers calculated for both [dThd+Na]+ and [Thd+Na]+ are M1(O4) conformers, which are 39.4 and 35.4 kJ/mol less stable than the corresponding ground conformers, respectively (see Supplementary Figures S4 and S5). Both anti and syn oriented O4 binding conformers are found with a strong preference for the anti orientation. The most stable syn oriented O4 binding conformers calculated for [dThd+Na]+ and [Thd+Na]+ are 53.1 and 45.6 kJ/mol less favorable than the corresponding ground conformers, respectively (see the M10(O4) conformer in Supplementary Figure S4 and the M4(O4) conformer in Supplementary Figure S5). No stable bidentate or tridentate O4 binding conformers are found for either [dThd+Na]+ or [Thd+Na]+. Clearly, the change in preference for O2 over O4 binding to Thy for the Thy nucleosides is driven by the additional stabilization provided by multiple chelation interactions of Na+ with Thy and the sugar moiety.
Sodium Cation Binding to the Sugar Moiety
The most stable sugar binding conformers calculated for [dThd+Na]+ and [Thd+Na]+ are the B1(O3'O5') conformer of [dThd+Na]+ and the B1(O2'O3') conformer of [Thd+Na]+, which are 46.8 and 25.0 kJ/mol less stable than the corresponding ground conformers, respectively. These results suggest that the stability and binding preferences of these species are dominated by the Thy nucleobase, and also by additional chelation interactions with the sugar moiety. However, the 2'-hydroxyl substituent of Thd is also an important determinant of the conformation and stability of [Thd+Na]+.
Sodium Cation Binding to Tautomeric Thymine Nucleosides
All tautomeric [dThd+Na]+ and [Thd+Na]+ are found to be less favorable than the canonical forms of these species. The most stable sodium cationized t4 conformers of [dThd+Na]+ and [Thd+Na]+ are 42.0 and 37.6 kJ/mol less favorable than the corresponding ground conformers, respectively (see the B1(O2O5')t4 conformers in Supplementary Figures S4 and S5). The sodium cationized t2 conformers are even less favorable than the sodium cationized t4 conformers. The most stable sodium cationized t2 conformers are the syn oriented B1(N3O4)t2 conformer of [dThd+Na]+, which is 53.7 kJ/mol less stable, and the anti oriented B1(N3O4)t2 conformer of [Thd+Na]+, which is 51.7 kJ/mol less stable than the corresponding ground conformers (see Supplementary Figures S4 and S5). Therefore, conformers involving Na+ binding to the tautomeric Thy nucleosides are unlikely to be important in the experiments.
Sugar Puckerings of [dThd+Na]+ and [Thd+Na]+
C1'-exo, C2'-endo, C2'-exo, C3'-endo, C3'-exo, C4'-endo, C4'-exo, and O4'-endo sugar puckerings are found among the stable conformers of both [dThd+Na]+ and [Thd+Na]+. O4'-exo sugar puckering is not found for either species. C1'-endo sugar puckering is only found for [Thd+Na]+ [see the B1(O2'O4') conformer at 91.7 kJ/mol in Supplementary Figure S5]. The ground conformers of [dThd+Na]+ and [Thd+Na]+ exhibit slightly different O4'-endo sugar puckering (OT1 and OT4), respectively. The effect of sugar puckering on the relative Gibbs free energies of the sodium cationized Thy nucleosides is clearly not as significant as the orientation and tautomeric form of the Thy nucleobase or the mode of Na+ binding.
Solvation of [Thd+Na]+
Figure 4 compares the conformations and relative Gibbs free energies of the T1(O2O4'O5') and B1(O2O2') conformers of [Thd+Na]+ and their mono and bis water adducts. The anti oriented B1(O2O2') conformer is only 6.6 kJ/mol less stable than the syn oriented T1(O2O4'O5') conformer. Binding of one water molecule to [Thd+Na]+ preferentially stabilizes the B1(O2O2') conformer, such that the anti oriented B1(O2O2')•H2O conformer is now very slightly favored (by 0.1 kJ/mol) over the syn oriented T1(O2O4'O5')•H2O conformer. Binding of a second water molecule further stabilizes the anti oriented B1(O2O2') conformer of [Thd+Na]+ relative to the syn oriented T1(O2O4'O5') conformer such that T1(O2O4'O5')•2H2O is 9.8 kJ/mol less stable than the anti oriented B1(O2O2')•2H2O conformer. These results suggest that [Thd+Na]+ prefers an anti orientation with the sodium cation binding to the O2 and O2' atoms of Thd in aqueous solution. This behavior parallels that found for the other sodium cationized pyrimidine nucleosides with Cyt and Ura nucleobases [72, 73].
Discussion
Comparison of Experimental IRMPD and Calculated IR Spectra of [dThd+Na]+
Figure 5 compares the experimental IRMPD and calculated IR spectra of [dThd+Na]+ for several conformers that exhibit good agreement. Supplementary Figures S6–S10 show analogous comparisons for representative conformers of [dThd+Na]+ that exhibit spectral misalignments (shaded in red). Overall, the calculated IR spectra of the T1, T2, and T3(O2O4'O5') conformers exhibit the best agreement with the experimental IRMPD spectrum. These conformers all exhibit tridentate binding of Na+, with a syn orientation of Thy, but with different sugar puckerings, O4'-endo (OT1), C1'-exo (1TO), and C2'-exo (2T3), respectively (see Figure 5). Changes of the sugar puckering and rotation of the N-glycosidic bond lead to the splitting of the IR feature predicted at ~3664 cm–1, indicating that the T4, T5, T6, and T7(O2O4'O5') conformers do not have measurable populations in the experiments (see Supplementary Figure S6). Minor shifts of the features in the range from ~949 to 1050 cm–1 are predicted in the calculated IR spectrum for the B1(O2O4') conformer, suggesting that if present, this conformer is in low abundance in the experiments (see Supplementary Figure S7). The IR spectra calculated for the B1(O2O3') and M1(O2) conformers exhibit significant spectral misalignment of the strong IR absorption observed at ~1686 cm–1, and a slight splitting of the IR absorption observed at ~3664 cm–1, indicating that all syn orientated bidentate and monodentate O2 conformers of [dThd+Na]+ do not have measurable populations in the experiments (see Supplementary Figure S7). Interestingly, the 3′-hydroxyl is red shifted to 3650 cm–1 versus the 5′-hydroxyl at 3670 cm–1 in the bidentate B1(O2O3') conformer, whereas the 3′-hydroxyl is blue shifted to 3678 cm–1 versus the 5′-hydroxyl at 3671 cm–1 in the monodentate M1(O2') conformer. A significant red shift of the strong IR feature at ~1686 cm–1 is observed for all O4 binding conformers, indicating that these conformers are not measurably populated in the experiments (see the example comparisons for both anti and syn oriented M1(O4) conformers in Supplementary Figure S8). For all sugar binding conformers, obvious spectral mismatches between the calculated IR and experimental IRMPD spectra are also found and demonstrate that Na+ binding solely to the sugar moiety is not sufficiently favorable for [dThd+Na]+ to measurably populated these species in the experiments, consistent with their computed free energies at 298 K (see Supplementary Figure S9). Likewise, neither t2 nor t4 conformers are present in the experiments because of the obvious spectral misalignments in the high end of the FELIX region and the lack of the IR feature at ~3420 cm–1 in the OPO region (see the example comparisons for the B1(O2O5')t4 and B1(N3O4)t2 conformers in Supplementary Figure S10).
Comparison of the experimental IRMPD action spectrum of [dThd+Na]+ with the B3LYP/6-311+G(d,p) optimized structures and calculated linear IR spectra for representative low-energy conformers that best match the measured spectrum. The Thy nucleobase orientation, sugar puckering, and B3LYP/6-311+G(2d,2p) relative Gibbs free energies at 298 K are also shown
In summary, the T1, T2, and T3(O2O4'O5') conformers are dominantly populated in the experiments, whereas the T4, T5, T6, and T7(O2O4'O5') conformers are not measurably populated. If the B1(O2O4') conformer is present, it has low abundance. All O4 binding and sugar binding conformers are not present in the experiments. Likewise, it is very clear that all sodium cationized tautomeric dThd are also not populated in the experiments. These findings are also consistent with the computed relative Gibbs free energies for these conformers of [dThd+Na]+.
Comparison of Experimental IRMPD and Calculated IR Spectra of [Thd+Na]+
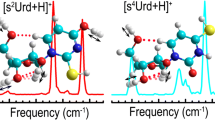
Figure 6 compares the experimental IRMPD and calculated IR spectra of [Thd+Na]+ for conformers that exhibit good agreement. Supplementary Figures S11–S15 show analogous comparisons of representative conformers of [Thd+Na]+ that exhibit spectral misalignments (shaded in red). The IR spectrum calculated for the B1(O2O2') conformer of [Thd+Na]+, which exhibits an anti orientation of the Thy nucleobase and C4'-exo (4TO) sugar puckering, exhibits good agreement with the experimental IRMPD spectrum; the slight broadening of the IR feature calculated at ~3420 cm–1 well reproduces the weak shoulder to the blue of the IR feature observed (see Figure 6). The other stable structures that exhibit a very similar conformation, i.e., B4 and B5(O2O2'), exhibit virtually identical IR spectra (not shown) and thus may also be populated in the experiments. For all T(O2O4'O5') conformers, a moderate IR feature is predicted between ~3540 and 3630 cm–1 in the calculated IR spectra, which contributes to the very weak IR absorption at ~3605 cm–1 in the experimental IRMPD spectrum of [Thd+Na]+. Among all T(O2O4'O5') conformers, the IR spectrum calculated for the T2(O2O4'O5') conformer shows the best agreement with the experimental IRMPD spectrum, but slight red and blue shifts are observed for the two intense peaks predicted at ~3420 and 3682 cm–1, respectively, suggesting that the T2(O2O4'O5') conformer of [Thd+Na]+ is a less important contributor than B1(O2O2') in the experiments (see Figure 6). For the ground T1(O2O4'O5') conformer, this IR feature shifts to the red compared with the measured IRMPD spectrum, suggesting that if present, the ground T1(O2O4'O5') conformer is likely present in low abundance in the experiments (see Supplementary Figure S11). Overall, the T(O2O4'O5') conformers have low abundances in the experiments. The calculated IR spectrum of the anti oriented tridentate O2 binding conformer, T1(O2O2'O3'), exhibits reasonably good agreement with the experimental IRMPD spectrum, where slight misalignments of the calculated IR spectrum from ~971 to 1166 cm–1 are observed, indicating that the T(O2O2'O3') conformers may also be populated in the experiments, but are probably present in lower abundance than the B1(O2O2') conformer (see Figure 6). The hydrogen-bonding interaction between the 3'- and 5'-hydroxyl substituents of the B2(O2O2') conformer produces an additional IR band at ~1166 cm–1 and a shift of the IR spectral features in the OPO region that does not agree with the IR features observed in the experiments, indicating that this conformer is not measurably populated (see Supplementary Figure S11). The B3(O2O2') conformer with C3'-exo (3T4) sugar puckering and a noncanonical hydrogen-bonding interaction between the O5' and C6H atoms leads to misalignment in the range from ~1150 to 1250 cm–1 and a further splitting of the IR feature at ~3420 cm–1, suggesting that this conformer, if present, is only present in very low population (see Supplementary Figure S11). The IR spectrum predicted for the B1(O2O4') conformer is somewhat parallel to that of the T1(O2O4'O5') conformer, but additional disagreements observed in the FELIX region indicate that all B(O2O4') conformers, if present, have even lower populations than the T(O2O4'O5') conformers (see Supplementary Figure S11). Similar to [dThd+Na]+, the IR spectra calculated for all O4 binding and sugar binding conformers exhibit obvious spectral mismatches, such that these conformers are not present in the experiments (see Supplementary Figures S12, S13, and S14). The two intense IR features in the high end of the FELIX region are associated with the C2=O and C4=O stretches, and the IR feature at ~3420 cm–1 is associated with the N3–H stretch (see Table 2). Thus, all t2 and t4 conformers where the N3 proton has shifted to the O2 or O4 atoms are not populated because the significant shifts of the IR bands associated with the C2=O, C4=O, and N3–H stretches (see the example comparisons for the B1(O2O5')t4 and B1(N3O4)t2 conformers in Supplementary Figure S15).
Comparison of the experimental IRMPD action spectrum of [Thd+Na]+ with the B3LYP/6-311+G(d,p) optimized structures and calculated linear IR spectra for representative low-energy conformers that best match the measured spectrum. The Thy nucleobase orientation, sugar puckering, and B3LYP/6-311+G(2d,2p) relative Gibbs free energies at 298 K are also shown
In summary, anti oriented bidentate O2 binding conformers of [Thd+Na]+ are dominantly populated, and anti oriented tridentate O2 binding conformers may also be populated, but have lower abundance than the B(O2O2') conformers; syn oriented tridentate and bidentate O2 binding conformers may also be minor contributors in the experiments. Kinetic trapping of B(O2O2') conformers occurs in the ESI desolvation process, consistent with the inversion of the relative stabilities of the solvated T1(O2O4'O5') and B1(O2O2') conformers. Additional hydrogen-bonding interactions in the B2(O2O2') conformer slightly shift the IR bands in both FELIX and OPO regions, suggesting that similar B(O2O2') conformers are not populated in the experiments. Minor disagreements are observed for the IR spectrum calculated for the B3(O2O2') conformer, suggesting that if present, this conformer has low abundance in the experiments. In contrast, all conformers involving O4 binding, sugar binding, and tautomeric [Thd+Na]+ are not important in the experiments. These findings are again consistent with the computed relative Gibbs free energies for the conformers of [Thd+Na]+.
Vibrational Mode Assignments for [dThd+Na]+ and [Thd+Na]+
Table 3 lists vibrational mode assignments for [dThd+Na]+ and [Thd+Na]+ based on the ground T1(O2O4'O5') conformer of [dThd+Na]+ and the dominantly populated B1(O2O2') conformer of [Thd+Na]+. In general, the IR features observed for [dThd+Na]+ and [Thd+Na]+ in the experiments are highly similar. The relatively strong IR features at ~772, 862, and 944 cm–1 associated with Thy twisting, C6–H wagging, and sugar breathing, respectively, are only observed in the measured IRMPD spectrum of [dThd+Na]+. This may be associated with inefficient dissociation of [Thd+Na]+ at these low frequencies. The intense IR feature with a shoulder to the red at ~1680 cm–1 is contributed by C5=C6 and C2=O stretches. The intense IR feature at ~1750 cm–1 is associated with the C4=O stretch. Additionally, the strong IR band at 3420 cm–1 represents N3–H stretching, which only occurs in the canonical forms of dThd and Thd. These spectral signatures provide strong evidence that all tautomeric [dThd+Na]+ and [Thd+Na]+ are not populated. In the measured IRMPD spectrum of [Thd+Na]+, the IR band at 3420 cm–1 along with the small shoulder to the blue is contributed by the N3–H and 2'-hydroxyl stretches.
Comparisons of the Experimental IRMPD Spectra of Sodium Cationized and Protonated dThd and Thd
IRMPD action spectroscopy and theoretical studies of protonated dThd and Thd were previously reported by Wu and coworkers [41]. Figure 2 shows comparisons of the experimental IRMPD spectra of sodium cationized and protonated dThd and Thd. As is obvious in the figure, the IRMPD spectra of these species exhibit significant differences. The experimental IRMPD spectra of [dThd+H]+ and [Thd+H]+ are contributed by both 2,4-dihydroxy (t24) and O2 protonated thymine nucleosides. The IR bands at ~1780 and ~3395 cm–1 are only contributed by O2 protonated conformers, whereas the IR bands at ~1210 and ~3580 cm–1 as well as the shoulder at ~3560 cm–1 are mainly contributed by the t24 conformers. The IR features measured at ~1520 and ~1495 cm–1 represent C2–O and C4–O stretches, respectively. In contrast, the two intense peaks in the high end of the FELIX region and the IR feature at ~3420 cm–1 measured for [dThd+Na]+ and [Thd+Na]+ are only contributed by the canonical 2,4-diketo forms of the Thy nucleosides. The relatively low IRMPD yields of the sodium cationized Thy nucleosides in the FELIX region suggest that sodium cationization activates the N-glycosidic bond less effectively than protonation, and that sodium cation binding to both the Thy nucleobase and sugar moiety may reduce the flexibility of the nucleoside in these complexes.
Comparisons of Stable Low-Energy Conformers of Sodium Cationized and Protonated dThd and Thd
Figure 1 also shows the ground conformers of the protonated Thy nucleosides calculated using the same theoretical approach as employed in this work [41]. For both [dThd+H]+ and [Thd+H]+, the ground conformers are 2,4-dihydroxy tautomers with anti orientations of the Thy nucleobase, where the 2- and 4-hydroxyl hydrogen atoms point toward the N3 atom, and exhibit C2'-endo sugar puckering. The most stable O2 protonated Thy nucleosides are only 4.1 and 5.2 kJ/mol less favorable than the corresponding ground conformers, respectively, whereas the most stable O4 protonated Thy nucleosides are 3.3 and 6.2 kJ/mol less stable, respectively, i.e., the protonation preferences of [dThd+H]+ follow the order t24 > O4 > O2. In contrast, the protonation preferences of [Thd+H]+ follow the order t24 > O2 > O4. In spite of the similar computed relative stabilities, evidence for O4 protonated species was not found. For the sodium cationized Thy nucleosides, the ground conformer of [dThd+Na]+ involves tridentate binding of Na+ to the canonical form of dThd via the O2, O4', and O5' atoms, with a syn orientation of the Thy nucleobase and O4'-endo (OT1) sugar puckering. The ground conformer of [Thd+Na]+ exhibits a very similar conformation, but the O4'-endo sugar puckering changes to OT4 because of the hydrogen-bonding interaction between the 2'- and 3'-hydroxyl substituents. The relative stabilities of different forms of [dThd+Na]+ follow the order O2 > O4 > t4 > sugar > t2. In contrast, the relative stabilities of different forms of [Thd+Na]+ follow the order O2 > sugar > O4 > t4 > t2. The difference in the order of the relative stabilities of [dThd+Na]+ and [Thd+Na]+ indicates that the 2'-hydroxyl substituent of Thd stabilizes the sugar binding conformers relative to dThd. Compared with protonation, sodium cationization is unable to sufficiently stabilize the minor tautomers of the Thy nucleosides, such that the canonical forms of [dThd+Na]+ and [Thd+Na]+ have lower relative Gibbs free energies and are the only forms populated in the experiments.
Survival Yield Analyses of the Sodium Cationized and Protonated Thymine Nucleosides
Figure 7 shows comparisons of the survival yield curves of the sodium cationized and protonated Thy nucleosides. Based on the CID50% values, the overall relative stabilities of these species follow the order [dThd+H]+ < [Thd+H]+ < [dThd+Na]+ < [Thd+Na]+. In particular, the dominant fragmentation pathways observed for [Thd+H]+, [dThd+Na]+, and [Thd+Na]+ involve only N-glycosidic bond cleavage. Thus, the CID50% values of these species directly correlate with the relative stabilities of the N-glycosidic bonds in these complexes. However, the CID behavior of [dThd+H]+ is much richer. Although glycosidic bond cleavage remains the dominant CID pathway (see Figure 3), a variety of water and ketene elimination reactions are also observed and become increasingly competitive with glycosidic bond cleavage, such that the survival yield falls off more rapidly and the CID50% value shifts to lower rfEA than it would be in the absence of these neutral loss pathways, and is representative of the overall stability rather than the glycosidic bond stability of [dThd+H]+. The CID50% value of [Thd+Na]+ is higher than that of [Thd+H]+, indicating that sodium cationization activates the N-glycosidic bond less effectively than protonation. Likewise, the CID50% value of [Thd+Na]+ exceeds that of [dThd+Na]+, indicating that the 2'-hydroxy modification of Thd stabilizes the N-glycosidic bond compared with dThd. These conclusions show good agreement with previous ER-CID and threshold collision-induced dissociation (TCID) studies [94,95,96,97], which all demonstrate that sodium cationization does not activate the N-glycosidic bond as effectively as protonation, and that the N-glycosidic bonds of the RNA nucleosides are stabilized by the additional 2'-hydroxyl substituent relative to the analogous DNA nucleosides.
Comparisons of the Sodium Cationized Uracil and Thymine Nucleosides
We have previously reported IRMPD action spectroscopy and ER-CID studies of the sodium cationized Ura nucleosides, [dUrd+Na]+ and [Urd+Na]+ [73]. The same theoretical approach as employed here finds that both ground conformers of [dUrd+Na]+ and [Urd+Na]+ exhibit tridentate Na+ binding to O2, O4', and O5' atoms with a syn orientation of Ura and O4'-endo sugar puckering. However, comparisons between the calculated IR and experimental IRMPD spectra of these complexes indicate that the syn oriented tridentate T(O2O4'O5') binding conformers of [dUrd+Na]+ are mainly populated, whereas the anti oriented bidentate B(O2O2') binding conformers of [Urd+Na]+ are dominantly populated in the experiments. These findings were explained by the fact that the anti oriented B(O2O2') conformers of [Urd+Na]+ are preferentially stabilized by hydration in aqueous solution. Thus, kinetic trapping of the B(O2O2') conformers during the ESI desolvation process results in these conformers being dominantly populated in the experiments. This conclusion was further validated for [Urd+Na]+ at the B3LYP/def2-TZVPPD, MP2(full)/6-311+G(2d,2p), and MP2(full)/def2-TZVPPD levels of theory [73]. Similar behavior is observed here for [dThd+Na]+ and [Thd+Na]+. Theory and experiment both suggest that the syn oriented tridentate O2 binding conformers of [dThd+Na]+ are mainly populated. In contrast, T(O2O4'O5') conformers of [Thd+Na]+ are minor contributors in the experiments, whereas the anti oriented bidentate (O2,O2') binding conformers of [Thd+Na]+ are dominantly populated in the experiments. The inversion of the order of the relative Gibbs free energies calculated for the solvated T1(O2O4'O5') and B1(O2O2') conformers suggests that B(O2O2') conformers are dominantly present in solution, and are kinetically trapped in the ESI desolvation process. Thus, the additional 5-methyl substituent does exert a minor influence on the stability of the T(O2O4'O5') versus B(O2O2') conformers.
Conclusions
DFT calculations find that the ground conformers of [dThd+Na]+ and [Thd+Na]+ exhibit tridentate Na+ binding to the O2, O4', and O5' atoms with a syn orientation of Thy and O4'-endo (OT1 for [dThd+Na]+ versus OT4 for [Thd+Na]+) sugar puckering. Comparisons of the experimental IRMPD and the theoretical IR spectra of [dThd+Na]+ and [Thd+Na]+ indicate that the sodium cation shows a preference for binding to both the Thy nucleobase and sugar moiety. The T1, T2, and T3(O2O4'O5') conformers of [dThd+Na]+ are dominantly populated in the experiments. In contrast, T(O2O4'O5') conformers of [Thd+Na]+ are minor contributors. Anti oriented B(O2O2') conformers of [Thd+Na]+ are stabilized by hydration in aqueous solution, and are kinetically trapped in the ESI desolvation process such that they are dominantly populated in the experiments. Anti oriented T(O2O2'O3') conformers may also be populated, but if so they are in lower abundance. All O4 and sugar binding conformers of [dThd+Na]+ and [Thd+Na]+ are not present in the experiments as evidenced by the significant differences between the calculated IR and experimental IRMPD spectra. Absence of the IR features associated with the 2-hydroxyl or 4-hydroxyl substituents demonstrate that all tautomeric [dThd+Na]+ and [Thd+Na]+ are also not important contributors to the experiments. The overall stabilities of protonated and sodium cationized dThd and Thd determined by survival yield analyses follow the order [dThd+H]+ < [Thd+H]+ < [dThd+Na]+ < [Thd+Na]+. Because N-glycosidic bond cleavages are the major fragmentation pathways observed for [Thd+H]+, [dThd+Na]+, and [Thd+Na]+, the relative N-glycosidic bond stabilities also follow the same order [Thd+H]+ < [dThd+Na]+ < [Thd+Na]+. These results indicate that the 2'-hydroxyl substituent stabilizes the N-glycosidic bond of Thd versus that of dThd, and that sodium cationization activates the N-glycosidic bond less effectively than protonation. By comparing the gas-phase conformations and energetics of sodium cationized Ura and Thy nucleosides, it is clear that the additional 5-methyl substituent exerts only a minor influence on the relative stabilities of the T(O2O4'O5') and B(O2O2') modes of binding.
References
van Dam, L., Ouwerkerk, N., Brinkmann, A., Raap, J., Levitt, M.H.: Solid-state NMR determination of sugar ring pucker in C-13-labeled 2'-deoxynucleosides. Biophys. J. 83, 2835–2844 (2002)
Rich, A., Nordheim, A., Wang, A.H.J.: The chemistry and biology of left-handed Z-DNA. Annu. Rev. Biochem. 53, 791–846 (1984)
McCullough, A.K., Dodson, M.L., Lloyd, R.S.: Initiation of base excision repair: glycosylase mechanisms and structures. Annu. Rev. Biochem. 68, 255–285 (1999)
Krokan, H.E., Standal, R., Slupphaug, G.: DNA glycosylases in the base excision repair of DNA. Biochem. J. 325, 1–16 (1997)
Roberts, R.J.: On base flipping. Cell 82, 9–12 (1995)
Kohwi, Y., Kohwishigematsu, T.: Magnesium ion-dependent triple-helix structure formed by homopurine-homopyrimidine sequences in supercoiled plasmid DNA. Proc. Natl. Acad. Sci. USA 85, 3781–3785 (1988)
Day, H.A., Pavlou, P., Waller, Z.A.E.: i-Motif DNA: structure, stability and targeting with ligands. Bioorgan. Med. Chem. 22, 4407–4418 (2014)
Eichhorn, G.L.: Conformational change induced by metal-ions through coordination. Coord. Chem. Rev. 128, 167–173 (1993)
Sen, D., Gilbert, W.: Formation of parallel 4-stranded complexes by guanine-rich motifs in DNA and its Implications for meiosis. Nature 334, 364–366 (1988)
Sundquist, W.I., Klug, A.: Telomeric DNA dimerizes by formation of guanine tetrads between hairpin loops. Nature 342, 825–829 (1989)
Parkinson, G.N., Lee, M.P.H., Neidle, S.: Crystal structure of parallel quadruplexes from human telomeric DNA. Nature 417, 876–880 (2002)
Burge, S., Parkinson, G.N., Hazel, P., Todd, A.K., Neidle, S.: Quadruplex DNA: sequence, topology, and structure. Nucleic Acids Res. 34, 5402–5415 (2006)
Saenger, W.: Principles of nucleic-acid structure, pp. 51–104. Springer-Verlag, New York (1984)
Shin, Y.A., Eichhorn, G.L.: Reversible change in Ψ Structure of DNA-poly(lys)complexes induced by metal-binding. Biopolymers 16, 225–230 (1977)
Stivers, J.T., Jiang, Y.L.: A mechanistic perspective on the chemistry of DNA repair glycosylases. Chem. Rev. 103, 2729–2759 (2003)
O'Brien, P.J.: Catalytic promiscuity and the divergent evolution of DNA repair enzymes. Chem. Rev. 106, 720–752 (2006)
Moody, E.M., Lecomte, J.T.J., Bevilacqua, P.C.: Linkage between proton binding and folding in RNA: a thermodynamic framework and its experimental application for investigating pK(a) shifting. RNA 11, 157–172 (2005)
Srivastava, D.K., Vande Berg, B.J., Prasad, R., Molina, J.T., Beard, W.A., Tomkinson, A.E., Wilson, S.H.: Mammalian abasic site base excision repair-identification of the reaction sequence and rate-determining steps. J. Biol. Chem. 273, 21203–21209 (1998)
Dianov, G., Lindahl, T.: Reconstitution of the DNA-base excision-repair pathway. Curr. Biol. 4, 1069–1076 (1994)
Zheng, Y., Xue, Y., Yan, S.G.: The effects of oxidation and protonation on the N-glycosidic bond stability of 8-oxo-2'-deoxyguanosine: DFT study. J. Mol. Struc.-Theochem. 860, 52–57 (2008)
Margison, G.P., Oconnor, P.J.: Biological implications of instability of N-glycosidic bond of 3-methyldeoxyadenosine in DNA. Biochim. Biophys. Acta 331, 349–356 (1973)
Das, R.S., Samaraweera, M., Morton, M., Gascon, J.A., Basu, A.K.: Stability of N-glycosidic bond of (5'S)-8,5'-cyclo-2'-deoxyguanosine. Chem. Res. Toxicol. 25, 2451–2461 (2012)
Tehrani, Z.A., Fattahi, A., Pourjavadi, A.: DFT Study of the interaction of cytidine and 2'-deoxycytidine with Li+, Na+, and K+: effects of metal cationization on sugar puckering and stability of the N-glycosidic bond. Carbohydr. Res. 344, 771–778 (2009)
Tehrani, Z.A., Fattahi, A., Pourjavadi, A.: Interaction of Mg2+, Ca2+, Zn2+, and Cu+ with cytosine nucleosides: influence of metal on sugar puckering and stability of N-glycosidic bond, a DFT study. J. Mol. Struct.-Theochem. 913, 117–125 (2009)
Rios-Font, R., Bertran, J., Rodriguez-Santiago, L., Sodupe, M.: Effects of ionization, metal cationization, and protonation on 2'-deoxyguanosine: changes on sugar puckering and stability of the N-glycosidic bond. J. Phys. Chem. B 110, 5767–5772 (2006)
Kang, H., Lee, K.T., Jung, B., Ko, Y.J., Kim, S.K.: Intrinsic lifetimes of the excited state of DNA and RNA bases. J. Am. Chem. Soc. 124, 12958–12959 (2002)
Cadet, J., Sage, E., Douki, T.: Ultraviolet radiation-mediated damage to cellular DNA. Mutat. Res. Fundam. Mol. Mech. Mutagen. 571, 3–17 (2005)
Kim, S.T., Sancar, A.: Effect of base, pentose, and phosphodiester backbone structures on binding and repair of pyrimidine dimers by Escherichia coli DNA photolyase. Biochemistry-US 30, 8623–8630 (1991)
Nosenko, Y., Kunitski, M., Brutschy, B.: Specific photodynamics in thymine clusters: the role of hydrogen bonding. J. Phys. Chem. A 115, 9429–9439 (2011)
Harris, V.H., Smith, C.L., Cummins, W.J., Hamilton, A.L., Adams, H., Dickman, M., Hornby, D.P., Williams, D.M.: The effect of tautomeric constant on the specificity of nucleotide incorporation during DNA replication: support for the rare tautomer hypothesis of substitution mutagenesis. J. Mol. Biol. 326, 1389–1401 (2003)
Alyoubi, A.O., Hilal, R.H.: A theoretical and experimental investigation of the electronic-spectra and tautomerization of nucleobases. Biophys. Chem. 55, 231–237 (1995)
Colarusso, P., Zhang, K.Q., Guo, B.J., Bernath, P.F.: The infrared spectra of uracil, thymine, and adenine in the gas phase. Chem. Phys. Lett. 269, 39–48 (1997)
Zhang, S.L., Michaelian, K.H., Loppnow, G.R.: Vibrational spectra and experimental assignments of thymine and nine of its isotopomers. J. Phys. Chem. A 102, 461–470 (1998)
Salpin, J.Y., Guillaumont, S., Tortajada, J., MacAleese, L., Lemaire, J., Maitre, P.: Infrared spectra of protonated uracil, thymine and cytosine. ChemPhysChem 8, 2235–2244 (2007)
Rodgers, M.T., Armentrout, P.B.: Noncovalent interactions of nucleic acid bases (uracil, thymine, and adenine) with alkali metal ions. Threshold collision-induced dissociation and theoretical studies. J. Am. Chem. Soc. 122, 8548–8558 (2000)
Kabelac, M., Hobza, P.: Na+, Mg2+, and Zn2+ binding to all tautomers of adenine, cytosine, and thymine and the eight most stable keto/enol tautomers of guanine: a correlated ab initio quantum chemical study. J. Phys. Chem. B 110, 14515–14523 (2006)
Zamir, A., Holley, R.W., Marquise, M.: Evidence for occurrence of a common pentanucleotide sequence in structures of transfer ribonucleic acids. J. Biol. Chem. 240, 1267–1273 (1965)
Bjork, G.R., Neidhardt, F.C.: Physiological and biochemical studies on function of 5-methyluridine in transfer ribonucleic-acid of Escherichia coli. J. Bacteriol. 124, 99–111 (1975)
Chen, B.C., Quinlan, S.L., Reid, J.G.: A new synthesis of the anti-aids drug AZT from 5-methyluridine. Tetrahedron Lett. 36, 7961–7964 (1995)
Chen, B.C., Quinlan, S.L., Stark, D.R., Reid, J.G., Audia, V.H., George, J.G., Eisenreich, E., Brundidge, S.P., Racha, S., Spector, R.H.: 5'-Benzoyl-2'alpha-bromo-3'-O-methanesulfonylthymidine: a superior nucleoside for the synthesis of the anti-aids drug D4T (stavudine). Tetrahedron Lett. 36, 7957–7960 (1995)
Wu, R.R., Yang, B., Frieler, C.E., Berden, G., Oomens, J., Rodgers, M.T.: 2,4-Dihydroxy and O2 protonated tautomers of dThd and Thd coexist in the gas phase: methylation alters protonation preferences versus dUrd and Urd. J. Am. Soc. Mass Spectrom. 27, 410–421 (2016)
Salpin, J.Y., Scuderi, D.: Structure of protonated thymidine characterized by infrared multiple photon dissociation and quantum calculations. Rapid Commun. Mass Spectrom. 29, 1898–1904 (2015)
Nei, Y.-w., Akinyemi, T.E., Steill, J.D., Oomens, J., Rodgers, M.T.: Infrared multiple photon dissociation action spectroscopy of protonated uracil and thiouracils: effects of thioketo-substitution on gas-phase conformation. Int. J. Mass Spectrom. 297, 139–151 (2010)
Nei, Y.-w., Akinyemi, T.E., Kaczan, C.M., Steill, J.D., Berden, G., Oomens, J., Rodgers, M.T.: Infrared multiple photon dissociation action spectroscopy of sodiated uracil and thiouracils: effects of thioketo-substitution on gas-phase conformation. Int. J. Mass Spectrom. 308, 191–202 (2011)
Crampton, K.T., Rathur, A.I., Nei, Y.-w., Berden, G., Oomens, J., Rodgers, M.T.: Protonation preferentially stabilizes minor tautomers of the halouracils: IRMPD action spectroscopy and theoretical studies. J. Am. Soc. Mass Spectrom. 23, 1469–1478 (2012)
Kaczan, C.M., Rathur, A.I., Wu, R.R., Chen, Y., Austin, C.A., Berden, G., Oomens, J., Rodgers, M.T.: Infrared multiple photon dissociation action spectroscopy of sodium cationized halouracils: effects of sodium cationization and halogenation on gas-phase conformation. Int. J. Mass Spectrom. 378, 76–85 (2015)
Oomens, J., Moehlig, A.R., Morton, T.H.: Infrared multiple photon dissociation (IRMPD) spectroscopy of the proton-bound dimer of 1-methylcytosine in the gas phase. J. Phys. Chem. Lett. 1, 2891–2897 (2010)
Yang, B., Wu, R.R., Berden, G., Oomens, J., Rodgers, M.T.: Infrared multiple photon dissociation action spectroscopy of proton-bound dimers of cytosine and modified cytosines: effects of modifications on gas-phase conformations. J. Phys. Chem. B 117, 14191–14201 (2013)
Salpin, J.Y., Haldys, V., Guillaumont, S., Tortajada, J., Hurtado, M., Lamsabhi, A.: Gas-phase interactions between lead(II) ions and cytosine: tandem mass spectrometry and infrared multiple-photon dissociation spectroscopy study. ChemPhysChem 15, 2959–2971 (2014)
Yang, B., Wu, R.R., Polfer, N.C., Berden, G., Oomens, J., Rodgers, M.T.: IRMPD action spectroscopy of alkali metal cation-cytosine complexes: effects of alkali metal cation size on gas phase conformation. J. Am. Soc. Mass Spectrom. 24, 1523–1533 (2013)
Rajabi, K., Gillis, E.A.L., Fridgen, T.D.: Structures of alkali metal ion-adenine complexes and hydrated complexes by IRMPD spectroscopy and electronic structure calculations. J. Phys. Chem. A 114, 3449–3456 (2010)
Rajabi, K., Theel, K., Gillis, E.A.L., Beran, G., Fridgen, T.D.: The structure of the protonated adenine dimer by infrared multiple photon dissociation spectroscopy and electronic structure calculations. J. Phys. Chem. A 113, 8099–8107 (2009)
Gillis, E.A.L., Demireva, M., Nanda, K., Beran, G., Williams, E.R., Fridgen, T.D.: Structures and energetics of electrosprayed uracil(n)Ca(2+) clusters (n = 14-4) in the gas phase. Phys. Chem., Chem. Phys 14, 3304–3315 (2012)
Power, B., Haldys, V., Salpin, J.Y., Fridgen, T.D.: Structures of [M(Ura-H)(H2O)(n)](+) (M = Mg, Ca, Sr, Ba; n = 1–3) complexes in the gas phase by IRMPD spectroscopy and theoretical studies. J. Mass Spectrom. 51, 236–244 (2016)
Ali, O.Y., Fridgen, T.D.: Structures and fragmentation of [Cu(Uracil-H)(Uracil)]+ in the gas phase. ChemPhysChem 13, 588–596 (2012)
Ali, O.Y., Randell, N.M., Fridgen, T.D.: Primary fragmentation pathways of gas phase [M(Uracil-H)(Uracil)]+ complexes (M = Zn, Cu, Ni, Co, Fe, Mn, Cd, Pd, Mg, Ca, Sr, Ba, and Pb): loss of uracil versus HNCO. ChemPhysChem. 13, 1507–1513 (2012)
Power, B., Rowe, S., Fridgen, T.D.: Ammoniated complexes of uracil and transition metal ions: structures of [M(Ura-H)(Ura)(NH3)](+) by IRMPD spectroscopy and computational methods (M = Fe, Co, Ni, Cu, Zn, Cd). J. Phys. Chem. B 121, 58–65 (2017)
Azargun, M., Jami-Alahmadi, Y., Fridgen, T.D.: The intrinsic stabilities and structures of alkali metal cationized guanine quadruplexes. Phys. Chem., Chem. Phys. 19, 1281–1287 (2017)
Azargun, M., Fridgen, T.D.: Guanine tetrads: an IRMPD spectroscopy, energy resolved SORI-CID, and computational study of M(9-Ethylguanine)(4)(+) (M = Li, Na, K, Rb, Cs) in the gas phase. Phys. Chem., Chem. Phys. 17, 25778–25785 (2015)
Gillis, E.A.L., Rajabi, K., Fridgen, T.D.: Structures of hydrated Li+-thymine and Li+-uracil complexes by IRMPD spectroscopy in the N-H/O-H stretching region. J. Phys. Chem. A 113, 824–832 (2009)
Bakker, J.M., Sinha, R.K., Besson, T., Brugnara, M., Tosi, P., Salpin, J.Y., Maitre, P.: Tautomerism of uracil probed via infrared spectroscopy of singly hydrated protonated uracil. J. Phys. Chem. A 112, 12393–12400 (2008)
van Zundert, G.C.P., Jaeqx, S., Berden, G., Bakker, J.M., Kleinermanns, K., Oomens, J., Rijs, A.M.: IR spectroscopy of isolated neutral and protonated adenine and 9-methyladenine. ChemPhysChem 12, 1921–1927 (2011)
Bakker, J.M., Salpin, J.Y., Maitre, P.: Tautomerism of cytosine probed by gas phase IR spectroscopy. Int. J. Mass Spectrom. 283, 214–221 (2009)
Wu, R.R., Yang, B., Frieler, C.E., Berden, G., Oomens, J., Rodgers, M.T.: Diverse mixtures of 2,4-dihydroxy tautomers and O4 protonated conformers of uridine and 2'-deoxyuridine coexist in the gas phase. Phys. Chem., Chem. Phys. 17, 25978–25988 (2015)
Wu, R.R., Yang, B., Berden, G., Oomens, J., Rodgers, M.T.: Gas-phase conformations and energetics of protonated 2'-deoxyguanosine and guanosine: IRMPD action spectroscopy and theoretical studies. J. Phys. Chem. B 118, 14774–14784 (2014)
Wu, R.R., Yang, B., Berden, G., Oomens, J., Rodgers, M.T.: Gas-phase conformations and energetics of protonated 2'-deoxyadenosine and adenosine: IRMPD action spectroscopy and theoretical studies. J. Phys. Chem. B 119, 2795–2805 (2015)
Wu, R.R., Yang, B., Frieler, C.E., Berden, G., Oomens, J., Rodgers, M.T.: N3 and O2 protonated tautomeric conformations of 2'-deoxycytidine and cytidine coexist in the gas phase. J. Phys. Chem. B 119, 5773–5784 (2015)
Ung, H.U., Huynh, K.T., Poutsma, J.C., Oomens, J., Berden, G., Morton, T.H.: Investigation of proton affinities and gas phase vibrational spectra of protonated nucleosides, deoxynucleosides, and their analogs. Int. J. Mass Spectrom. 378, 294–302 (2015)
Filippi, A., Fraschetti, C., Rondino, F., Piccirillo, S., Steinmetz, V., Guidoni, L., Speranza, M.: Protonated pyrimidine nucleosides probed by IRMPD spectroscopy. Int. J. Mass Spectrom. 354, 54–61 (2013)
Zhu, Y., Hamlow, L.A., He, C.C., Strobehn, S.F., Lee, J.K., Gao, J., Berden, G., Oomens, J., Rodgers, M.T.: Influence of sodium cationization versus protonation on the gas-phase conformations and glycosidic bond stabilities of 2'-deoxyadenosine and adenosine. J. Phys. Chem. B 120, 8892–8904 (2016)
Zhu, Y., Hamlow, L.A., He, C.C., Lee, J.K., Gao, J., Berden, G., Oomens, J., Rodgers, M.T.: Gas-phase conformations and N-glycosidic bond stabilities of sodium cationized 2'-deoxyguanosine and guanosine: sodium cations preferentially bind to the guanine residue. J. Phys. Chem. B 121, 4048–4060 (2017)
Zhu, Y., Hamlow, L.A., He, C.C., Roy, H.A., Cunningham, N.A., Munshi, M.U., Berden, G., Oomens, J., Rodgers, M.T.: Conformations and N-glycosidic bond stabilities of sodium cationized 2'-deoxycytidine and cytidine: solution conformation of [Cyd+Na]+ is preserved upon ESI. Int. J. Mass Spectrom. (2017). doi:10.1016/j.ijms.2017.04.005
Zhu, Y., Roy, H.A., Cunningham, N.A., Strobehn, S.F., Gao, J., Munshi, M.U., Berden, G., Oomens, J., Rodgers, M.T.: Effects of sodium cationization versus protonation on the conformations and N-glycosidic bond stabilities of sodium cationized uridine and 2'-deoxyuridine: solution conformation of [Urd+Na]+ is preserved upon ESI. Phys. Chem. Chem. Phys. 19 17637–17652 (2017)
Nei, Y.-w., Hallowita, N., Steill, J.D., Oomens, J., Rodgers, M.T.: Infrared multiple photon dissociation action spectroscopy of deprotonated DNA mononucleotides: gas-phase conformations and energetics. J. Phys. Chem. A 117, 1319–1335 (2013)
Nei, Y.-w., Crampton, K.T., Berden, G., Oomens, J., Rodgers, M.T.: Infrared multiple photon dissociation action spectroscopy of deprotonated RNA mononucleotides: gas-phase conformations and energetics. J. Phys. Chem. A 117, 10634–10649 (2013)
Wu, R.R., He, C.C., Hamlow, L.A., Nei, Y.-w., Berden, G., Oomens, J., Rodgers, M.T.: Protonation induces base rotation of purine nucleotides pdGuo and pGuo. Phys. Chem., Chem. Phys. 18, 15081–15090 (2016)
Wu, R.R., He, C.C., Hamlow, L.A., Nei, Y.-w., Berden, G., Oomens, J., Rodgers, M.T.: N3 protonation induces base rotation of 2'-deoxyadenosine-5'-monophosphate and adenosine-5'-monophosphate. J. Phys. Chem. B 120, 4616–4624 (2016)
Chiavarino, B., Crestoni, M.E., Fornarini, S., Lanucara, F., Lemaire, J., Maitre, P., Scuderi, D.: Infrared spectroscopy of isolated nucleotides. 1. The cyclic 3',5'-adenosine monophosphate anion. Int. J. Mass Spectrom. 270, 111–117 (2008)
Lanucara, F., Crestoni, M.E., Chiavarino, B., Fornarini, S., Hernandez, O., Scuderi, D., Maitre, P.: Infrared spectroscopy of nucleotides in the gas phase. 2. The protonated cyclic 3',5'-adenosine monophosphate. Rsc. Adv. 3, 12711–12720 (2013)
Ligare, M.R., Rijs, A.M., Berden, G., Kabelac, M., Nachtigallova, D., Oomens, J., de Vries, M.S.: Resonant infrared multiple photon dissociation spectroscopy of anionic nucleotide monophosphate clusters. J. Phys. Chem. B 119, 7894–7901 (2015)
Salpin, J.Y., MacAleese, L., Chirot, F., Dugourd, P.: Structure of the Pb2+-deprotonated dGMP complex in the gas phase: a combined MS-MS/IRMPD spectroscopy/ion mobility study. Phys. Chem., Chem. Phys. 16, 14127–14138 (2014)
Wu, R.R., Hamlow, L.A., He, C.C., Nei, Y.-w., Berden, G., Oomens, J., Rodgers, M.T.: N3 and O2 protonated conformers of the cytosine mononucleotides coexist in the gas phase. J. Am. Soc. Mass Spectrom. 28, 1638–1646 (2017)
Valle, J.J., Eyler, J.R., Oomens, J., Moore, D.T., van der Meer, A.F.G., von Helden, G., Meijer, G., Hendrickson, C.L., Marshall, A.G., Blakney, G.T.: Free electron laser-fourier transform ion cyclotron resonance mass spectrometry facility for obtaining infrared multiphoton dissociation spectra of gaseous ions. Rev. Sci. Instrum. 76, 023103 (2005)
Polfer, N.C., Oomens, J., Moore, D.T., von Helden, G., Meijer, G., Dunbar, R.C.: Infrared spectroscopy of phenylalanine Ag(I) and Zn(II) complexes in the gas phase. J. Am. Chem. Soc. 128, 517–525 (2006)
Polfer, N.C., Oomens, J.: Reaction products in mass spectrometry elucidated with infrared spectroscopy. Phys. Chem., Chem. Phys. 9, 3804–3817 (2007)
Oepts, D., Vandermeer, A.F.G., Vanamersfoort, P.W.: The free-electron-laser user facility FELIX. Infrared Phys. Techn. 36, 297–308 (1995)
Wolinski, K., Hinton, J.F., Wishart, D.S., Sykes, B.D., Richards, F.M., Pastone, A., Saudek, V., Ellis, P.D., Maciel, G.E., McIver, J.W., Blizzard, A.C., Santry, D.P., Pople, J.A., Ostlund, N.S., Ducasse, L., Hoarau, J., Pesquer, M., Kondo, M., Ando, I., Chujo, R., Nishioka, A., Vauthier, E.C., Odiot, S., Tonnard, F., Baker, J.D., Zerner, M.C., Beveridge, D.V., Anderson, W.P., Cundari, T.R., Bingham, R.C., Dewar, M.J.S., Lo, D.H., Li, J., Mello, P.C., Jug, K., Tihiel, W., Zoebisch, E.G., Healy, E.F., Stewart, J.J.P., Fraser, M., Hayes, D.M.; HyperChem Computational Chemistry Software Package; ver. 8.0: Hypercube, Inc., Gainesville, FL, (2004)
Frisch, M.J., Trucks, G.W., Schlegel, H.B., Scuseria, G.E., Robb, M.A., Cheeseman, J.R., Scalmani, G., Barone, V., Mennucci, B., Petersson, G.A., Nakatsuji, H., Caricato, M., Li, X., Hratchian, H.P., Izmaylov, A.F., Bloino, J., Zheng, G., Sonnenberg, J.L., Hada, M., Ehara, M., Toyota, K., Fukuda, R., Hasegawa, J., Ishida, M., Nakajima, T., Honda, Y., Kitao, O., Nakai, H., Vreven, T., Montgomery J.A. Jr., Peralta, J.E., Ogliaro, F., Bearpark, M.J., Heyd, J., Brothers, E.N., Kudin, K.N., Staroverov, V.N., Kobayashi, R., Normand, J., Raghavachari, K., Rendell, A.P., Burant, J.C., Iyengar, S.S., Tomasi, J., Cossi, M., Rega, N., Millam, N.J., Klene, M., Knox, J.E., Cross, J.B., Bakken, V., Adamo, C., Jaramillo, J., Gomperts, R., Stratmann, R.E., Yazyev, O., Austin, A.J., Cammi, R., Pomelli, C., Ochterski, J.W., Martin, R.L., Morokuma, K., Zakrzewski, V.G., Voth, G.A., Salvador, P., Dannenberg, J.J., Dapprich, S., Daniels, A.D., Farkas, Ö., Foresman, J.B., Ortiz, J.V., Cioslowski, J., Fox, D.J.; Gaussian 09, revision C.01: Gaussian, Inc., Wallingford, CT, (2009)
Memboeuf, A., Nasioudis, A., Indelicato, S., Pollreisz, F., Kuki, A., Keki, S., van den Brink, O.F., Vekey, K., Drahos, L.: Size effect on fragmentation in tandem mass spectrometry. Anal. Chem. 82, 2294–2302 (2010)
Derwa, F., Depauw, E., Natalis, P.: New basis for a method for the estimation of secondary ion internal energy-distribution in soft ionization techniques. Org. Mass Spectrom. 26, 117–118 (1991)
Guo, X.H., Duursma, M.C., Kistemaker, P.G., Nibbering, N.M.M., Vekey, K., Drahos, L., Heeren, R.M.A.: Manipulating internal energy of protonated biomolecules in electrospray ionization Fourier transform ion cyclotron resonance mass spectrometry. J. Mass Spectrom. 38, 597–606 (2003)
Memboeuf, A., Jullien, L., Lartia, R., Brasme, B., Gimbert, Y.: Tandem mass spectrometric analysis of a mixture of isobars using the survival yield technique. J. Am. Soc. Mass Spectrom. 22, 1744–1752 (2011)
Kertesz, T.M., Hall, L.H., Hill, D.W., Grant, D.F.: CE50: Quantifying collision induced dissociation energy for small molecule characterization and identification. J. Am. Soc. Mass Spectrom. 20, 1759–1767 (2009)
Wu, R.R., Chen, Y., Rodgers, M.T.: Mechanisms and energetics for N-glycosidic bond cleavage of protonated 2'-deoxyguanosine and guanosine. Phys. Chem., Chem. Phys. 18, 2968–2980 (2016)
Wu, R.R., Rodgers, M.T.: Tautomerization lowers the activation barriers for N-glycosidic bond cleavage of protonated uridine and 2'-deoxyuridine. Phys. Chem., Chem. Phys. 18, 24451–24459 (2016)
Wu, R.R., Rodgers, M.T.: Mechanisms and energetics for N-glycosidic bond cleavage of protonated adenine nucleosides: N3 protonation induces base rotation and enhances N-glycosidic bond stability. Phys. Chem., Chem. Phys. 18, 16021–16032 (2016)
Wu, R.R., Rodgers, M.T.: O2 protonation controls threshold behavior for N-glycosidic bond cleavage of protonated cytosine nucleosides. J. Phys. Chem. B 120, 4803–4811 (2016)
Acknowledgements
This work was financially supported by the National Science Foundation, grants OISE-0730072 and OISE-1357887 (for the IRMPD measurements and international travel), DBI-0922819 (for the Bruker amaZon ETD QITMS employed in this work), and CHE-1409420 (for other research costs). Y.Z. gratefully acknowledges support from Wayne State University Thomas C. Rumble Graduate Fellowship. S.F.S. acknowledges support as an MSPIRE REU.
The authors also thank Wayne State University C&IT for computational resources and support. This work is part of the research program of FOM, which is financially supported by the Nederlandse Organisatie voor Wetenschappelijk Onderzoek (NWO). The skillful assistance of the FELIX staff is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 6.63 MB)
Rights and permissions
About this article
Cite this article
Zhu, Y., Roy, H.A., Cunningham, N.A. et al. IRMPD Action Spectroscopy, ER-CID Experiments, and Theoretical Studies of Sodium Cationized Thymidine and 5-Methyluridine: Kinetic Trapping During the ESI Desolvation Process Preserves the Solution Structure of [Thd+Na]+ . J. Am. Soc. Mass Spectrom. 28, 2423–2437 (2017). https://doi.org/10.1007/s13361-017-1753-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-017-1753-5