Abstract
Electrospray ionization mass spectrometry (ESI-MS) binding studies between proteins and ligands under native conditions require that instrumental ESI source conditions are optimized if relative solution-phase equilibrium concentrations between the protein–ligand complex and free protein are to be retained. Instrumental ESI source conditions that simultaneously maximize the relative ionization efficiency of the protein–ligand complex over free protein and minimize the protein–ligand complex dissociation during the ESI process and the transfer from atmospheric pressure to vacuum are generally specific for each protein–ligand system and should be established when an accurate equilibrium dissociation constant (KD) is to be determined via titration. In this paper, a straightforward and systematic approach for ESI source optimization is presented. The method uses statistical design of experiments (DOE) in conjunction with response surface methodology (RSM) and is demonstrated for the complexes between Plasmodium vivax guanylate kinase (PvGK) and two ligands: 5′-guanosine monophosphate (GMP) and 5′-guanosine diphosphate (GDP). It was verified that even though the ligands are structurally similar, the most appropriate ESI conditions for KD determination by titration are different for each.

ᅟ
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Binding interactions between ligands and proteins are crucial for a wide variety of biological processes and for the effectiveness of many therapeutic compounds. Among the methods available for the identification and characterization of noncovalent interactions between proteins and their ligands, electrospray ionization mass spectrometry (ESI-MS) offers advantages of high sensitivity, selectivity, simplicity, and speed [1–4]. With ESI-MS, native proteins and protein–ligand complexes can be transferred from solution to the gas phase as ions without changing their relative equilibrium concentrations [5, 6]. ESI-MS can probe specificity and stoichiometry of binding [7–10], protein conformational changes [11, 12], kinetics [13, 14], thermodynamics of binding [15, 16], and quantify equilibrium dissociation constants (KD) [17–25]. However, before establishing any correlation between the relative abundance of the gas-phase ions in the mass spectra and relative concentrations in solution, differences in ionization and detection efficiencies (response factors) between the free protein and protein–ligand complex, as well as processes such as nonspecific ligand binding and protein–ligand complex dissociation, need to be considered and, if possible, minimized.
To take into account any differences in ionization and detection efficiencies between the free protein and the protein–ligand complex, several methods that quantify their response factors (R) have been proposed [26–29]. The charge residue model (CRM) explains that high concentrations of ligand in the final droplet can lead to nonspecific interactions between free ligand and free protein, resulting in formation of nonspecific protein–ligand complexes on droplet evaporation. These complexes are indistinguishable from specific complexes originating from solution in the mass spectra [6, 30]. Based on the observation that nonspecific ligand binding is independent of the size and structure of the protein and follows a random distribution (Poisson distribution) [30–34], different strategies have been developed to correct the ESI mass spectra for the occurrence of nonspecific ligand binding [33–38].
Although most reports of KD determination by ESI-MS recognize the importance of adjusting the instrumental source parameters to preserve the protein–ligand complexes formed in solution [39–43], none has provided a straightforward statistical approach for successful systematic ESI source optimization. Statistical design of experiments (DOE) and response surface methodology (RSM) have previously been shown to be useful to optimize signal sensitivity [44–49]. We have, for example, previously reported the use of a three-level fractional factorial design to maximize the absolute intensity of the noncovalent complex bCAII-ethoxzolamide and thus find the best conditions for screening natural product extracts against this protein [47].
Optimization of the ESI source was performed for the interactions between Plasmodium vivax guanylate kinase (PvGK) and its natural substrates 5′-guanosine monophosphate (GMP) and 5′-guanosine diphosphate (GDP). The effects of determined preselected ESI source parameters on relative ion abundances, defined as the ratio of protein–ligand complex (1:1 stoichiometry) to free protein ion abundances (PL/P), were evaluated in inscribed central composite designs (CCIs). MS response analysis by RSM allowed the establishment of optimal ESI source conditions for KD determination by titration, which was validated by competition experiments and literature KD values.
Experimental
Materials
GMP and GDP were purchased from Sigma-Aldrich (Castle Hill, NSW, Australia). Ammonium acetate was purchased from Fluka (Castle Hill, NSW, Australia). GMP and GDP stock solutions were prepared in Milli-Q water (Millipore, North Ryde, NSW, Australia). For protein buffer exchange, Nalgene NAP-5 size G25, from GE Healthcare, (Parramatta, NSW, Australia) were used (purchased through Thermo Fisher Scientific Australia Pty Ltd, Scoresby, VIC, Australia).
Protein Production
Full-length guanylate kinase with an N-terminal 6-histidine tag (23545 Da) was cloned, produced, and purified as previously described from Plasmodium vivax cDNA [50].
ESI Source and FT-ICR Mass Spectrometry
All experiments were performed on a Bruker APEX III 4.7 T FT-ICR mass spectrometer equipped with an external Apollo ESI source (Supplementary Figure S1). The 14 experimental ESI source parameters that can be tuned are: sample flow rate, nebulizer gas (N2) pressure, end plate voltage, capillary voltage, drying gas (N2) flow rate, drying gas temperature, capillary exit voltage, skimmer 1 and skimmer 2 voltages, trapping and extracting voltages, and hexapole rf amplitude, hexapole DC offset voltage, and hexapole accumulation time (Supplementary Figure S1). All sample solutions were injected manually using a Cole-Parmer syringe pump. The nebulizer was off-axis and grounded. The glass capillary was 15 cm in length and had an internal diameter of 0.5 mm. Between the capillary exit and the skimmer 1, the background pressure was ~2 mbar. The ESI source housing, to where the ions are released after being accumulated inside the hexapole during a predetermined time, had a pressure of ~2 × 10–6 mbar. The analyzer was an Infinity cell.
PvGK protein was buffer exchanged with 10 mM ammonium acetate buffer (pH 6.8) and stored in aliquots at –28 °C until needed. For optimization of the ESI source by DOE and RSM, working solutions of PvGK:GMP (2:4.8 μM) and PvGK:GDP (2:2.4 μM) were prepared prior to MS analysis by appropriately diluting the protein (thawed unassisted at room temperature) and the ligand in 10 mM ammonium acetate buffer (pH 6.8). For titration experiments, samples with a fixed concentration of PvGK (2 μM) and varying concentrations of ligand (0–28.8 μM of GMP and 0–7.2 μM of GDP) were prepared. For competition experiments, samples with a fixed concentration of PvGK and GMP (2 μM and 9.6 μM, respectively) and varying concentrations of GDP (0–9.6 μM) were prepared. Incubation time was 1 h at room temperature (23–25 °C).
The data were acquired in Bruker’s Xmass software. All spectra were recorded in positive ion mode, with a sum of 32 scans per acquisition and 512 k data points per transient. The obtained spectra were analyzed in mMass 5.5.0. software. The protein–ligand complex and free protein ion abundances were calculated as the sum of all charge states intensity peaks (I) normalized for charge state (n) (∑ I(PL) n+ /n and ∑ I (P) n+ /n, respectively).
Inscribed Central Composite Designs (CCI) for ESI Source Optimization
Optimization of the ESI source was carried out using central composite designs (CCD). Inscribed central composite designs (CCI) were chosen because the specified limits (higher and lower factor levels) corresponded to or were very close to truly instrumental limits for most of the selected factors. Each design included a full or fractional factorial portion created within the specified factor limits (by dividing the distance from the design center to the factor limits by α), from which the first order or linear effects could be estimated; a central point, from which curvature due to two-way interactions or quadratic terms could be evaluated; and a star portion with experimental points at the specified factor limits, from which the factors responsible for response surface curvature could be known. With this type of design, all factors were studied in five levels. The number of required experiments was given by 2 K-p + 2K + C, where K is the number of factors included in the design, p the fraction of the full factorial design to be run, and C the number of replicates at the central point; α values were dependent on the number of variables and the desired design characteristics, such as rotatability and orthogonality. In this study, all CCIs were designed to be orthogonal and rotatable. All steps of the experimental design selection, statistical data analysis, and prediction of optimal factor settings were performed in R software [51], with the “rsm” package [52].
KD Determination by Titration
To determine PvGK–GMP and PvGK–GDP KDs by titration, the relative abundances of bound to total protein in the mass spectra were correlated to the relative equilibrium concentrations of bound to total protein in solution, as follows:
By plotting experimentally observed ratios between bound and total protein ion abundances against total concentration of ligand, KD could be obtained as a parameter of a nonlinear least squares curve fitting [22]:
KD Determination by Competition
To determine PvGK–GDP KD by competition, the ion abundance of the reference protein –ligand complex with a known binding constant (PvGK–GMP, with KD determined by titration) was monitored in the presence and in the absence of the competitor ligand (GDP) and correlated to the corresponding solution equilibrium concentrations.
By plotting the observed ratios between the reference protein –ligand complex in the presence and absence of competitor ligand as a function of total concentration of competitor ligand, the KD of the competitor ligand could be obtained as a parameter of a nonlinear least squares curve fitting [23]:
Where,
Results and Discussion
Optimization of PvGK-GMP Complex Over Free PvGK Ion Abundances
Based on preliminary screening experiments (Supplementary Figure S2), nine out of the 14 ESI source parameters were selected for statistical optimization. Optimization of the ESI source was carried out in two stages because of the large number of variables. In the first, sample flow rate (120 to 630 μL/h), drying gas flow rate (20 to 60 L/min), drying gas temperature (100 to 150 °C) and nebulizer gas pressure (40 to 70 psi) were studied. In the second, capillary exit voltage (50 to 200 V), skimmer 1 voltage (15 to 25 V), skimmer 2 voltage (5 to 25 V), capillary voltage (–4000 to –6000 V), and end plate voltage (–2500 to –3500 V) were studied. It was assumed that if any interactions between factors included in the first and second CCI existed, these would not contribute significantly to the MS response. While the first stage CCI was carried out in 30 runs (24 + 2 × 4 + 6), the second was carried out in 50 runs (25 + 2 × 5 + 8).
After MS response collection, a second order model was used to fit the data using the least squares method. Different MS response transformations were evaluated, including square root, logarithmic, and squared transformations. Raw data was chosen last because residuals had a closer distribution to normality. Supplementary Tables S1 and S3 provide, for first and second stage CCI, respectively, the fitted model terms coefficients and respective statistical significances. Similarly, Supplementary Tables S2 and S4 show the ANOVA employed to evaluate the quality of the model fitted.
For the first stage CCI, a P-value of 5.82 × 10–9 was obtained, indicating the model was statistically significant. Moreover, the model had an adjusted R2 of 0.94 and a lack of fit P-value of 0.15 (indicating that lack of fit errors were due to random errors and were not considered statistically different from pure errors) (Supplementary Table S2). The residuals were normally distributed and equally dispersed for all predicted values, with no systematic trends observed. All variables had significant main effect on PL/P, with sample flow rate showing a significant quadratic term and no interaction with any other parameter (Supplementary Table S1).
Sample flow rate could be tuned independently of the settings of other variables. A threefold increase in sample flow rate from 120 μL/h up to around 400 μL/h favored PvGK–GMP complex over free PvGK ion abundances, after which the tendency was inverted.
Drying gas temperature significantly interacted with drying gas flow rate. Thus, these parameters needed to be tuned together. The heated drying gas, N2, flows counter-current to the charged droplets trajectory and is very important for desolvation. Lower drying gas temperatures combined with higher drying gas flow rates provided the best PL/P. The three-dimensional response surface plots are presented in Figure 1.
Three-dimensional response surface plots of the experimental region studied by first stage CCI, PvGK-GMP system. Slices of the response surface are displayed for two variables at a time, with other variables fixed at the central point (flow rate 375 μL/h; drying gas flow rate 40 L/min; drying gas temperature 125 °C; nebulizer gas pressure 55 psi). (a) Effect of flow rate and drying gas flow rate. (b) Effect of flow rate and drying gas temperature. (c) Effect of flow rate and nebulizer gas pressure. (d) Effect of drying gas flow rate and drying gas temperature. (e) Effect of drying gas flow rate and nebulizer gas pressure. (f) Effect of drying gas temperature and nebulizer gas pressure
According to these results, better PL/P responses could theoretically be achieved outside the experimental region, especially if drying gas temperatures less than 100 °C were used with flow rates in the order of 400–420 μL/h. However, due to the observed trade-off between the relative ion abundances and intensities of both free protein and protein–ligand complex, such combination of factor levels could not be tested or used in practice. A second order model was also fitted to the signal-to-noise ratios (S/N) of free PvGK and PvGK–GMP complex with the only statistically significant interaction being between drying gas temperature and sample flow rate. The obtained three-dimensional surface plot for these two variables is shown in Figure 2 [(a) for PvGK-GMP complex and (b) for free PvGK]. It is interesting to note that at the lowest drying gas temperature (100 °C), the S/N of both free PvGK and PvGK–GMP complex decrease with increasing sample flow rate (probably due to a decrease in desolvation efficiency). However, if enough desolvation energy is provided by increasing the drying gas temperature, this trend is inverted (S/N increase with increasing sample flow rates).
Experimental verification confirmed that the maximum response was obtained at sample flow rate 400 μL/h, drying gas flow rate 45 L/min, drying gas temperature 105 °C, and nebulizer gas pressure 58 psi.
For the second stage CCI, the regression model was not significant (P-value of 0.73), even though the fit to the experimental data was good (lack of fit test was nonsignificant with a P-value of 0.66 and residuals had a normal distribution) (Supplementary Table S4).
Given the nonsignificant impact of capillary exit, skimmer 1, skimmer 2, end plate, and capillary voltages on relative ion abundances, these variables were used to simultaneously maximize the S/N of both free PvGK and the PvGK–GMP complex (1:1 stoichiometry). This approach involved the construction of a desirability function for each individual response and their combination by using the weighted geometric average to obtain an overall desirability function (R package “desirability” [53]) to establish capillary exit 126.5 V; skimmer 1 15.42 V; skimmer 2 15.55 V; end plate –3085 V; capillary voltage –4033 V. The improvement in S/N was 76% and 87% for free PvGK and PvGK-GMP complex, respectively.
The overall optimized conditions for PvGK and the screening conditions are compared in Table 1. The screening conditions allowed the observation of protein and protein–ligand complexes under native conditions, even though relative solution-phase equilibrium concentrations between the protein–ligand complex and free protein might not be preserved. Indeed, the optimized conditions for PvGK-GMP led to a relative ion abundance of 1.75, whereas screening conditions gave a relative ion abundance of 0.85.
PvGK–GMP Titration Experiments and KD Determination
Titration experiments demonstrated that under ESI optimized conditions the relative ion abundances were higher compared with screening conditions. Representative spectra obtained from samples containing PvGK (2 μM) and three different concentrations of GMP (2.4, 4.8, and 9.6 μM) acquired under both optimized and screening ESI source conditions are shown and compared in Figure 3.
At GMP concentrations of 9.6 μM and above, nonspecific protein–ligand binding was detected. The model developed by Daubenfeld et al. [37] was used to separate specific and nonspecific binding. The plot of the fraction of bound protein ([PL]/[P]t) versus total concentration of GMP and the best curve fit is shown in Figure 4. The KD obtained after nonlinear least squares curve fitting using Equation (2) was 1.84 ± 0.11 μM and 3.67 ± 0.30 μM, respectively, for ESI source optimized and screening conditions.
Fraction of bound protein ([PL]/[P]t) obtained under both ESI optimized and screening (pre-optimized) conditions plotted against total ligand concentration, GMP. The curves of best fit using the method of least squares with Equation (2) are present
The KD obtained under ESI source optimized conditions for PvGK–GMP was in good agreement with the results obtained for GMP binding to Plasmodium falciparum guanylate kinase (1.45 ± 0.05 μM at 25 °C, determined by isothermal titration calorimetry) [54]. By contrast, the KD determined under screening conditions was an overestimate.
Competition Experiments for PvGK-GDP KD Determination
Competition experiments revealed a clear competition between GMP and GDP for protein binding (Figure 5). As the concentration of GDP was increased in solution GMP was proportionally displaced from the protein, as can be observed from the decrease of PvGK–GMP and increase of PvGK-GDP complexes in Figure 5a and the linear relationship between the ratio of their ion abundances and total GDP concentration in Figure 5c. The degree of GMP displacement from the protein due to the presence of GDP was used to quantitatively estimate GDP KD (Figure 5b). The GDP KD was estimated to be 0.31 ± 0.07μM. Given GDP has one more phosphate group compared with GMP, it contains more possibilities for electrostatic/Coulombic interactions with the protein and, thus, for binding more tightly. To cross-validate the ESI source optimization approach, this was independently optimized for the PvGK–GDP system. The KD determined by titration thereafter was compared with the KD determined by competition.
Competiton experiments: spectra obtained from samples containing 2 μM PvGK, 9.6 μM GMP, and increasing concentrations of GDP [(0 μM in 1), (1.2 μM in 2), (2.4 μM in 3), (4.8 μM in 4), and (9.6 μM in 5)] are shown in (a). Reference protein–ligand complex [PvGK:GMP (2:9.6 μM)] concentration in the presence of competitor ligand (GDP) over reference protein–ligand complex concentration in the absence of competitor ligand is plotted as a function of total concentration of GDP in (b). The solid line represents the best fit obtained using Equation (4). The linear relationship between the ion abundances of PvGK-GDP complex over PvGK-GMP complex and total GDP concentration is shown in (c)
Optimization of PvGK-GDP Complex Over Free PvGK Ion Abundances
Optimization of the ESI source for the PvGK–GDP system was carried out in two stages; in the first, sample flow rate (120 to 630 μL/h), drying gas flow rate (20 to 60 L/min), drying gas temperature (100 to 150 °C), and nebulizer gas pressure (30 to 80 psi) were studied. In the second, capillary exit voltage (50 to 200 V), skimmer 1 voltage (15 to 25 V), skimmer 2 voltage (5 to 25 V), capillary voltage (–4000 to –7000 V), and end plate voltage (–2500 to –3500 V) were studied.
The coefficients of the second-order model estimated with the raw MS responses of the first stage CCI is shown in Supplementary Table S5. Its ANOVA table (Supplementary Table S6) shows that the model was statistically significant (P-value 2.00 × 10–9), presenting an adjusted R2 of 0.85 and nonsignificant lack of fit (P-value 0.08). All variables except drying gas flow rate had significant main effect on PL/P. Similarly to the PvGK–GMP system, sample flow rate had a significant quadratic term and a significant interaction with nebulizer gas pressure. Drying gas flow rate and drying gas temperature did not influence PL/P significantly. Confirmation experiments conducted at specified distances from the design center (0, 0.5, 1, 1.5) verified that maximum PL/P response could be obtained at sample flow rate 145 μL/h, drying gas flow rate 38 L/min, drying gas temperature 101 °C, and nebulizer gas pressure 72 psi.
For the second-stage CCI, the ANOVA demonstrates that the regression model estimated with the raw MS responses was statistically significant (P-value 2.27 × 10–4), presenting an adjusted R2 of 0.32 and nonsignificant lack of fit (P-value 0.17) (Supplementary Table S8). Contrary to what was observed for the PvGK–GMP system, capillary exit had a significant main effect, as did its interaction with skimmer 1 (Supplementary Table S7). At lower skimmer 1 settings, an increase in capillary exit promoted a decrease in relative ion abundances, whereas at higher skimmer 1 values, an increase in capillary exit increased considerably relative ion abundances (Figure 6a). A similar effect was observed for a statistically significant interaction between end plate and capillary voltages (Figure 6b). For the PvGK–GDP system no trade-off between relative ion abundances and S/N of both free PvGK and PvGK-GDP complex was found. We believe that the additional phosphate group of GDP contributes to the increased stability of the protein–ligand complex, thus tolerating higher voltages, which improve the declustering efficiency without promoting the complex dissociation. Experimental results confirmed that the maximum response could be obtained at capillary –5100 V, end plate –3392 V, capillary exit 200 V, skimmer 1, 23.60 V, and skimmer 2, 15.00 V. The overall optimized and screening conditions are compared in Table 1.
Three-dimensional response surface plots of skimmer 1 versus capillary exit (a) and capillary versus end plate voltages (b) (second stage CCI, PvGK-GDP system). Other variables are fixed at the central point (skimmer 1, 20 V; skimmer 2, 15 V; end plate –3500 V; capillary voltage –5500 V; capillary exit 125 V)
PvGK–GDP Titration Experiments and KD Determination
The KD determined after nonlinear least squares curve fitting using Equation (2) was 0.30 ± 0.03 μM. The KD was in excellent agreement with the KD found by competition experiments (0.31 ± 0.07 μM). These results confirm the value of ESI source optimization by statistical DOE and RSM in order to improve the accuracy of KD determinations by ESI-MS.
Conclusions
A systematic approach, combining statistical DOE with RSM, simultaneously maximizes the relative ionization efficiency of a protein–ligand complex over free protein, minimizes the dissociation of protein–ligand complex and enables the establishment of appropriate ESI source settings for KD determination by titration. If the fractional protein occupancy tends to one as ligand concentrations are increased, KD can directly be determined by nonlinear least squares curve fitting using Equation (2). However, if nonspecific ligand binding is observed, this needs to be considered before KD determination. In this study, the model introduced by Daubenfeld et al. [37], which explains ligand binding as a convolution of two different statistical distributions (a binomial distribution for specific ligand binding and a Poisson distribution for nonspecific ligand binding), was used to compute the contributions from only specific ligand binding. If the fractional protein occupancy does not tend to one even after ESI source optimization, we propose that the horizontal asymptote on the right side is determined and used to correct the experimental data, in a similar way presented by Jaquillard et al. [25]. Here, the assumption that protein–ligand complex dissociation is only dependent on its gas-phase stability under vacuum and thus the type of intermolecular interactions involved in binding is made. Hence, a constant correction factor can be applied to the experimental fraction of bound protein obtained over the range of ligand concentrations used for the titration experiment.
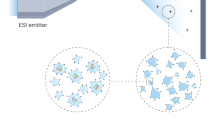
The optimization was demonstrated for the noncovalent interactions between PvGK and its natural substrates GMP and GDP. The results show that the ESI conditions, which better retain relative solution-phase equilibrium concentrations between a protein–ligand complex and free protein, depend on the protein–ligand system itself (Scheme 1). These results also showed that if ESI source instrumental conditions are carefully chosen, the accuracy of KD determinations by ESI-MS can be improved.
Summary of ESI source parameters studied by statistical DOE and RSM and their influence on relative ion abundances for (a) PvGK-GMP system and (b) PvGK-GDP system: parameters inside white boxes had no significant impact on relative ion abundances; parameters inside colored boxes had significant main effect on relative ion abundances; connecting lines denote significant interactions or quadratic effects
References
Hofstadler, S.A., Sannes-Lowery, K.A.: Applications of ESI-MS in drug discovery: interrogation of noncovalent complexes. Nat. Rev. Drug Discov. 5, 585–595 (2006)
Kitova, E.N., El-Hawiet, A., Schnier, P.D., Klassen, J.S.: Reliable determinations of protein–ligand interactions by direct ESI-MS measurements. Are we there yet? J. Am. Soc. Mass Spectrom. 23, 431–441 (2012)
Cole, R.B.: Electrospray and MALDI mass spectrometry: fundamentals, instrumentation, practicalities, and biological applications, 2nd edn. John Wiley and Sons, Inc., Hoboken, NJ (2010)
Pacholarz, K.J., Garlish, R.A., Taylor, R.J., Barran, P.E.: Mass spectrometry based tools to investigate protein–ligand interactions for drug discovery. Chem. Soc. Rev. 41, 4335–4355 (2012)
Erba, E.B., Zenobi, R.: Mass spectrometric studies of dissociation constants of noncovalent complexes. Annu. Rep. C (Phys. Chem.) 107, 199–228 (2011)
Peschke, M., Verkerk, U.H., Kebarle, P.: Features of the ESI mechanism that affect the observation of multiply charged non-covalent protein complexes and the determination of the association constant by the titration method. J. Am. Soc. Mass Spectrom. 15, 1424–1434 (2004)
Canon, F., Pate, F., Meudec, E., Marlin, T., Cheynier, V., Giuliani, A., Sarni-Manchado, P.: Characterization, stoichiometry, and stability of salivary protein–tannin complexes by ESI-MS and ESI-MS/MS. Anal. Bioanal. Chem. 395, 2535–2545 (2009)
Marcotte, D., Zeng, W., Hus, J.C., McKenzie, A., Hession, C., Jin, P., Bergeron, C., Lugovskoy, A., Enyedy, I., Cuervo, H., Wang, D., Atmanene, C., Roecklin, D., Vecchi, M., Vivat, V., Kraemer, J., Winkler, D., Hong, V., Chao, J., Lukashev, M., Silvian, L.: Small molecules inhibit the interaction of Nrf2 and the Keap1 Kelch domain through a noncovalent mechanism. Bioorg. Med. Chem. 21, 4011–4019 (2013)
Sanglier, S., Atmanene, C., Chevreux, G., Van Dorsselaer, A.: Nondenaturing mass spectrometry to study noncovalent protein/protein and protein/ligand complexes: technical aspects and application to the determination of binding stoichiometries. In: Thompson, J., Ueffing, M., Schaeffer-Reiss, C. (eds.) Functional Proteomics vol. 484. Methods in Molecular Biology, pp. 217–243. Humana Press: (2008)
Kitova, E.N., Soya, N., Klassen, J.S.: Identifying specific small-molecule interactions using electrospray ionization mass spectrometry. Anal. Chem. 83, 5160–5167 (2011)
Eckart, K., Spiess, J.: Electrospray ionization mass spectrometry of biotin binding to streptavidin. J. Am. Soc. Mass Spectrom. 6, 912–919 (1995)
Lin, X., Zhao, W., Wang, X.: Characterization of conformational changes and non-covalent complexes of myoglobin by electrospray ionization mass spectrometry, circular dichroism and fluorescence spectroscopy. J. Mass Spectrom. 45, 618–626 (2010)
Simmons, D.A., Wilson, D.J., Lajoie, G.A., Doherty-Kirby, A., Konermann, L.: Subunit disassembly and unfolding kinetics of hemoglobin studied by time-resolved electrospray mass spectrometry. Biochemistry 43, 14792–14801 (2004)
Natan, E., Hirschberg, D., Morgner, N., Robinson, C.V., Fersht, A.R.: Ultraslow oligomerization equilibria of p53 and its implications. Proc. Natl. Acad. Sci. 106, 14327–14332 (2009)
Daneshfar, R., Kitova, E.N., Klassen, J.S.: Determination of protein−ligand association thermochemistry using variable-temperature nanoelectrospray mass spectrometry. J. Am. Chem. Soc. 126, 4786–4787 (2004)
Shoemaker, G.K., Soya, N., Palcic, M.M., Klassen, J.S.: Temperature-dependent cooperativity in donor-acceptor substrate binding to the human blood group glycosyltransferases. Glycobiology 18, 587–592 (2008)
Lim, H.-K., Hsieh, Y.L., Ganem, B., Henion, J.: Recognition of cell-wall peptide ligands by vancomycin group antibiotics: studies using ion spray mass spectrometry. J. Mass Spectrom. 30, 708–714 (1995)
Greig, M.J., Gaus, H., Cummins, L.L., Sasmor, H., Griffey, R.H.: Measurement of macromolecular binding using electrospray mass spectrometry. determination of dissociation constants for oligonucleotide: serum albumin complexes. J. Am. Chem. Soc. 117, 10765–10766 (1995)
Loo, J.A., Peifeng, H., McConnell, P., Tom Mueller, W., Sawyer, T.K., Thanabal, V.: A study of Src SH2 domain protein–phosphopeptide binding interactions by electrospray ionization mass spectrometry. J. Am. Soc. Mass Spectrom. 8, 234–243 (1997)
Griffey, R.H., Hofstadler, S.A., Sannes-Lowery, K.A., Ecker, D.J., Crooke, S.T.: Determinants of aminoglycoside-binding specificity for rRNA by using mass spectrometry. Proc. Natl. Acad. Sci. 96, 10129–10133 (1999)
Sannes-Lowery, K.A., Griffey, R.H., Hofstadler, S.A.: Measuring dissociation constants of RNA and aminoglycoside antibiotics by electrospray ionization mass spectrometry. Anal. Biochem. 280, 264–271 (2000)
Bligh, S.W.A., Haley, T., Lowe, P.N.: Measurement of dissociation constants of inhibitors binding to Src SH2 domain protein by non-covalent electrospray ionization mass spectrometry. J. Mol. Recognit. 16, 139–148 (2003)
Tjernberg, A., Carno, S., Oliv, F., Benkestock, K., Edlund, P.O., Griffiths, W.J., Hallen, D.: Determination of dissociation constants for protein–ligand complexes by electrospray ionization mass spectrometry. Anal. Chem. 76, 4325–4331 (2004)
Fryčák, P., Schug, K.A.: Dynamic titration: determination of dissociation constants for noncovalent complexes in multiplexed format using HPLC-ESI-MS. Anal. Chem. 80, 1385–1393 (2008)
Jaquillard, L., Saab, F., Schoentgen, F., Cadene, M.: Improved accuracy of low affinity protein-ligand equilibrium dissociation constants directly determined by electrospray ionization mass spectrometry. J. Am. Soc. Mass Spectrom. 23, 908–922 (2012)
Erba, E.B., Barylyuk, K., Yang, Y., Zenobi, R.: Quantifying protein–protein interactions within noncovalent complexes using electrospray ionization mass spectrometry. Anal. Chem. 83, 9251–9259 (2011)
Gabelica, V., Galic, N., Rosu, F., Houssier, C., De Pauw, E.: Influence of response factors on determining equilibrium association constants of noncovalent complexes by electrospray ionization mass spectrometry. J. Mass Spectrom. 38, 491–501 (2003)
Chitta, R., Rempel, D., Gross, M.: Determination of affinity constants and response factors of the non-covalent dimer of gramicidin by electrospray ionization mass spectrometry and mathematical modeling. J. Am. Soc. Mass Spectrom. 16, 1031–1038 (2005)
Gabelica, V., Rosu, F., De Pauw, E.: A simple method to determine electrospray response factors of noncovalent complexes. Anal. Chem. 81, 6708–6715 (2009)
Wang, W., Kitova, E.N., Klassen, J.S.: Nonspecific protein−carbohydrate complexes produced by nanoelectrospray ionization. Factors influencing their formation and stability. Anal. Chem. 77, 3060–3071 (2005)
Wang, W., Kitova, E., Sun, J., Klassen, J.: Blackbody infrared radiative dissociation of nonspecific protein-carbohydrate complexes produced by nanoelectrospray ionization: the nature of the non-covalent interactions. J. Am. Soc. Mass Spectrom. 16, 1583–1594 (2005)
Deng, L., Sun, N., Kitova, E.N., Klassen, J.S.: Direct quantification of protein−metal ion affinities by electrospray ionization mass spectrometry. Anal. Chem. 82, 2170–2174 (2010)
Sun, N., Soya, N., Kitova, E., Klassen, J.: Nonspecific interactions between proteins and charged biomolecules in electrospray ionization mass spectrometry. J. Am. Soc. Mass Spectrom. 21, 472–481 (2010)
Sun, N., Sun, J., Kitova, E., Klassen, J.: Identifying nonspecific ligand binding in electrospray ionization mass spectrometry using the reporter molecule method. J. Am. Soc. Mass Spectrom. 20, 1242–1250 (2009)
Sun, J., Kitova, E.N., Wang, W., Klassen, J.S.: Method for distinguishing specific from nonspecific protein−ligand complexes in nanoelectrospray ionization mass spectrometry. Anal. Chem. 78, 3010–3018 (2006)
Sun, J., Kitova, E.N., Sun, N., Klassen, J.S.: Method for identifying nonspecific protein−protein interactions in nanoelectrospray ionization mass spectrometry. Anal. Chem. 79, 8301–8311 (2007)
Daubenfeld, T., Bouin, A.-P., van der Rest, G.: A deconvolution method for the separation of specific versus nonspecific interactions in non-covalent protein–ligand complexes analyzed by ESI-FT-ICR mass spectrometry. J. Am. Soc. Mass Spectrom. 17, 1239–1248 (2006)
Shimon, L., Sharon, M., Horovitz, A.: A method for removing effects of nonspecific binding on the distribution of binding stoichiometries: application to mass spectroscopy data. Biophys. J. 99, 1645–1649 (2010)
van Dongen, W.D., Heck, A.J.R.: Binding of selected carbohydrates to apo-concanavalin A studied by electrospray ionization mass spectrometry. Analyst 125, 583–589 (2000)
Benkestock, K., Sundqvist, G., Edlund, P.-O., Roeraade, J.: Influence of droplet size, capillary–cone distance and selected instrumental parameters for the analysis of noncovalent protein–ligand complexes by nano-electrospray ionization mass spectrometry. J. Mass Spectrom. 39, 1059–1067 (2004)
Sun, J., Kitova, E.N., Klassen, J.S.: Method for stabilizing protein−ligand complexes in nanoelectrospray ionization mass spectrometry. Anal. Chem. 79, 416–425 (2006)
Wang, W., Kitova, E.N., Klassen, J.S.: Influence of solution and gas phase processes on protein−carbohydrate binding affinities determined by nanoelectrospray Fourier transform ion cyclotron resonance mass spectrometry. Anal. Chem. 75, 4945–4955 (2003)
Håkansson, K., Axelsson, J., Palmblad, M., Håkansson, P.: Mechanistic studies of multipole storage assisted dissociation. J. Am. Soc. Mass Spectrom. 11, 210–217 (2000)
Wang, J., Schnute, W.C.: Optimizing mass spectrometric detection for ion chromatographic analysis. I. Common anions and selected organic acids. Rapid Commun. Mass Spectrom. 23, 3439–3447 (2009)
Jiao, J., Carella, A.J., Steeno, G.S., Darrington, R.T.: Optimization of triple quadrupole mass spectrometer for quantitation of trace degradants of pharmaceutical compounds. Int. J. Mass Spectrom. 216, 209–218 (2002)
Barry, J.A., Muddiman, D.C.: Global optimization of the infrared matrix-assisted laser desorption electrospray ionization (IR MALDESI) source for mass spectrometry using statistical design of experiments. Rapid Commun. Mass Spectrom. 25, 3527–3536 (2011)
Vu, H., Quinn, R.J.: Direct screening of natural product extracts using mass spectrometry. J. Biomol. Screen 13, 265–275 (2008)
Riter, L.S., Vitek, O., Gooding, K.M., Hodge, B.D., Julian, R.K.: Statistical design of experiments as a tool in mass spectrometry. J. Mass Spectrom. 40, 565–579 (2005)
Bezerra, M.A., Santelli, R.E., Oliveira, E.P., Villar, L.S., Escaleira, L.A.: Response surface methodology (RSM) as a tool for optimization in analytical chemistry. Talanta 76, 965–977 (2008)
Mehlin, C., Boni, E., Buckner, F.S., Engel, L., Feist, T., Gelb, M.H., Haji, L., Kim, D., Liu, C., Mueller, N., Myler, P.J., Reddy, J.T., Sampson, J.N., Subramanian, E., Van Voorhis, W.C., Worthey, E., Zucker, F., Hol, W.G.J.: Heterologous expression of proteins from Plasmodium falciparum: results from 1000 genes. Mol. Biochem. Parasitol. 148, 144–160 (2006)
R Core Team: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria (2013)
Lenth, R.V.: Response-surface methods in R, using rsm. J. Stat Softw 32, 1–17 (2009)
Kuhn, M.: Desirability: Desirabiliy Function Optimization and Ranking. Available at: http://CRAN.R-project.org/package=desirability (2014) Accessed 22 Jan 2014
Kandeel, M., Kitade, Y.: Binding dynamics and energetic insight into the molecular forces driving nucleotide binding by guanylate kinase. J. Mol. Recognit. 24, 322–332 (2011)
Acknowledgments
The authors thank Dr. James McBroom for support on statistical data analysis, Dr. Gregory J. Crowther for organizing protein shipment, and Dr. Hoan Vu for support in instrument troubleshooting. They thank the Australian Research Council for funding the purchase of the 4.7 Tesla FTMS (LE0237908) and a Linkage grant (LP120100485). They acknowledge the Bill & Melinda Gates Foundation Grand Challenges Explorations Grant Phase II OPP1035218 GCE. L.P. acknowledges Griffith University for the provision of the Griffith University International Postgraduate Research Scholarship and Griffith University Postgraduate Research Scholarship.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 588 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Pedro, L., Van Voorhis, W.C. & Quinn, R.J. Optimization of Electrospray Ionization by Statistical Design of Experiments and Response Surface Methodology: Protein–Ligand Equilibrium Dissociation Constant Determinations. J. Am. Soc. Mass Spectrom. 27, 1520–1530 (2016). https://doi.org/10.1007/s13361-016-1417-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13361-016-1417-x