Abstract
Background
Dabrafenib and irinotecan are two drugs that can be utilized to treat melanoma. A previous in vivo study has shown that dabrafenib enhances the antitumor activity of irinotecan in a xenograft model with unclear mechanism.
Objectives
This study aims to investigate the inhibition of dabrafenib on SN-38 (the active metabolite of irinotecan) glucuronidation, trying to elucidate the possible mechanism underlying the synergistic effect and to provide a basis for further development and optimization of this combination in clinical research.
Methods
Recombinant human uridine diphosphate glucuronosyltransferase 1A1 (UGT1A1) and human liver microsomes (HLMs) were employed to catalyze the glucuronidation of SN-38 in vitro. Inhibition kinetic analysis and quantitative prediction study were combined to predict drug–drug interaction (DDI) potential in vivo.
Results
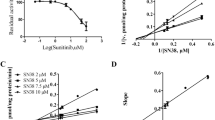
Dabrafenib noncompetitively inhibited SN-38 glucuronidation in pooled HLMs and recombinant UGT1A1 with unbound inhibitor constant (Ki,u) values of 12.43 ± 0.28 and 3.89 ± 0.40 μM, respectively. Based on the in vitro Ki,u value and estimation of kinetic parameters, dabrafenib administered at 150 mg twice daily may result in about a 1−2% increase in the area under the curve (AUC) of SN-38 in vivo. However, the ratios of intra-enterocyte concentration of dabrafenib to Ki,u ([I]gut/Ki,u) are 2.73 and 8.72 in HLMs and recombinant UGT1A1, respectively, indicating a high risk of intestinal DDI when dabrafenib was used in combination with irinotecan.
Conclusion
Dabrafenib is a potent noncompetitive inhibitor of UGT1A1 and may bring potential risk of DDI when combined with irinotecan.





Similar content being viewed by others
References
Maldonado JL, Fridlyand J, Patel H, Jain AN, Busam K, Kageshita T, et al. Determinants of BRAF mutations in primary melanomas. J Natl Cancer Inst. 2003;95(24):1878–90.
Davies H, Bignell GR, Cox C, Stephens P, Edkins S, Clegg S, et al. Mutations of the BRAF gene in human cancer. Nature. 2002;417(6892):949–54.
Eggermont AM, Robert C. New drugs in melanoma: it’s a whole new world. Eur J Cancer. 2011;47(14):2150–7.
Hauschild A, Grob JJ, Demidov LV, Jouary T, Gutzmer R, Millward M, et al. Dabrafenib in BRAF-mutated metastatic melanoma: a multicentre, open-label, phase 3 randomised controlled trial. Lancet. 2012;380(9839):358–65.
Corcoran RB, Andre T, Atreya CE, Schellens JHM, Yoshino T, Bendell JC, et al. Combined BRAF, EGFR, and MEK inhibition in patients with BRAF(V600E)-mutant colorectal cancer. Cancer Discov. 2018;8(4):428–43.
Long GV, Stroyakovskiy D, Gogas H, Levchenko E, de Braud F, Larkin J, et al. Dabrafenib and trametinib versus dabrafenib and placebo for Val600 BRAF-mutant melanoma: a multicentre, double-blind, phase 3 randomised controlled trial. Lancet. 2015;386(9992):444–51.
Robert C, Karaszewska B, Schachter J, Rutkowski P, Mackiewicz A, Stroiakovski D, et al. Improved overall survival in melanoma with combined dabrafenib and trametinib. N Engl J Med. 2015;372(1):30–9.
Lawrence SK, Nguyen D, Bowen C, Richards-Peterson L, Skordos KW. The metabolic drug–drug interaction profile of dabrafenib: in vitro investigations and quantitative extrapolation of the P450-mediated DDI risk. Drug Metab Dispos. 2014;42(7):1180–90.
Suttle AB, Grossmann KF, Ouellet D, Richards-Peterson LE, Aktan G, Gordon MS, et al. Assessment of the drug interaction potential and single- and repeat-dose pharmacokinetics of the BRAF inhibitor dabrafenib. J Clin Pharmacol. 2015;55(4):392–400.
Gao K, Lockwood WW, Li J, Lam W, Li G. Genomic analyses identify gene candidates for acquired irinotecan resistance in melanoma cells. Int J Oncol. 2008;32(6):1343–9.
Hanioka N, Ozawa S, Jinno H, Ando M, Saito Y, Sawada J. Human liver UDP-glucuronosyltransferase isoforms involved in the glucuronidation of 7-ethyl-10-hydroxycamptothecin. Xenobiotica. 2001;31(10):687–99.
Araki E, Ishikawa M, Iigo M, Koide T, Itabashi M, Hoshi A. Relationship between development of diarrhea and the concentration of SN-38, an active metabolite of CPT-11, in the intestine and the blood plasma of athymic mice following intraperitoneal administration of CPT-11. Jpn J Cancer Res. 1993;84(6):697–702.
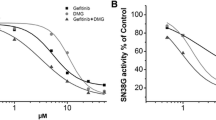
Li W, Xing Y, Liu Y. Inhibition of SN-38 glucuronidation by gefitinib and its metabolite. Cancer Chemother Pharmacol. 2015;75(6):1253–60.
Jiang L, Wang L, Zhang Z, Wang Z, Wang X, Wang S, et al. The pharmacokinetic interaction between irinotecan and sunitinib. Cancer Chemother Pharmacol. 2020;85(2):443–8.
Iwase M, Fujita KI, Nishimura Y, Seba N, Masuo Y, Ishida H, et al. Pazopanib interacts with irinotecan by inhibiting UGT1A1-mediated glucuronidation, but not OATP1B1-mediated hepatic uptake, of an active metabolite SN-38. Cancer Chemother Pharmacol. 2019;83(5):993–8.
Yanagihara K, Kubo T, Mihara K, Kuwata T, Ochiai A, Seyama T, et al. Establishment of a novel cell line from a rare human duodenal poorly differentiated neuroendocrine carcinoma. Oncotarget. 2018;9(92):36503–14.
Korprasertthaworn P, Chau N, Nair PC, Rowland A, Miners JO. Inhibition of human UDP-glucuronosyltransferase (UGT) enzymes by kinase inhibitors: effects of dabrafenib, ibrutinib, nintedanib, trametinib and BIBF 1202. Biochem Pharmacol. 2019;169:113616.
Yin H, Wang Z, Wang X, Lv X, Fan X, Yan M, et al. Inhibition of human UDP-glucuronosyltransferase enzyme by dabrafenib: implications for drug–drug interactions. Biomed Chromatogr. 2021;35:e5205.
Zhang N, Liu Y, Jeong H. Drug–drug interaction potentials of tyrosine kinase inhibitors via inhibition of UDP-glucuronosyltransferases. Sci Rep. 2015;5:17778.
Copeland AR. Enzymes: a practical introduction to structure, mechanism, and data analysis. Sov Appl Mech. 2000;26(6):515–23.
Ito K, Brown HS, Houston JB. Database analyses for the prediction of in vivo drug–drug interactions from in vitro data. Br J Clin Pharmacol. 2004;57(4):473–86.
Rostami-Hodjegan A, Tucker G. ‘In silico’ simulations to assess the ‘in vivo’ consequences of ‘in vitro’ metabolic drug–drug interactions. Drug Discov Today Technol. 2004;1(4):441–8.
Ouellet D, Gibiansky E, Leonowens C, O’Hagan A, Haney P, Switzky J, et al. Population pharmacokinetics of dabrafenib, a BRAF inhibitor: effect of dose, time, covariates, and relationship with its metabolites. J Clin Pharmacol. 2014;54(6):696–706.
Davies B, Morris T. Physiological parameters in laboratory animals and humans. Pharm Res. 1993;10(7):1093–5.
USFDA. Approved label. https://www.accessdata.fda.gov/drugsatfda_docs/label/2018/202806s008lbl.pdf. Accessed 8 Jan 2021.
Obach RS, Walsky RL, Venkatakrishnan K. Mechanism-based inactivation of human cytochrome p450 enzymes and the prediction of drug–drug interactions. Drug Metab Dispos. 2007;35(2):246–55.
USFDA. In vitro drug interaction studies—cytochrome P450 enzyme- and transporter-mediated drug interactions guidance for industry; 2020.
Austin RP, Barton P, Cockroft SL, Wenlock MC, Riley RJ. The influence of nonspecific microsomal binding on apparent intrinsic clearance, and its prediction from physicochemical properties. Drug Metab Dispos. 2002;30(12):1497–503.
Liu Y, Ramirez J, House L, Ratain MJ. Comparison of the drug–drug interactions potential of erlotinib and gefitinib via inhibition of UDP-glucuronosyltransferases. Drug Metab Dispos. 2010;38(1):32–9.
Burns K, Nair PC, Rowland A, Mackenzie PI, Knights KM, Miners JO. The nonspecific binding of tyrosine kinase inhibitors to human liver microsomes. Drug Metab Dispos. 2015;43(12):1934–7.
Puszkiel A, Noé G, Bellesoeur A, Kramkimel N, Paludetto M-N, Thomas-Schoemann A, et al. Clinical pharmacokinetics and pharmacodynamics of dabrafenib. Clin Pharmacokinet. 2019;58(4):451–67.
Fuchs CS, Moore MR, Harker G, Villa L, Rinaldi D, Hecht JR. Phase III comparison of two irinotecan dosing regimens in second-line therapy of metastatic colorectal cancer. J Clin Oncol. 2003;21(5):807–14.
Berg AK, Buckner JC, Galanis E, Jaeckle KA, Ames MM, Reid JM. Quantification of the impact of enzyme-inducing antiepileptic drugs on irinotecan pharmacokinetics and SN-38 exposure. J Clin Pharmacol. 2015;55(11):1303–12.
Ramchandani RP, Wang Y, Booth BP, Ibrahim A, Johnson JR, Rahman A, et al. The role of SN-38 exposure, UGT1A1*28 polymorphism, and baseline bilirubin level in predicting severe irinotecan toxicity. J Clin Pharmacol. 2007;47(1):78–86.
Gupta E, Mick R, Ramirez J, Wang X, Lestingi TM, Vokes EE, et al. Pharmacokinetic and pharmacodynamic evaluation of the topoisomerase inhibitor irinotecan in cancer patients. J Clin Oncol. 1997;15(4):1502–10.
Gupta E, Lestingi TM, Mick R, Ramirez J, Vokes EE, Ratain MJ. Metabolic fate of irinotecan in humans: correlation of glucuronidation with diarrhea. Cancer Res. 1994;54(14):3723–5.
Mick R, Gupta E, Vokes EE, Ratain MJ. Limited-sampling models for irinotecan pharmacokinetics–pharmacodynamics: prediction of biliary index and intestinal toxicity. J Clin Oncol. 1996;14(7):2012–9.
Innocenti F, Undevia SD, Iyer L, Chen PX, Das S, Kocherginsky M, et al. Genetic variants in the UDP-glucuronosyltransferase 1A1 gene predict the risk of severe neutropenia of irinotecan. J Clin Oncol. 2004;22(8):1382–8.
Bosma PJ, Chowdhury JR, Bakker C, Gantla S, de Boer A, Oostra BA, et al. The genetic basis of the reduced expression of bilirubin UDP-glucuronosyltransferase 1 in Gilbert’s syndrome. N Engl J Med. 1995;333(18):1171–5.
Sugatani J, Yamakawa K, Yoshinari K, Machida T, Takagi H, Mori M, et al. Identification of a defect in the UGT1A1 gene promoter and its association with hyperbilirubinemia. Biochem Biophys Res Commun. 2002;292(2):492–7.
Ando Y, Saka H, Asai G, Sugiura S, Shimokata K, Kamataki T. UGT1A1 genotypes and glucuronidation of SN-38, the active metabolite of irinotecan. Ann Oncol. 1998;9(8):845–7.
Wang Y, Shen L, Xu N, Wang JW, Jiao SC, Liu ZY, et al. UGT1A1 predicts outcome in colorectal cancer treated with irinotecan and fluorouracil. World J Gastroenterol. 2012;18(45):6635–44.
Iyer L, Hall D, Das S, Mortell MA, Ramírez J, Kim S, et al. Phenotype-genotype correlation of in vitro SN-38 (active metabolite of irinotecan) and bilirubin glucuronidation in human liver tissue with UGT1A1 promoter polymorphism. Clin Pharmacol Ther. 1999;65(5):576–82.
Iyer L, Das S, Janisch L, Wen M, Ramírez J, Karrison T, et al. UGT1A1*28 polymorphism as a determinant of irinotecan disposition and toxicity. Pharmacogenom J. 2002;2(1):43–7.
Rostami-Hodjegan A, Tucker GT. Simulation and prediction of in vivo drug metabolism in human populations from in vitro data. Nat Rev Drug Discov. 2007;6(2):140–8.
Yang N, Sun R, Liao X, Aa J, Wang G. UDP-glucuronosyltransferases (UGTs) and their related metabolic cross-talk with internal homeostasis: a systematic review of UGT isoforms for precision medicine. Pharmacol Res. 2017;121:169–83.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interests regarding the publication of this article.
Ethics approval
Not applicable.
Funding
This study was financially supported by the National Key Research and Development Program of China (2017YFC1702006), the Fundamental Research Funds for the Central Universities (DUT21LK11).
Availability of data and material
The material supporting the conclusion has been included within the article.
Code availability
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Author contributions
ZW, JC, and YL contributed to the study conception and design. ZW, XW, and ZW contributed to the acquisition of data. XF and MY contributed to the analysis and interpretation of the data. ZW, LJ, and YX participated in the analysis and interpretation of the data and in the drafting of this article. All authors provided critical revision of the article for important intellectual content.
Rights and permissions
About this article
Cite this article
Wang, Z., Wang, X., Wang, Z. et al. Prediction of Drug–Drug Interaction Between Dabrafenib and Irinotecan via UGT1A1-Mediated Glucuronidation. Eur J Drug Metab Pharmacokinet 47, 353–361 (2022). https://doi.org/10.1007/s13318-021-00740-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13318-021-00740-x