Abstract
Introduction
As one of the main yield components, grain number per panicle (GNP) played critical role in the rice yield improvement. The identification of natural advantageous variations under different situations will promote the sustainable genetic improvement in rice yield.
Objectives
This study was designed to identify natural genotypes in a rice mini-core collection, to examine the genotypic effects across the indica and japonica genetic background in different environments, and excavating the superior genotypes that had drove the modern genetic improvement.
Methods
The association mapping of GNP was carried out using a mini-core collection including 154 indica and 119 japonica accessions in seven different environments. Genotypic effects of each genotype for each QTL were calculated and genotype frequency distortion between the commercial rice cultivars and landraces was screened by χ2-test.
Results
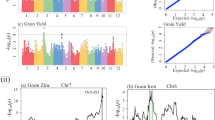
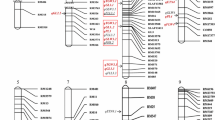
In total, 74 QTLs containing stable and sensitive QTLs in various environments were detected. Within them, 20 positive and 24 negative genotypes in indica, and 24 positive and 16 negative genotypes in japonica were identified. When checking the accumulation of positive genotypes identified in indica across cultivars in each of the two subspecies, it indicated that increased number of positive genotypes identified in indica results in the substantially increased GNP in both indica and japonica across all of the environments, while this trend was not obvious for the positive genotypes identified in japonica especially in short day environments. Moreover, the positive and negative genotype frequency distortion between the landraces and commercial rice cultivars indicated that both positive selection of positive genotypes and negative selection of negative genotypes had driven the genetic improvement on GNP.
Conclusion
Our findings suggested that the accumulation of positive genotypes and purifying negative genotypes played equivalently important roles in the improvement of rice yield, but the efficient use for some QTLs or genotypes depends on the comprehensive evaluation of their effect under diverse genetic backgrounds and environments.



Similar content being viewed by others
References
Ashikari M, Sakakibara H, Lin S, Yamamoto T, Takashi T, Nishimura A et al (2005) Cytokinin oxidase regulates rice grain production. Science 309:741–745. https://doi.org/10.1126/science.1113373
Cai H, Diao S, He Y, Zhang L, Liu S, Zhu Y et al (2011) Genetic and physical mapping of qHY-8, a pleiotropic QTL for heading date and yield-related traits in rice. Euphytica 184:109–118. https://doi.org/10.1007/s10681-011-0581-0
Chow CY (2016) Bringing genetic background into focus. Nat Rev Genet 17:63–64. https://doi.org/10.1038/nrg.2015.9
Gong J, Du J, Fan Y, Wu J, Zhuang J (2010) Quantitative trait loci for panicle size and grain yield detected in interval RM111-RM19784 on the short arm of rice chromosome 6. Agric Sci China 9:1085–1092. https://doi.org/10.1016/s1671-2927(09)60194-2
Huang X, Qian Q, Liu Z, Sun H, He S, Luo D et al (2009) Natural variation at the DEP1 locus enhances grain yield in rice. Nat Genet 41:494–497. https://doi.org/10.1038/ng.352
Huang X, Wei X, Sang T, Zhao Q, Feng Q, Zhao Y et al (2010) Genome-wide association studies of 14 agronomic traits in rice landraces. Nat Genet 42:961–967. https://doi.org/10.1038/ng.695
Huang X, Zhao Y, Wei X, Li C, Wang A, Zhao Q et al (2011) Genome-wide association study of flowering time and grain yield traits in a worldwide collection of rice germplasm. Nat Genet 44:32–39. https://doi.org/10.1038/ng.1018
Huang X, Kurata N, Wei X, Wang ZX, Wang A, Zhao Q et al (2012) A map of rice genome variation reveals the origin of cultivated rice. Nature 490:497–501. https://doi.org/10.1038/nature11532
Hwang E, Song Q, Jia G, Specht JE, LHyten D, Costa D, J., et al (2014) A genome-wide association study of seed protein and oil content in soybean. BMC Genomics 15:1471–2164
Jia L, Yan W, Zhu C, Agrama HA, Jackson A, Yeater K et al (2011) Allelic analysis of sheath blight resistance with association mapping in rice. PLoS ONE 7:e32703. https://doi.org/10.1371/journal.pone.0032703.g001
Jiang G, Xiang Y, Zhao J, Yin D, Zhao X, Zhu L et al (2014) Regulation of inflorescence branch development in rice through a novel pathway involving the pentatricopeptide repeat protein sped1-D. Genetics 197:1395–1407. https://doi.org/10.1534/genetics.114.163931/-/DC1
Komatsu M, Maekawa M, Shimamoto K, Kyozuka J (2001) The LAX1 and FRIZZY PANICLE 2 genes determine the inflorescence architecture of rice by controlling rachis-branch and spikelet development. Dev Biol 231:364–373. https://doi.org/10.1006/dbio.2000.9988
Lawson MJ, Zhang L (2006) Distinct patterns of SSR distribution in the Arabidopsis thaliana and rice genomes. Genome Biol 7:1–11. https://doi.org/10.1186/gb-2006-7-2-r14
Li Y, Huang Y, Bergelson J, Nordborg M, Borevitz JO (2010) Association mapping of local climate-sensitive quantitative trait loci in Arabidopsis thaliana. Proc Natl Acad Sci USA 107:21199–21204. https://doi.org/10.1073/pnas.1007431107
Li S, Zhao B, Yuan D, Duan M, Qian Q, Tang L et al (2013) Rice zinc finger protein DST enhances grain production through controlling Gn1a/OsCKX2 expression. Proc Natl Acad Sci USA 110:3167–3172. https://doi.org/10.1073/pnas.1300359110
Liu T, Liu H, Zhang H, Xing Y (2013) Validation and characterization of Ghd7.1, a major quantitative trait locus with pleiotropic effects on spikelets per panicle, plant height, and heading date in rice (Oryza sativa L.). J Integr Plant Biol 55:917–927. https://doi.org/10.1111/jipb.12070
Londo JP, Chiang YC, Hung KH, Chiang TY, Schaal BA (2006) Phylogeography of Asian wild rice, Oryza rufipogon, reveals multiple independent domestications of cultivated rice, Oryza sativa. Proc Natl Acad Sci USA 103:9578–9583. https://doi.org/10.1073/pnas.0603152103
Narayanan NN, Niranjan B, Norman PO, Casiana MV, Samuel SG, Karabi D et al (2004) Molecular breeding marker-assisted selection combined with biolistic transformation for blast and bacterial blight resistance in indica rice. Mol Breed 14:61–71
Pan Y, Zhang H, Zhang D, Li J, Xiong H, Yu J et al (2015) Genetic analysis of cold tolerance at the germination and booting stages in rice by association mapping. PLoS ONE 10:e0120590. https://doi.org/10.1371/journal.pone.0120590
Piao R, Jiang W, Ham TH, Choi MS, Qiao Y, Chu SH et al (2009) Map-based cloning of the ERECT PANICLE 3 gene in rice. Theor Appl Genet 119:1497–1506. https://doi.org/10.1007/s00122-009-1151-x
Qiao Y, Piao R, Shi J, Lee SI, Jiang W, Kim BK et al (2011) Fine mapping and candidate gene analysis of dense and erect panicle 3, DEP3, which confers high grain yield in rice (Oryza sativa L.). Theor Appl Genet 122:1439–1449. https://doi.org/10.1007/s00122-011-1543-6
Rogers OS, Bendich AJ (1989) Extraction of DNA from plant tissues. Plant Mol Biol Manual 6:1–10
Sakamoto T, Matsuoka M (2008) Identifying and exploiting grain yield genes in rice. Curr Opin Plant Biol 11:209–214. https://doi.org/10.1016/j.pbi.2008.01.009
Sweeney M, McCouch S (2007) The complex history of the domestication of rice. Ann Bot 100:951–957. https://doi.org/10.1093/aob/mcm128
Tabuchi H, Zhang Y, Hattori S, Omae M, Shimizu-Sato S, Oikawa T et al (2011) LAX PANICLE2 of rice encodes a novel nuclear protein and regulates the formation of axillary meristems. Plant Cell 23:3276–3287. https://doi.org/10.1105/tpc.111.088765
Thiel T, Michalek W, Varshney RK, Graner A (2003) Exploiting EST databases for the development and characterization of gene-derived SSR-markers in barley (Hordeum vulgare L.). Theor Appl Genet 106:411–422. https://doi.org/10.1007/s00122-002-1031-0
Tian F, Bradbury PJ, Brown PJ, Hung H, Sun Q, Flint-Garcia S et al (2011) Genome-wide association study of leaf architecture in the maize nested association mapping population. Nat Genet 43:159–162. https://doi.org/10.1038/ng.746
Wu J, Qi Y, Hu G, Li J, Li Z, Zhang H (2016) Genetic architecture of flag leaf length and width in rice (Oryza sativa L.) revealed by association mapping. Genes Genomics 39:341–352. https://doi.org/10.1007/s13258-016-0501-8
Xue W, Xing Y, Weng X, Zhao Y, Tang W, Wang L et al (2008) Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40:761–767. https://doi.org/10.1038/ng.143
Yamagishi M, Takeuchi Y, Kono L, Yano M (2002) QTL analysis for panicle characteristics in temperate japonica rice. Euphytica 128:219–224
Yan WH, Wang P, Chen HX, Zhou HJ, Li QP, Wang CR et al (2011) A major QTL, Ghd8, plays pleiotropic roles in regulating grain productivity, plant height, and heading date in rice. Mol Plant 4:319–330. https://doi.org/10.1093/mp/ssq070
Yan W, Liu H, Zhou X, Li Q, Zhang J, Lu L et al (2013) Natural variation in Ghd7.1 plays an important role in grain yield and adaptation in rice. Cell Res 23:969–971. https://doi.org/10.1038/cr.2013.43
Yoshida A, Sasao M, Yasuno N, Takagi K, Daimon Y, Chen R et al (2013) TAWAWA1, a regulator of rice inflorescence architecture, functions through the suppression of meristem phase transition. Proc Natl Acad Sci USA 110:767–772. https://doi.org/10.1073/pnas.1216151110
Zeng D, Tian Z, Rao Y, Dong G, Yang Y, Huang L et al (2017) Rational design of high-yield and superior-quality rice. Nat Plants 3:1–5. https://doi.org/10.1038/nplants.2017.31
Zhang Y, Luo L, Xu C, Zhang Q, Xing Y (2006) Quantitative trait loci for panicle size, heading date and plant height co-segregating in trait-performance derived near-isogenic lines of rice (Oryza sativa L.). Theor Appl Genet 113:361–368. https://doi.org/10.1007/s00122-006-0305-3
Zhang D, Zhang H, Wang M, Sun J, Qi Y, Wang F et al (2009) Genetic structure and differentiation of Oryza sativa L. in China revealed by microsatellites. Theor Appl Genet 119:1105–1117. https://doi.org/10.1007/s00122-009-1112-4
Zhang H, Zhang D, Wang M, Sun J, Qi Y, Li J et al (2011) A core collection and mini core collection of Oryza sativa L. in China. Theor Appl Genet 122:49–61. https://doi.org/10.1007/s00122-010-1421-7
Funding
This research was supported by China National Key Technologies R&D Program (2015BAD02B01 and 2013BAD01B02-15).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Jianyin Xie, Fengmei Li, Najeeb Ullah Khan, Xiaoyang Zhu, Xueqiang Wang, Zhifang Zhang, Xiaoqian Ma, Yan Zhao, Quan Zhang, Shuyang Zhang, Zhanying Zhang, Jinjie Li, Zichao Li and Hongliang Zhang declare that they have no conflict of interest.
Research involving human and animal participants
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xie, J., Li, F., Khan, N.U. et al. Identifying natural genotypes of grain number per panicle in rice (Oryza sativa L.) by association mapping. Genes Genom 41, 283–295 (2019). https://doi.org/10.1007/s13258-018-0758-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-018-0758-1