Abstract
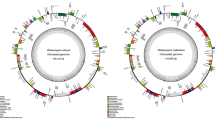
In this study, the chloroplast genome of Hariotina reticulata was fully sequenced and compared to other Sphaeropleales chloroplast genomes. It is 210,757 bp larger than most Sphaeropleales cpDNAs. It presents a traditional chloroplast structure, and contains 103 genes, including 68 protein-coding genes, six rRNA genes and 29 tRNA genes. The coding region constitutes of 43% of the whole cpDNA. Eighteen introns are found in 11 genes and six introns are unique for Hariotina. 11 open reading frames are identified among these introns. The synteny between Hariotina and Acutodesmus cpDNAs is in general identical, while within Sphaeropleales order, high variability in cpDNA architecture is indicated by general high DCJ distances. Ankyra judayi exhibits the greatest dissimilarity in gene synteny to the others and share some unique gene clusters with Treubaria triappendiculata. The phylogenomic analyses show that A. judayi is clustered with Treubariaceae species and sister to Chlorophyceae incertae sedis and other Sphaeropleales species. The monophyly of Sphaeropleales is rejected.



Similar content being viewed by others
References
Abe T, Ikemura T, Sugahara J, Kanai A, Ohara Y, Uehara H et al (2011) tRNADB-CE 2011: tRNA gene database curated manually by experts. Nucleic Acids Res 39:210–213. https://doi.org/10.1093/nar/gkq1007
An SS, Friedl T, Hegewald E (1999) Phylogenetic relationships of Scenedesmus and Scenedesmus-like coccoid green algae as inferred from ITS-2 rDNA sequence comparisons. Plant Biol 1:418–429
Besendahl A, Qiu YL, Lee J, Palmer JD, Bhattacharya D (2000) The cyanobacterial origin and vertical transmission of the plastid tRNALeu group-I intron. Curr Genet 37:12–23
Buchheim MA, Sutherland DM, Schleicher T, Forster F, Wolf M (2012) Phylogeny of Oedogoniales, Chaetophorales and Chaetopeltidales (Chlorophyceae): inferences from sequence-structure analysis of ITS2. Ann Bot 109:109–116
Caisová L, Marin B, Melkonian M (2013) A consensus secondary structure of ITS2 in the Chlorophyta identified by phylogenetic reconstruction. Protist 164:482–496
Cheng J, Zeng X, Ran G, Liu Z (2013) CGAP: a new comprehensive platform for the comparative analysis of chloroplast genomes. BMC Bioinform 14:95. https://doi.org/10.1186/1471-2105-14-95
Chodat R (1926) Scenedesmus. Étude de génétique, de systématique expérimentale et d’hydrobiologie. Rev Hydrobiol 3:71–258
Darriba D, Taboada GL, Doallo R, Posada D (2011) ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27:1164–1165
de Cambiaire JC, Otis C, Lemieux C, Turmel M (2006) The complete chloroplast genome sequence of the chlorophycean green alga Scenedesmus obliquus reveals a compact gene organization and a biased distribution of genes on the two DNA strands. BMC Evol Biol 6:37. https://doi.org/10.1186/1471-2148-6-37
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Eliáš M, Němcová Y, Škaloud P, Neustupa J, Kaufnerová V, Šejnohová L (2010) Hylodesmus singaporensis gen. et sp. nov., a new autosporic subaerial green alga (Scenedesmaceae, Chlorophyta) from Singapore. Int J Syst Evol Microbiol 60:1224–1235
Fučíková K, Lewis PO, Lewis LA (2016) Chloroplast phylogenomic data from the green algal order Sphaeropleales (Chlorophyceae, Chlorophyta) reveal complex patterns of sequence evolution. Mol Phylogenet Evol 98:176–183
Griffiths-Jones S, Bateman A, Marshall M, Khanna A, Eddy SR (2003) Rfam: an RNA family database. Nucleic Acids Res 31:439–441
Guiry MD, Guiry GM (2016) AlgaeBase. World-wide electronic publication. National University of Ireland, Galway
He LJ, Lou SL, Zhang F, Yang SJ, Zhang C, Lin XZ, Yang LH (2016) The complete mitochondrial DNA sequence of the green algae Hariotina sp. F30 (Scenedesmaceae, Sphaeropleales, Chlorophyceae). Mitochondrial DNA Part B: Resour 1:132–133
Hegewald E (1978) Eine neue Unterteilung der Gattung Scenedesmus Meyen. Nova Hedwig 30:343–376
Hegewald E (2000) New combinations in the genus Desmodesmus (Chlorophyceae, Scenedesmaceae). Algol Stud 96:1–18
Hegewald E, Silva PC (1988) Annotated catalogue of Scenedesmus and nomenclaturally related genera including original descriptions and figures. Bibl Phycol 80:1–587
Hegewald E, Wolf M, Keller A, Friedl T, Krienitz L (2010) ITS2 sequence-structure phylogeny in the Scenedesmaceae with special reference to Coelastrum (Chlorophyta, Chlorophyceae), including the new genera Comasiella and Pectinodesmus. Phycologia 49:325–335
Hilker R, Sickinger C, Pedersen CNS, Stoye J (2012) UniMoG—a unifying framework for genomic distance calculation and sorting based on DCJ. Bioinformatics 28:2509–2511
Keller A, Schleicher T, Förster F, Ruderisch B, Dandekar T, Müller T, Wolf M (2008) ITS2 data corroborate a monophyletic chlorophycean DO-group (Spaheropleales). BMC Evol Biol 8:218. https://doi.org/10.1186/1471-2148-8-218
Kessler E, Schäfer M, Hümmer C, Kloboucek A, Huss VAR (1997) Physiological, biochemical, and molecular characters for the taxonomy of the subgenera of Scenedesmus (Chlorococcales, Chlorophyta). Bot Acta 110:244–250
Krienitz L, Bock C (2012) Present state of the systematic of planktonic coccoid green algae of inland waters. Hydrobiologia 698:295–326
Lohse M, Drechsel O, Bock R (2007) Organellar Genome DRAW (OGDRAW): a tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr Genet 52:267–274
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, HÓ§hna S et al (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:W686–W689
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Trainor FR, Cain JR, Shubert EL (1976) Morphology and nutrition of the colonial green alga Scenedesmus: 80 years later. Bot Rev 42:5–25
Turmel M, Otis C, Lemieux C (2015) Dynamic evolution of the chloroplast genome in the green algal classes Pedinophyceae and Trebouxiophyceae. Genome Biol Evol 7:2062–2080
Van Hannen EJ, Fink P, Lürling M (2002) A revised secondary structure model for the internal transcribed spacer 2 of the green algae Scenedesmus and Desmodesmus and its implication for the phylogeny of these algae. Eur J Phycol 37:203–208
Vanormelingen P, Hegewald E, Braband A, Kitschke M, Friedl T, Sabbe K, Vyverman W (2007) The systematics of small spineless Desmodesmus species, D. costato-granulatus (Sphaeropleales, Chlorophyceae), based on ITS2 rDNA sequence analyses and cell wall morphology. J Phycol 43:378–396
Wolf M, Buchheim M, Hegewald E, Krienitz L, Hepperle D (2002) Phylogenetic position of the Sphaeropleaceae (Chlorophyta). Plant Syst Evol 230:161–171
Wyman SK, Jansen RK, Boore JL (2004) Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20:3252–3255
Acknowledgements
This study was supported by Demonstration Project of Marine Economy Innovative Development Project (Nos. 12PYY001SF08, 16PYY005SF09), Natural Science Foundation of Fujian Province (No. 2016J05100), Xiamen Southern Oceanographic Center (No. 14CZP028HJ02) and the Xiamen Science and Technology Project (No. 3502Z20172017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Lijuan He declares that she does not have conflict of interest. Zhaokai Wang declares that he does not have conflict of interest. Sulin Lou declares that she does not have conflict of interest. Xiangzhi Lin declares that he does not have conflict of interest. Fan Hu declares that he does not have conflict of interest.
Research involving with human and animal participants
This article does not contain any animal or human subjects. The research material is not in the endangered species list.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
He, L., Wang, Z., Lou, S. et al. The complete chloroplast genome of the green algae Hariotina reticulata (Scenedesmaceae, Sphaeropleales, Chlorophyta). Genes Genom 40, 543–552 (2018). https://doi.org/10.1007/s13258-018-0652-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-018-0652-x