Abstract
Purpose
Although segmentation of Abdominal Aortic Aneurysms (AAA) thrombus is a crucial step for both the planning of endovascular treatment and the monitoring of the intervention’s outcome, it is still performed manually implying time consuming operations as well as operator dependency. The present paper proposes a fully automatic pipeline to segment the intraluminal thrombus in AAA from contrast-enhanced Computed Tomography Angiography (CTA) images and to subsequently analyze AAA geometry.
Methods
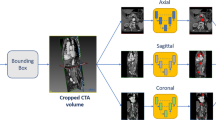
A deep-learning-based pipeline is developed to localize and segment the thrombus from the CTA scans. The thrombus is first identified in the whole sub-sampled CTA, then multi-view U-Nets are combined together to segment the thrombus from the identified region of interest. Polygonal models are generated for the thrombus and the lumen. The lumen centerline is automatically extracted from the lumen mesh and used to compute the aneurysm and lumen diameters.
Results
The proposed multi-view integration approach returns an improvement in thrombus segmentation with respect to the single-view prediction. The thrombus segmentation model is trained over a training set of 63 CTA and a validation set of 8 CTA scans. By comparing the thrombus segmentation predicted by the model with the ground truth data, a Dice Similarity Coefficient (DSC) of 0.89 ± 0.04 is achieved. The AAA geometry analysis provided an Intraclass Correlation Coefficient (ICC) of 0.92 and a mean-absolute difference of 3.2 ± 2.4 mm, for the measurements of the total diameter of the aneurysm. Validation of both thrombus segmentation and aneurysm geometry analysis is performed over a test set of 14 CTA scans.
Conclusion
The developed deep learning models can effectively segment the thrombus from patients affected by AAA. Moreover, the diameters automatically extracted from the AAA show high correlation with those manually measured by experts.
Graphical abstract








Similar content being viewed by others
Data Availability
None.
Code Availability
None.
References
Antiga, L., M. Piccinelli, L. Botti, B. Ene-Iordache, A. Remuzzi, and D. A. Steinman. An image-based modeling framework for patient-specific computational hemodynamics. Med. Biol. Eng. Comput. 46(11):1097–1112, 2008. https://doi.org/10.1007/s11517-008-0420-1.
Arash Salarian, Intraclass Correlation Coefficient (icc). https://www.mathworks.com/matlabcentral/fileexchange/22099-intraclass-correlation-coefficient-icc
Caradu, C., B. Spampinato, A. M. Vrancianu, X. Bérard, and E. Ducasse. Fully automatic volume segmentation of infrarenal abdominal aortic aneurysm computed tomography images with deep learning approaches versus physician controlled manual segmentation. J. Vasc. Surg. 74(1):246-256.e6, 2021. https://doi.org/10.1016/j.jvs.2020.11.036.
Chaikof, E. L., et al. The Society for Vascular Surgery practice guidelines on the care of patients with an abdominal aortic aneurysm. J. Vasc. Surg. 67(1):2-77.e2, 2018. https://doi.org/10.1016/j.jvs.2017.10.044.
Chollet, Francois et al. Keras. 2015. https://keras.io
Developers, TensorFlow. TensorFlow. Zenodo 2021. https://doi.org/10.5281/ZENODO.5043456.
Fantazzini, A., et al. 3D automatic segmentation of aortic computed tomography angiography combining multi-view 2D Convolutional Neural Networks. Cardiovasc. Eng. Technol. 11(5):576–586, 2020. https://doi.org/10.1007/s13239-020-00481-z.
Haller, S. J., et al. Intraluminal thrombus is associated with early rupture of abdominal aortic aneurysm. J. Vasc. Surg. 67(4):1051-1058.e1, 2018. https://doi.org/10.1016/j.jvs.2017.08.069.
Hong, H. A. and U. U. Sheikh. Automatic detection, segmentation and classification of abdominal aortic aneurysm using deep learning. In 2016 IEEE 12th International Colloquium on Signal Processing & Its Applications (CSPA), Melaka, Malaysia, Mar. 2016, pp. 242–246. https://doi.org/10.1109/CSPA.2016.7515839.
Kaladji, A., A. Lucas, G. Kervio, P. Haigron, and A. Cardon. Sizing for endovascular aneurysm repair: clinical evaluation of a new automated three-dimensional software. Ann. Vasc. Surg. 24(7):912–920, 2010. https://doi.org/10.1016/j.avsg.2010.03.018.
Lareyre, F., C. Adam, M. Carrier, C. Dommerc, C. Mialhe, and J. Raffort. A fully automated pipeline for mining abdominal aortic aneurysm using image segmentation. Sci. Rep. 9(1):13750, 2019. https://doi.org/10.1038/s41598-019-50251-8.
López-Linares, K., et al. Fully automatic detection and segmentation of abdominal aortic thrombus in post-operative CTA images using Deep Convolutional Neural Networks. Med. Image Anal. 46:202–214, 2018. https://doi.org/10.1016/j.media.2018.03.010.
López-Linares, K., I. García, A. García-Familiar, I. Macía, and M. A. G. Ballester. 3D convolutional neural network for abdominal aortic aneurysm segmentation. 2019. https://doi.org/10.1016/j.media.2018.03.010.
Lorensen, W. E., and H. E. Cline. Marching cubes: A high resolution 3D surface construction algorithm. ACM SIGGRAPH Comput. Graph. 21(4):163–169, 1987. https://doi.org/10.1145/37402.37422.
Maiora, J., B. Ayerdi, and M. Graña. Random forest active learning for AAA thrombus segmentation in computed tomography angiography images. Neurocomputing. 126:71–77, 2014. https://doi.org/10.1016/j.neucom.2013.01.051.
Maurer, C. R., R. Qi, and V. Raghavan. A linear time algorithm for computing exact Euclidean distance transforms of binary images in arbitrary dimensions. IEEE Trans. Pattern Anal. Mach. Intell. 25(2):265–270, 2003. https://doi.org/10.1109/TPAMI.2003.1177156.
McCormick, M., X. Liu, J. Jomier, C. Marion, and L. Ibanez. ITK: enabling reproducible research and open science. Front. Neuroinform. 2014. https://doi.org/10.3389/fninf.2014.00013.
Óleary, S. A., E. G. Kavanagh, P. A. Grace, T. M. McGloughlin, and B. J. Doyle. The biaxial mechanical behaviour of abdominal aortic aneurysm intraluminal thrombus: classification of morphology and the determination of layer and region specific properties. J. Biomech. 47(6):1430–1437, 2014. https://doi.org/10.1016/j.jbiomech.2014.01.041.
Ronneberger, O., P. Fischer, and T. Brox, ‘U-Net: Convolutional Networks for Biomedical Image Segmentation’, ArXiv150504597 Cs, May 2015, doi: https://doi.org/10.1007/978-3-319-24574-4_28.
Schroeder, W., K. Martin, and B. Lorensen. The visualization toolkit: an object-oriented approach to 3D graphics; visualize data in 3D—medical, engineering or scientific; build your own applications with C++, Tcl, Java or Python; includes source code for VTK (supports Unix, Windows and Mac), 4th ed. Clifton Park, NY: Kitware Inc, 2006.
Shen, D., G. Wu, and H.-I. Suk. Deep learning in medical image analysis. Annu. Rev. Biomed. Eng. 19(1):221–248, 2017. https://doi.org/10.1146/annurev-bioeng-071516-044442.
Singh, K., et al. Intra- and interobserver variability in the measurements of abdominal aortic and common iliac artery diameter with computed tomography. The Tromsø study. Eur. J. Vasc. Endovasc. Surg. 25(5):399–407, 2003. https://doi.org/10.1053/ejvs.2002.1856.
Wolf, I., et al. The medical imaging interaction toolkit. Med. Image Anal. 9(6):594–604, 2005. https://doi.org/10.1016/j.media.2005.04.005.
Zhang, Y., Q. Liao, and J. Zhang. Exploring efficient volumetric medical image segmentation using 2.5D method: an empirical study. ArXiv201006163 Cs Eess, 2020. http://arxiv.org/abs/2010.06163
Funding
This study did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None.
Additional information
Associate Editor Igor Efimov oversaw the review of this article.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Brutti, F., Fantazzini, A., Finotello, A. et al. Deep Learning to Automatically Segment and Analyze Abdominal Aortic Aneurysm from Computed Tomography Angiography. Cardiovasc Eng Tech 13, 535–547 (2022). https://doi.org/10.1007/s13239-021-00594-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13239-021-00594-z