Abstract
Ebola virus (EBOV) is an enveloped negative-sense RNA virus and a member of the filovirus family. Nucleoprotein (NP) expression alone leads to the formation of inclusion bodies (IBs), which are critical for viral RNA synthesis. The matrix protein, VP40, not only plays a critical role in virus assembly/budding, but also can regulate transcription and replication of the viral genome. However, the molecular mechanism by which VP40 regulates viral RNA synthesis and virion assembly/budding is unknown. Here, we show that within IBs the N-terminus of NP recruits VP40 and is required for VLP-containing NP release. Furthermore, we find four point mutations (L692A, P697A, P698A and W699A) within the C-terminal hydrophobic core of NP result in a stronger VP40–NP interaction within IBs, sequestering VP40 within IBs, reducing VP40–VLP egress, abolishing the incorporation of NC-like structures into VP40–VLP, and inhibiting viral RNA synthesis, suggesting that the interaction of N-terminus of NP with VP40 induces a conformational change in the C-terminus of NP. Consequently, the C-terminal hydrophobic core of NP is exposed and binds VP40, thereby inhibiting RNA synthesis and initiating virion assembly/budding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Ebola virus (EBOV) is an enveloped non-segmented negative strand RNA virus (NNSV) and a member of the filovirus family, which consists of three genera, Ebolavirus, Marburgvirus, and Cuevavirus. The species Zaire, Sudan, Bundibugyo, Tai Forest, and Reston ebolavirus are found within the genus of Ebolavirus. Marburgvirus and Cuevavirus each include one species: Marburg virus (MARV) and Lloviu virus (LLOV), respectively (Negredo et al., 2011; Misasi and Sullivan, 2014; Martin et al., 2016; Yang et al., 2019). EBOV can cause a severe fever with a high fatality rate. The outbreak of EBOV in the Democratic Republic Congo from August 2018 to November 2019 caused 3,296 infections and 2,196 deaths (67%), making it the second largest outbreak since the 2014–2016 outbreak in West Africa (Hartman et al., 2010; Messaoudi et al., 2015; Vetter et al., 2016; Aruna et al., 2019; Hoenen et al., 2019). The EBOV genome is approximately 19 kb in length, encoding seven structural proteins: nucleoprotein (NP), cofactor of polymerase L (VP35), matrix protein (VP40), glycoprotein (GP), transcription activator (VP30), minor matrix protein (VP24), and RNA-dependent RNA polymerase (L) (Kirchdoerfer et al., 2017).
The viral genome of EBOV is encapsidated by NP to form an NP–RNA template that then interacts with the RNA-dependent RNA polymerase complex consisting of VP35, VP30, and L to initiate viral transcription and replication (Muhlberger et al., 1998; Weik et al., 2002; Hartlieb et al., 2007; Groseth et al., 2009; Jasenosky et al., 2010; Trunschke et al., 2013). Similar to many negative strand RNA viruses (NSVs), EBOV can form IBs, which are the viral RNA synthesis factories in infected cells (Hoenen et al., 2012; Nanbo et al., 2013). For some NNSVs, such as vesicular stomatitis virus (VSV), human respiratory syncytial virus (hRSV), and human parainfluenza virus type 3 (HPIV3), the co-expression of NP and phosphoprotein (P) is the minimum requirement for the formation of viral IBs (Garcia-Barreno et al., 1996; Heinrich et al., 2010; Zhang et al., 2013; Richard et al., 2018). By contrast, regarding EBOV, NP expression alone can lead to the formation of IBs (Nelson et al., 2016). Moreover, NP along with VP35 and VP24 are indispensable for the formation of nucleocapsid (NC)-like structures (Huang et al., 2002; Noda et al., 2007a; Shi et al., 2008; Bharat et al., 2012). In addition, a study suggests that NC is formed at the periphery of the IBs, which indicates that NP can recruit other viral proteins for viral replication and NC assembly (Nanbo et al., 2013). This suggests that NP plays a critical role in the lifecycle of EBOV. Like the NP of other NNSVs, the Ebola NP consists of a highly conserved N-terminus, which is critical for NP oligomerization and RNA binding, and a hypervariable C-terminus, which is involved in the association with VP40 and the incorporation of NCs into virions. The C-terminus of NP has no homology with any known proteins, which suggests that the NP C-terminus plays a critical role in protein–protein interactions (Buchholz et al., 1993; Longhi et al., 2003; Watanabe et al., 2006; Noda et al., 2007b; Noda et al., 2010; Dziubanska et al., 2014; Zhang et al., 2017).
Assembly and budding are key steps in the viral lifecycle. For many enveloped viruses, the virus-like particle (VLP) system is useful for studying the mechanisms of viral assembly and budding. The matrix (M) protein is indispensable for the formation and release of VLPs. For most NNSVs, like HPIV1 (Coronel et al., 1999), Sendai virus (SeV) (Sakaguchi et al., 1999; Sugahara et al., 2004), Measles virus (MeV) (Runkler et al., 2007), Nipah virus (NiV) (Patch et al., 2007), HPIV3 (Zhang et al., 2014), and VSV (Li et al., 1993; Justice et al., 1995), expression of the M protein alone can direct budding and formation of VLPs. However, in the case of paramyxovirus simian virus 5 (SV5) M protein, NP and fusion protein (F) or hemagglutinin-neuraminidase (HN) are required for the formation and release VLPs (Schmitt et al., 2002). VP40 is the major matrix protein of EBOV; when expressed alone, it can be released into the culture medium as a type of VLP (Jasenosky et al., 2001; Timmins et al., 2001).
The M proteins of many NNSVs can regulate viral replication and/or transcription. The purified M protein of VSV condenses the nucleocapsid into a tight structure causing in vitro inhibition of viral transcription (De et al., 1982). The M protein of MeV retains the RNP complex at the plasma membrane by interacting with NP, thus inhibiting RNA synthesis and facilitating viral assembly and budding (Iwasaki et al., 2009). The M protein of HPIV3 suppresses the formation of IBs by interacting with NP, thereby reducing viral replication (Zhang et al., 2018). A previous study showed that VP40 inhibits replication and transcription of the Ebola minigenome (Hoenen et al., 2010). However, the detailed mechanism by which VP40 regulates viral RNA synthesis and NC assembly/budding remains largely unknown. In addition, from the perspective of virus evolution, in EBOV infected cells, when the virus synthesizes enough elements (RNAs and proteins) for packaging, the virus should shut down the synthesis of RNAs and proteins, and concentrate on the assembly/budding of virus particles to achieve the purpose of virion amplification. However, the molecular mechanism of how viruses transform from RNA synthesis to viral particle assembly is not clear.
In this study, we show that VP40 is first recruited to IBs by association with the N-terminus of NP, which results in a conformational change and exposure of the hydrophobic core within the NP C-terminus consisting of L692, P697, P698, and W699. Consequently, the exposed hydrophobic core within the NP C-terminus interacts with VP40 for the incorporation of NC-like structures into the VLPs of VP40. When the L692, P697, P698, and W699 within the hydrophobic core are mutated into alanines, stronger VP40–NP interactions occur, and the budding of VP40 is inhibited by sequestration of VP40 into IBs. Furthermore, we found that the hydrophobic core is critical for NP encapsidating viral RNA, and VP40 inhibits viral replication and transcription by association with this domain. These results suggest that the two-stage interaction of VP40 with NP plays a critical role in the transition from RNA synthesis to NC assembly/budding.
Results
N-terminal amino acids 26 to 150 of the NP are required for interactions with VP40 and for NP incorporation into VP40–VLP
Given the essential role played by NP and VP40 in EBOV RNA synthesis and NC assembly/budding, we first sought to study the relationship between VP40 and NP. We performed co-immunoprecipitation (coIP) and VLP budding assays, and the results showed that VP40 can interact with NP and incorporate NP into VP40–VLP, which was confirmed by the protease protection assay (Fig. S1A–C).
A previous study showed that both the N- and C-termini of NP are important for interaction with VP40, and a 150 aa region within the NP N-terminus was found to be responsible for mediating the interaction with VP40 (Noda et al., 2007b). Based on this result, we sought to narrow down the specific amino acids within this region responsible for interaction, and generated two NP truncated mutants with an HA tag at the N-terminus (Leung et al., 2015; Su et al., 2018). We found that HA-NP∆N25 can interact with VP40-Flag as efficiently as WT NP, whereas the mutant HA-NP∆N26–150 is barely co-immunoprecipitated by VP40-Flag (Fig. 1A, lanes 2, 4, and 6). Meanwhile, Myc-VP40 can incorporate HA-NP∆N25 but not HA-NP∆N26–150 into VLPs, which is consistent with the coIP result (Fig. 1B, bottom panel, lanes 4 and 6). In addition, we observed that NP∆N26–150 still maintains its associations with VP35 and VP30 (Fig. 1C, lane 4 and 1D, lane 3), which excludes the possibility that the structure of NP∆N26–150 has been changed and can no longer be recognized by VP40. Next, to further narrow down the region within the N-terminus of NP involved in the interaction with VP40, we constructed a series of NP N-terminal truncated mutants based on a previous study (Sugita et al., 2018). We found that all the mutants with shorter truncations could be co-immunoprecipitated and incorporated into VLP s by VP40 (Fig. 1E–H), suggesting that more than one domain within amino acids 26 to 150 in the NP N-terminus is involved in the association of NP–VP40 and the incorporation of NP into VP40–VLP.
N-terminal amino acids 26 to 150 of the NP are required for interactions with VP40 and for NP incorporation into VP40–VLP. (A) HEK293T cells were co-transfected with HA-NP/NP N-terminal mutants and VP40-Flag. At 30 h p.t, lysates were subjected to IP with anti-Flag antibodies and analyzed via Western blot (WB). (B) HEK293T cells were co-transfected with HA-NP/NP N-terminal mutants and Myc-VP40 for 36 h. Cell lysates and VLPs from supernatants were subjected to WB with anti-Myc and anti-HA antibodies. (C and D) HEK293T cells were co-transfected with HA-NP/NP∆N26–150 and Flag-VP35 or Flag-VP30. Lysates were subjected to IP with anti-Flag antibodies and analyzed via WB. (C) The coIP of VP35 with NP∆N26–150; (D) the coIP of VP30 with NP∆N26–150. (E and F) HEK293T cells were transfected with the indicated plasmids. Lysates were subjected to IP with anti-Flag antibodies and analyzed via WB. (E) The coIP of VP40 with NP∆N26–38; (F) the coIP of VP40 with NP∆N39–90, NP∆N91–120, or NP∆N121–150. (G and H) HEK293T cells were transfected with the indicated plasmids. Cell lysates and VLPs from supernatants were subjected to WB. (G) The VLP budding assay of VP40 with NP∆N26–38; (H) the VLP budding assay of VP40 with NP∆N39–90, NP∆N91–120, or NP∆N121–150
Because NP expression alone can induce the formation of IBs, we sought to examine whether the expression of the above NP N-terminal mutants can also form IBs. Toward this aim, HeLa cells were transfected with NP or N-terminal mutants. We also observed that the mutants NP∆N25 and NP∆N26–38 could aggregate in the cytoplasm, similar to those formed by NP. However, the mutants NP∆N39–90, NP∆N91–120, and NP∆N121–150 were all homogeneously distributed in the cytoplasm (Fig. S1D). To determine whether the failure of NP mutants to form IBs is related to their oligomerization, we performed a coIP assay. As expected, HA-NP∆N25 and HA-NP∆N26–38 interacted with Myc-NP, while HA-NP∆N39–90, HA-NP∆N91–120, and HA-NP∆N121–150 failed to interact with Myc-NP (Fig. S1E–F), suggesting that amino acids 39 to 150 in the N-terminus are critical for the formation of IBs and the oligomerization of NP.
Taken together, our results demonstrate that N-terminal amino acids 26 to 150 are critical for the interaction of VP40 and NP as well as the incorporation of NP into VP40–VLP.
Incorporation of NP into VP40–VLP via the NP C-terminal hydrophobic core
We thus established that the NP N-terminus is required for the incorporation of NP into VP40–VLP. In addition, a previous study showed that NP C-terminal 50 amino acids are critical for NP’s incorporation into VP40–VLP (Licata et al., 2004). Next, we sought to determine how the N- and C-termini of NP participate in the incorporation of NP into VP40–VLP. Using a VLP budding assay, we first confirmed that a truncated NP whereby the last 50 amino acids in the C-terminus were deleted (NP∆C50) failed to incorporate NP into VP40–VLP (Fig. 2A, bottom panel, lane 4). Moreover, we found that NP∆C50 still maintains the associations with VP35 and VP30 (Fig. S2A and S2B, lanes 4, 3), suggesting that the structure of NP∆C50 is intact and undamaged. Then, to narrow down the specific amino acids within this 50 amino acid region responsible for incorporation of NP into VP40–VLP, we generated five other mutants by sequential deletion, whereby each contained 10 amino acids as shown in Fig. S2C, and found that for the same level of expression, NP∆C41–50 clearly lost the ability to be incorporated into VP40–VLP (Fig. 2B, bottom panel, lane 12), suggesting that amino acids 690 to 699, located at the NP C-terminus, are critical for the incorporation of NP into VP40–VLP (Fig. 2C). To further explore the critical motif for NP incorporation into VP40–VLP in amino acids 690 to 699, we constructed three different triple or quadruple-point mutants A1 (NP690–692AAA), A2 (NP693–695AAA), and A3(NP696–699AAAA) (Fig. 2C) and found that neither A1 nor A3 could be incorporated into VP40–VLP, in contrast to A2 (Fig. 2D, lanes 6, 8, and 10). To determine the precise binding site(s), we generated seven point mutants containing unique alanine mutations within A1 and A3 (Fig. 2C), and these mutants were used for the VLP assay. We found that, similar to A1 and A3, four mutants (NPL692A, NPP697A, NPP698A, and NPW699A) could not be incorporated into VP40–VLP (Fig. 2E, lanes 4, 10 and 2F, lanes 4, 8, 10 and 12). Sequence alignments show that L692, P697, P698, and W699 are relatively conserved among the filovirus family (Fig. S2D) and exist in NP as side chains which constitute a hydrophobic core, according to an analysis of NP C-terminal structure (PDB: 4QB0) (Fig. S2E). Taken together, these results show that four sites (L692, P697, P698, and W699) of the hydrophobic core in the NP C-terminus are indispensable for incorporation of NP into VP40–VLP.
Incorporation of NP into VP40–VLP via the NP C-terminal hydrophobic core. (A) HEK293T cells were co-transfected with HA-NP/NP∆C50 and Myc-VP40 for 36 h. Cell lysates and VLPs from supernatants were subjected to WB analysis. (B) VLP budding assay for the co-transfection of HA-NP/NP C-terminal truncated mutants with Myc-VP40. HEK293T cells were transfected with the indicated plasmids for 36 h. Cell lysates and VLPs were analyzed via WB. (C) Schematic diagrams of C-terminally truncated or point mutants of NP. (D–F) VLP budding assay of HA-NP/NP C-terminal point mutants with Myc-VP40. HEK293T cells were transfected with the indicated plasmids for 36 h. Cell lysates and VLPs were analyzed via WB. (G) coIP assay of the VP40-Flag with HA-NP/NPL692A, NPP697A, NPP698A, or NPW699A. HEK293T cells were transfected with the indicated plasmids. At 30 h p.t, cell lysates were subjected to IP with anti-Flag antibodies and then analyzed via WB. (H) HEK293T cells were treated as above. The interactions of HA-NP/NPL692A, NPP697A, NPP698A, or NPW699A with VP40-Flag were quantified using the Quantity One software. (I) The colocalization of VP40 with NP/NPL692A, NPP697A, NPP698A, or NPW699A. HeLa cells were co-transfected with Myc-VP40 and HA-NP/NP mutants for 26 h. Cells were immunostained for VP40 (red) and NP or NP mutants (green) and analyzed using superresolution microscopy (Zeiss LSM 800). Scale bars = 5 μm. Images are representative of three independent experiments. Error bars, mean ± SD of three experiments (n = 3). Student’s t-test; *P < 0.05; **P < 0.01; ***P < 0.001; ns, non-significant
Because NP–VP40 interactions are a prerequisite for NP incorporation into VP40–VLP, and having found that NPL692A, NPP697A, NPP698A, and NPW699A cannot be incorporated into VP40–VLP, we sought to determine whether these mutants fail to interact with VP40. However, to our surprise, we found that all these mutants significantly enhanced interactions with VP40 compared to WT NP at the same level of expression (Fig. 2G–H). A previous study showed that VP40 can be recruited into IBs in EBOV-infected cells (Nanbo et al., 2013). Thus, we sought to determine whether the expression of these mutants could form IBs, and if so, whether the IBs formed by these mutants could recruit VP40. The results showed that the IBs formed by the expression of NPL692A, NPP697A, NPP698A, and NPW699A are similar in size and morphology to those of WT NP (Fig. S2F). When VP40 was expressed alone, it was homogeneously distributed in the cytoplasm and nucleus, and we observed a filamentous structure on the plasma membrane (Fig. S2G). We also found that VP40 was, indeed, recruited into the IBs but did not change the size and morphology of the IBs when co-expressed with WT NP. However, when VP40 was co-expressed with NPL692A, NPP697A, NPP698A, or NPW699A, VP40 was not only recruited into IBs but also resulted in the aggregation of IBs which were significantly larger in size than those formed by the co-expression of NP and VP40 (Fig. 2I). VP35 or VP30 could also colocalize with IBs but had no effect on the size and morphology of the IBs formed by NPL692A, NPP697A, NPP698A, or NPW699A (Fig. S2H and S2I). Taken together, these results show that both the N- and C-termini of NP are required for the incorporation of NP into VP40–VLP.
NPL692A, NPP697A, NPP698A, and NPW699A inhibit the release of VP40–VLP
Since VP40 has stronger interactions with NPL692A, NPP697A, NPP698A, and NPW699A and induces the aggregation of IBs but fails to incorporate these mutants into VLPs, we hypothesized that the interaction of the NP N-terminus with VP40 is a prerequisite for the incorporation of NP into VP40–VLP and only contributes to recruiting VP40 into IBs. If only the N-terminal interaction with VP40 is preserved, and the C-terminal interaction of NP with VP40 is deprived, VP40 will be trapped in the IBs, thereby inhibiting the assembly/release of VP40–VLP. To test this, we co-expressed VP40 with NPL692A, NPP697A, NPP698A, or NPW699A and performed a VLP budding assay. As expected, VP40 could be efficiently detected in the VLPs when co-expressed with NP. However, the release of VP40–VLP was reduced by 2–4 fold in the case of NPL692A, NPP697A, NPP698A, or NPW699A (Fig. 3A and 3B). NP∆N26–150, however, no longer interacted with VP40 and, therefore, had no effect on the release of VP40–VLP (Fig. 3C and 3D). Furthermore, we found that the numbers of cells containing filamentous structures in VP40-expressing cells were dramatically decreased in the presence of NPL692A, NPP697A, NPP698A, or NPW699A (Fig. 3E), suggesting that more VP40 is sequestered in the IBs formed by NPL692A, NPP697A, NPP698A, or NPW699A, thereby inhibiting the release of VP40–VLP. Furthermore, by using transmission electron microscopy, we observed that many VP40–VLP containing NP are located around the plasma membrane (PM), but that fewer empty VP40–VLP could be observed around the PM when VP40 was co-expressed with NPL692A, NPP697A, NPP698A, or NPW699A (Fig. 3F). Taken together, our results show that four sites, L692, P697, P698, and W699, located in the hydrophobic core of the C-terminus of NP, are indispensable for the assembly/budding of NP–VP40 VLPs, while NPL692A, NPP697A, NPP698A and NPW699A inhibit the release of VP40–VLP by sequestering VP40 into IBs.
NPL692A, NPP697A, NPP698A, and NPW699A inhibit the release of VP40–VLP. (A and B) VLP budding assay of VP40 in the presence of NP/NPL692A, NPP697A, NPP698A, or NPW699A. HEK293T cells were co-transfected with Myc-VP40 and HA-NP/NP mutants for 36 h. The VLP budding assay was performed as mentioned in Fig. 2A (A); the ability of VP40–VLP to be released was quantified using Quantity One software (B). (C and D) VLP budding assay of VP40 in the presence of NP/NP∆N26–150. HEK293T cells were co-transfected with Myc-VP40 and HA-NP/NP∆N26–150 for 36 h. (C) The VLP budding assay was performed as mentioned in Fig. 2A; (D) the ability of VP40–VLP egress was quantified using Quantity One software. (E) Budding cells under VP40 co-transfection with NP/NPL692A, NPP697A, NPP698A, or NPW699A. HeLa cells were treated as mentioned in Fig. 2I. The cells with filamentous structures were quantified (n = 3, 100 cells were counted). Images are representative of three independent experiments. Error bars, mean ± SD of three experiments (n = 3). Student’s t-test; *P < 0.05; **P < 0.01; ***P < 0.001; ns, non-significant. (F) Budding assay of VP40 via transmission electron microscopy. HEK293T cells were transfected with the indicated plasmids for 36 h. Budding cells were observed via electron microscopy. Black triangles indicate VLPs containing NP; black arrows indicate empty VP40–VLP. Scale bars = 500 nm
NPL692A, NPP697A, NPP698A, and NPW699A inhibit the migration of VP40 to the plasma membrane
The efficient egress of VP40 requires its transportation to the PM, where it finishes self-assembly/budding (Bornholdt et al., 2013). Since NPL692A, NPP697A, NPP698A, or NPW699A sequester VP40 into IBs, we hypothesized that alterations in VP40 localization would correlate with budding inhibition. To test this, live-cell imaging was performed. For this, we co-expressed mCherry-VP40 and GFP-NP or GFP-NPW699A in HeLa cells, and found that VP40 colocalized with IBs formed by NP; these IBs were able to co-migrate to the PM. Although VP40 could also colocalize with the IBs induced by NPW699A, their ability to migrate to the PM was inhibited (Fig. 4A; Video. S1A and S1B). Furthermore, we utilized a biochemical approach to determine whether the levels of VP40 in the PM fraction could be reduced in the presence of NPL692A, NPP697A, NPP698A, or NPW699A. For this, HEK293T cells were transfected with either VP40 and NP, or VP40 in combination with one of these mutants, and we found that when normalized to equivalent levels in the cytoplasm, the levels of VP40 in the PM fraction were reduced by 2–3 fold in the presence of NPL692A, NPP697A, NPP698A, or NPW699A compared to the levels in the presence of NP (Fig. 4B and 4C). Taken together, these results show that NPL692A, NPP697A, NPP698A, and NPW699A inhibit the migration of VP40 migration to the PM by sequestering VP40 into IBs.
NPL692A, NPP697A, NPP698A, and NPW699A inhibit the migration of VP40 to the plasma membrane. (A) The kinetic progress of IB migration to the PM. HeLa cells were cultured on 20 mm dishes and co-transfected with mCherry-VP40 and GFP-NP or GFP-NPW699A, together with their respective functional proteins, to exclude the effect of fluorescence tags. Live-cell imaging analysis was started at 24 h post-transfection. The pictures show the maximum intensity projection of time-lapse images of the cells. Images were captured every 120 s. Scale bar = 5 μm. (B and C) Isolation of cytosol and plasma membrane (PM) fractions. HEK293T cells were transfected with the indicated plasmids for 24 h. The cytosol and PM fractions were isolated and then subjected to a WB assay. GAPDH and Na/K ATPase served as controls for the cytosol or the PM fraction, respectively (B); quantification of the amount of PM-associated VP40 is shown (C). (D and E) The impact of NPP697A on the interaction of VP40 with Sec24C. HEK293T cells were co-transfected with VP40-Flag, HA-Sec24C, and Myc-NP/NPP697A for 30 h. IP was performed with anti-HA antibodies (D); quantification of the ability of VP40 binding Sec24C in the presence of NP/NPP697A is shown (E). Images are representative of three independent experiments. Error bars, mean ± SD of three independent experiments (n = 3). Student’s t-test; *P < 0.05; **P < 0.01; ***P < 0.001; ns, non-significant
A previous study demonstrated that VP40 can interact with Sec24C, a component of the COPII transport system which is critical for the intracellular transport of VP40 to the PM (Yamayoshi et al., 2008). Therefore, we sought to determine whether these mutants affect VP40–Sec24C interaction. We chose NPP697A and performed a coIP assay and found that NPP697A had no effect on VP40–Sec24C interaction (Fig. 4D, upper panel, lanes 2, 4 and 4E), suggesting that the NP–VP40 complex may use a different mechanism for transport, which requires further investigation.
The NP C-terminal hydrophobic core is critical for the incorporation of nucleocapsid-like structures into VP40–VLP
We found that VP40 incorporated NP into VLPs via the NP C-terminal hydrophobic core, and a previous study also showed that VP35 could be incorporated into VP40–VLP (Johnson et al., 2006). Therefore, we sought to determine what would happen when NP, VP35, and VP40 were co-expressed. We found that NP or VP35 could be efficiently incorporated into VP40–VLP (Fig. 5A, lanes 2 and 15), while the NP C-terminal mutants NPL692A, NPP697A, NPP698A, and NPW699A could not be incorporated into VP40–VLP in the presence of VP35. Furthermore, the incorporation of VP35 into VP40–VLP was also severely inhibited by NPL692A, NPP697A, NPP698A, and NPW699A (Fig. 5A, lanes 5, 8, 11 and 14), suggesting that the assembly of the NP–VP35 complex into VP40–VLP is mediated via the association of the NP hydrophobic core with VP40, rather than VP35. To confirm this, we performed coIP assay and found that NPL692A, NPP697A, NPP698A, and NPW699A interacted with VP35 as efficiently as NP (Fig. 5B, upper panel, lanes 2, 4, 6, 8, and 10). Moreover, VP40 did not affect the interactions of NPL692A, NPP697A, NPP698A, or NPW699A with VP35 (Fig. 5B, upper panel, lanes 4 to 11), but NPL692A, NPP697A, NPP698A, and NPW699A significantly inhibited the interaction of VP35 with VP40 (Fig. 5C, upper panel, lanes 2 to 6), suggesting that NP–VP35 complex is incorporated into VP40–VLP via NP–VP40 interaction.
The NP C-terminal hydrophobic core is critical for the incorporation of nucleocapsid-like structures into VP40–VLP. (A) VP40 assembles the NP–VP35 complex into VLPs via the NP C-terminal hydrophobic core. HEK293T cells were transfected with the indicated plasmids. Cell lysates and VLPs from the supernatants were subjected to WB assay with anti-Myc and anti-HA antibodies. (B) The effect of VP40 on the interactions of NPL692A, NPP697A, NPP698A, or NPW699A with VP35. HEK293T cells were transfected with the indicated plasmids, and IP was performed with the anti-HA antibodies. WB was performed with anti-Flag, anti-HA, and anti-Myc antibodies. (C) The effect of NPL692A, NPP697A, NPP698A, or NPW699A on the interaction of VP40 with VP35. HEK293T cells were transfected with the indicated plasmids. Cell lysates were subjected to IP with anti-Flag antibodies and analyzed via WB. (D–G) The association of the NP C-terminal hydrophobic core with VP40 mediates the NC-like structures assembly. NP or one of the C-terminal mutants with VP40, VP35, and VP24 were co-transfected into HEK293T cells. Cell lysates and VLPs from the supernatants were subjected to a WB assay with anti-Myc and anti-HA antibodies in the presence of NPL692A (D); in the presence of NPP697A (E); in the presence of NPP698A (F); and in the presence of NPW699A (G)
It has also been established that NP, VP35, and VP24 are the minimal requirements for the formation of NC-like structures of EBOV (Huang et al., 2002). Moreover, we found that VP24 can interact with VP40 and be incorporated into VLPs (Fig. S3A and S3B, lanes 2). Therefore, we sought to determine how VP40 assembles NC-like structures. For this, we expressed NP or one of the NP C-terminal mutants, VP35 and VP24, with VP40 and then performed VLP budding assays. The results show that VP40 could indeed incorporate the NP–VP35–VP24 complex into VLPs (Fig. 5D–G, lanes 7). However, the complex formed by one of the NP C-terminal mutants, VP35 and VP24, was not able to be incorporated into the VLPs (Fig. 5D–G, lanes 6). Taken together, these results show that the association of the NP C-terminal hydrophobic core with VP40 mediates assembly of the NC-like structures.
Interaction of VP40 with the NP C-terminal hydrophobic core prevents NP from encapsidating viral RNA
We established that the NP C-terminal hydrophobic core plays an essential role in the incorporation of NC-like structures into VP40–VLP. In addition, it was also reported that VP40 could regulate viral RNA synthesis (Hoenen et al., 2010), but the molecular mechanism through which VP40 regulates viral RNA is unclear. Therefore, we hypothesized that the hydrophobic core in the NP C-terminus may play an essential role in viral RNA synthesis and that its association with VP40 may inhibit viral RNA synthesis, thus causing a switch from viral RNA synthesis to virion assembly. To confirm this, we first performed an Ebola minigenome assay with increasing amounts of VP40 and found that VP40 inhibits viral RNA synthesis in a dose-dependent manner (Fig. 6A). Then, we further examined the RNA synthesis activity of NPL692A, NPP697A, NPP698A, and NPW699A and found that all four mutants barely support minigenome-encoded Renilla luciferase expression (Fig. 6B), suggesting that the NP C-terminal hydrophobic core is, indeed, indispensable for viral RNA synthesis.
Interaction of VP40 with the NP C-terminal hydrophobic core prevents NP from encapsidating viral RNA. (A) The effect of VP40 on Ebola minigenome RNA synthesis activity. HEK293T cells were co-transfected with plasmids encoding the minigenome assay components (NP, VP35, VP30, L, pBS-Rluc, Luc2, and T7) and increasing amounts of VP40. Renilla luciferase activity normalized to the firefly luciferase values was measured following the manufacturer’s instructions. The expression of VP40 was analyzed via WB. (B) The minigenome RNA synthesis ability of NPL692A, NPP697A, NPP698A, or NPW699A. Plasmids encoding the minigenome assay components (VP35, VP30, L, pBS-Rluc, T7, and Luc2) and Myc-NP/NP C-terminal point mutations were co-transfected into HEK293T cells. Renilla luciferase activity normalized to the firefly luciferase values was measured following the manufacturer’s instructions. The expression of Myc-NP and NP mutants was detected via WB. (C) The ability of NPL692A, NPP697A, NPP698A, or NPW699A binding minigenome RNA. Plasmids encoding the minigenome assay components (VP35, VP30, L, pBS-Rluc, and T7) and Myc-NP/NP C-terminal point mutations were co-transfected into HEK293T cells. Cell lysates were subjected to IP with anti-Myc antibodies, and then RNA was extracted, followed by RT-qPCR. The data are represented as relative scales, with the IP minigenome RNA normalized against the cell lysate’s minigenome RNA for each sample. The values were expressed as a percentage, where 100% was taken as abundance of immunoprecipitated Myc-NP. IP and cell lysate fractions were analyzed via WB. (D) VP40 inhibits NP binding the minigenome RNA. Plasmids encoding the minigenome assay components (Myc-NP, VP35, VP30, L, pBS-Rluc, and T7) and VP40-Flag were co-transfected into HEK293T cells. Cell lysates were subjected to IP with anti-Myc antibodies, and then RNA was extracted, followed by RT-qRCR. The data are represented as relative scales, with the IP minigenome RNA normalized against the cell lysate’s minigenome RNA for each sample. The values were expressed as a percentage, where 100% was taken as abundance of immunoprecipitated Myc-NP. IP and cell lysate fractions were analyzed via WB. Images are representative of three independent experiments. Error bars, mean ± SD of three independent experiments (n = 3). Student’s t-test; *P < 0.05; **P < 0.01; ***P < 0.001; ns, non-significant
Because the oligomerization of NP and NP–VP30 interaction are essential for the RNA synthesis function of NP (Kirchdoerfer et al., 2015; Leung et al., 2015; Kirchdoerfer et al., 2016; Xu et al., 2017; Batra et al., 2018; Sugita et al., 2018), we sought to determine whether these mutants disrupt the oligomerization of NP or NP–VP30 interaction. We found that all these mutants can interact with NP or VP30 as efficiently as NP (Fig. S4A and S4B), which is consistent with the immunofluorometric assay results (Fig. S2F and S2I). In addition, a previous study showed that NP acts as a bridge to recruit PP2A-B56 via the LxxIxE motif for VP30 dephosphorylation, thus activating viral transcription (Xu et al., 2006; Hertz et al., 2016; Kruse et al., 2018). To further determine whether the NP C-terminal mutants are still able to dephosphorylate VP30, we co-expressed VP30 with NP or NPP697A and found that the levels of VP30 Ser29 phosphorylation were similarly reduced by NPP697A, comparable to that in the presence of NP (Fig. S4C). Taken together, these results showed that the NP C-terminal mutants maintain their normal functions of oligomerization, interactions with other nucleocapsid proteins, and VP30 dephosphorylation.
Next, we sought to determine whether these mutants affect NP encapsidation of viral RNA. To test this, we performed an immunoprecipitation-RT-qPCR assay using the Ebola virus minigenome system and found that amount of immunoprecipitated minigenome RNA was severely reduced for the NP C-terminal mutants compared to NP (Fig. 6C), suggesting that the NP C-terminal hydrophobic core is essential for NP encapsidation of viral RNA. To determine how VP40 inhibits viral RNA synthesis, we examined the effects of VP40 on NP oligomerization and NP–VP30 or VP35–VP30 interactions. We found that VP40 had no effect on NP oligomerization, NP–VP30 or VP35–VP30 interactions (Fig. S4D–F), suggesting that VP40 did not affect the formation of NCs. However, as expected, we found that the ability of NP binding minigenome RNA was dramatically reduced in the presence of VP40 (Fig. 6D), suggesting that the interaction of VP40 with the NP C-terminal hydrophobic core prevents NP from encapsidating viral RNA, thus inhibiting viral RNA synthesis.
Discussion
In a previous study, both the N- and C-termini of NP were found to be important for the interaction of NP with VP40 and the incorporation of NP into VP40–VLP (Noda et al., 2007b). However, how both ends regulate the incorporation of NP into VP40–VLP remains unclear. Here, we show that VP40 is first recruited into IBs by interacting with the NP N-terminus, which subsequently results in conformational change of the NP C-terminus and exposure of the hydrophobic core that associates with VP40 for the incorporation of NP into VLPs. This conclusion is supported by the following evidence. First, the NP N-terminal mutant, NP∆N26–150, failed to interact with VP40 and prevented the incorporation of NP into VP40–VLP (Fig. 1A and 1B). In addition, the structural analysis of the NP and VP40 interacting regions was performed using the HDOCK server based on NP core structure (PDB: 4Z9P) and the VP40 structure (PDB: 1ES6) (Fig. S5A). The top three predicted results were analyzed and we found that the NP-VP40 model showed that with the third highest docking score exhibited the largest interacting area of the three (Fig. S5B), while the top two models had larger interacting region with the C-lobe (Fig. S5C–D). In addition, previous co-IP result showed that the region from amino acid 151 to 300 (within the C-lobe) of NP is dispensable for VP40 binding (Noda et al., 2007b). Taken together, the docking analysis was consistent with our functional data. Moreover, NP∆26–150 maintained its interactions with VP35 and VP30 (Fig. 1C and 1D), indicating that the C-terminal structure of NP∆26–150 is similar to that of NP, which also indicates that the intact C-terminus of NP is unable to interact with VP40. Second, we identified four key amino acids (L692, P697, P698, and W699) at the NP C-terminus which form the hydrophobic core (Fig. S2E); when mutated to alanines (NPL692A, NPP697A, NPP698A, and NPW699A) the incorporation of NP into VP40–VLP was abolished but the interaction with VP40 became stronger (Fig. 2E–H). Furthermore, we also observed that VP40 colocalized with the IBs formed by these mutants, resulting in the aggregation of IBs (Fig. 2I). Third, the release of VP40–VLP was also inhibited by NPL692A, NPP697A, NPP698A, and NPW699A (Fig. 3A, 3B, 3E and 3F), similar to a previous study that showed the final 50 amino acids in the NP C-terminus regulate the budding of VP40 (Licata et al., 2004), suggesting that any mutation within L692, P697, P698, and W699 would prevent conformational changes in the hydrophobic core resulting from interactions of the NP N-terminus with VP40. On the one hand, this results in the hydrophobic core of NP C-terminus being unable to expose itself and interact with VP40; on the other hand, the N-terminal interaction with VP40 is preserved, thereby sequestering VP40 in IBs and causing failure of VP40–VLP budding.
We found that NPP697A had no effect on the interaction of VP40 with Sec24C (Fig. 4D and 4E), suggesting NP–VP40 uses a different mechanism for cellular transport. A previous study revealed that EBOV nucleocapsids are dependent on actin for long-distance transport in Ebola virus-infected cells (Schudt et al., 2015), which indicates that mutations in the NP C-terminus may affect the interaction of VP40 with cytoskeleton proteins, thus inhibiting their cellular transport.
We also found that the NP C-terminal hydrophobic core is critical for NC-like structures to be recruited to viral particles (Fig. 5). Similar to these results, our previous result showed that the interaction of NP with the M protein of HPIV3 regulates the NC-like structures assembly (Zhang et al., 2015). Moreover, the NP C-terminus of HPIV1 and SeV was also reported to mediate interaction with M protein, which is critical for virion incorporation (Coronel et al., 2001). The NP of the influenza virus is essential for virion assembly, possibly via association with M1 (Noton et al., 2009). However, for RSV, the transcriptional antiterminator M2-1 acts as a bridge in linking RNPs with M protein, which is required for the incorporation of RNPs into virions (Li et al., 2008). These results suggest that the mechanisms by which RNPs are assembled and incorporated into viral particles seem to differ among the different viruses. In addition, we could not exclude the possibility that VP35 and VP24 play a role in the correct assembly of virions (Bharat et al., 2012). We also need to further investigate the regions within VP40 that regulate interaction with nucleocapsid proteins, which may provide essential insights into the mechanism underlying the incorporation of Ebola internal proteins into virions.
The M proteins of some NNSVs can regulate RNA synthesis. For EBOV, we confirmed that VP40 inhibits RNA synthesis in a dose-dependent manner (Fig. 6A). Different viruses utilize different mechanisms to regulate RNA synthesis. The M protein of HPIV3 and MeV inhibits RNA synthesis by interacting with NP and, especially in the case of MeV, it is possible that M protein inhibits RNA synthesis to promote viral particle production (Iwasaki et al., 2009; Zhang et al., 2018). Previous studies suggested that EBOV NP oligomerization and NP–RNA binding are cooperative and do not occur independently (Sugita et al., 2018). VP40 can be recruited into IBs via the NP N-terminus that is critical for NP oligomerization (Figs. S1D–F and 2). However, we found that VP40 has no effect on NP oligomerization (Fig. S4D). Moreover, the binding regions of RNA and VP40 at the NP N-terminus do not overlap (Dong et al., 2015; Sugita et al., 2018), making it unlikely that VP40 inhibits RNA synthesis by interacting with the NP N-terminus. Meanwhile, we found that VP40 has no effect on nucleocapsid formation (Fig. S4E and S4F).
Our results show that NPL692A, NPP697A, NPP698A, and NPW699A have almost complete inhibition of RNA synthesis (Fig. 6B), suggesting that the hydrophobic core is indispensable for RNA synthesis. Moreover, we found that these mutants maintain the functions of NP in oligomerization, interacting with other nucleocapsid proteins and engaging in the dephosphorylation of VP30 (Figs. S4A–C and 5B). It is possible that the hydrophobic core could regulate NP encapsidation of viral RNA by changing the state of folding and unfolding, which was confirmed by an IP-RT-qPCR assay (Fig. 6C). Since the NP C-terminal hydrophobic core is critical both for viral RNA synthesis and NC assembly/budding, it was important to determine whether VP40 regulates RNA synthesis and NC assembly/budding by interacting with this domain. We also found that VP40 inhibits NP encapsidation of viral RNA (Fig. 6D). These results suggest that VP40 inhibits RNA synthesis to facilitate NC assembly/budding by interacting with the NP C-terminal hydrophobic core. In addition, we need to further investigate the domains within VP40 that regulate the interaction of VP40 with the NP C-terminal hydrophobic core. Meanwhile, we cannot exclude possibility that cellular factor(s) participate in these processes by interacting with either VP40 or NP, or both.
In the evolution of EBOV, the mRNAs of NP are firstly transcribed, and then go through translation for protein synthesis. Moreover, the amount of NP is much more than VP40 in the cytoplasm and NP preferentially encapsidates viral RNA to form NP-RNA template for RNAs and proteins synthesis. When elements (RNAs and proteins) for packaging are enough, the virus is prepared for shutting down the synthesis of RNAs and proteins, and focusing on the assembly/budding. At this point, VP40 is recruited into IBs via the interaction with NP N-terminus, and then their interaction is switched to NP C-terminus, thereby preventing NP from encapsidating of newly synthesized RNA, thus inhibiting RNA synthesis. However, once having formed NP-RNA template, VP40 interacts with NP in the same way, thus initiating nucleocapsids (NCs) assembly/budding. Therefore, we think that VP40 inhibiting RNA encapsidation to initiate viral assembly is not a paradox in terms of viral lifecycle.
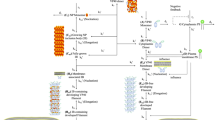
In conclusion, we suggest a model (Fig. 7) through which VP40 can be recruited into IBs mediated by the NP N-terminus, with such interaction then resulting in a conformational change in the NP C-terminus and exposure of the hydrophobic core. On the one hand, when VP40 associates with the NP C-terminal hydrophobic core, VP40 inhibits RNA synthesis by preventing NP from encapsidating of newly synthesized viral RNA (viral RNA encapsidation did not occur); on the other hand, VP40 incorporates NC (viral RNA encapsidation occurred) into VLPs for assembly/budding of virion. To our best knowledge, this is the first study to demonstrate how a two-stage VP40–NP interaction plays a critical role in switching from viral RNA synthesis to assembly/ budding, which suggests that this interaction may be a promising target for antiviral therapy.
Materials and methods
Cell culture
HEK293T and HeLa cells were purchased from China Center for Type Culture Collection (CCTCC) and maintained in Dulbecco’s modified Eagle’s medium (DMEM, Gibco), supplemented with 15% or 8% fetal bovine serum (FBS, Gibco) at 37 °C with 5% CO2.
Plasmid construction
The cDNA of the Zaire ebolavirus and the expression plasmids pCAGGS-NP, pCAGGS-VP35, pCAGGS-VP30, pCAGGS-T7, pCAGGS-Luc2 (encoding firefly luciferase), pCAGGS-L, and p4cis-vRNA-Rluc were provided by Dr. Bo Zhang (Wuhan Institute of Virology); Dr. Heinz Feldmann (National Institutes of Health, Hamilton, Montana, USA). VP40 and VP24 were generated from cDNA by PCR-based cloning techniques. VP40 was cloned into pCAGGS with an N-terminal Myc tag or C-terminal Flag tag, while VP24 was tagged with HA at the N-terminus. All mutants of NP were constructed by PCR amplification using NP as a template, and were cloned into pCAGGS with a Myc or HA tag at N-terminus. The minigenome, encoding Renilla luciferase, which was obtained from the plasmid p4cis-vRNA-Rluc via PCR, was generated as previously described (Muhlberger et al., 1998) and cloned into the T7 expression vector pBS, named pBS-Rluc. VP306A was generated by PCR amplification as previously described (Modrof et al., 2002). The coding region of Sec24C (NCBI accession no. NM_004922) was obtained from HeLa cells via RNA extraction, followed by RT-PCR, and then cloned into the pCAGGS with an HA tag at the N-terminus. The details, related to the construction process of all the plasmids, are available upon request. All the constructs were verified by DNA sequencing.
SDS-PAGE and Western blot
Cells were harvested and lysed with 500 μL of lysis buffer (150 nmol/L NaCl, 50 nmol/L Tris-HCl (pH 7.4), 1% Triton X-100, 1 mmol/L EDTA (pH 8.0), and 0.1% sodium dodecyl sulfate (SDS)) with a protease inhibitor cocktail and then incubated on ice for 30 min. The supernatants were collected by centrifugation at 4 °C at 12,000 ×g for 30 min and then boiled in 5× SDS-PAGE loading buffer for 10 min. The prepared samples were resolved on 10% SDS-PAGE and then transferred onto nitrocellulose membrane (GE Healthcare). The membrane was blocked with 5% skim milk powder in phosphate-buffered saline (PBS) with 0.1% Tween 20 (PBST) for 30 min before being incubated with primary antibodies, followed by incubation with horseradish peroxidase-conjugated goat anti-rabbit or anti-mouse IgG (Thermo Fisher Scientific) for 1 h. The primary antibodies were used as follows: mouse anti-Myc (1:10,000, MBL), mouse anti-HA (1:10,000, MBL), mouse anti-Flag (1:10,000, Sigma), rabbit anti-HA (1:10,000, Sigma), rabbit anti-Flag (1:10,000, CST), rabbit anti-Myc (1:10,000, Sigma), rabbit anti-VP30 pSer29 antibody (1:1000, ABclonal), rabbit anti-Na/K ATPase (1:1000, ABclonal), and mouse anti-GAPDH (1:1000, ABclonal). The secondary antibodies, goat anti-rabbit IgG and goat anti-mouse IgG, were used at a 1:5,000 dilution.
Virus-like particle assays
HEK293T cells were transfected with the indicated plasmids. At 36 h post-transfection, the supernatant was harvested and clarified by centrifugation at 12,000 ×g for 10 min and then pelleted through a 20% sucrose cushion; it was subsequently ultra centrifugated at 35,000 rpm at 4 °C for 2 h. Cell lysates were obtained as mentioned for the Western blots. The VLP pellets were resuspended in PBS at 4 °C overnight. The supernatants and cell lysates were boiled in 5× SDS-PAGE loading buffer for 10 min and then subjected to Western blot analysis.
Protease protection assay
The VLP pellets were prepared as described above and then divided into four aliquots and treated as follows: (i) left untreated, (ii) treated with Triton X-100 to a final concentration of 1%, (iii) treated with trypsin to a final concentration of 1 μg/mL, or (iv) treated with Triton X-100 plus trypsin. The prepared samples were incubated at 37 °C for 30 min and then boiled in 5× SDS-PAGE loading buffer for 10 min and analyzed via Western blot.
In vivo co-immunoprecipitation
HEK293T cells were transfected with the indicated plasmids using the calcium phosphate precipitation method. At 30 h post-transfection, cell lysates were prepared in a 500 μL lysis buffer as described for WB analysis. Then, 40 μL lysates were removed for input analysis, and the remaining cell lysates were incubated with the relevant antibodies (anti-Flag affinity gel (Sigma) or anti-Myc affinity gel (Biolegend)) overnight at 4 °C with gentle rotation. The beads were collected by centrifugation at 5,000 rpm for 2 min at 4 °C and washed three times with a lysis buffer. The collected beads and input were boiled in 1× or 5× SDS-PAGE loading buffer, respectively, for 10 min and then subjected to WB analysis.
Immunofluorescence analysis
HeLa cells were cultured on the coverslips in 24-well plates overnight and then transfected with the indicated plasmids with Lipofectamine 2000 reagent (Invitrogen). After 26 h of transfection, the cells were harvested and fixed with ice-cold 4% (w/v) paraformaldehyde at room temperature for 20 min and then washed with PBS three times (each time for 5 min) before being incubated with 0.2% Triton X-100 for 20 min. Subsequently, the cells were blocked with 3% BSA for 30 min after being washed three times with PBS. Then, the specific primary antibodies diluted in 1% BSA were added and incubated overnight at 4 °C. After being washed by PBS, the cells were incubated with secondary antibodies diluted in 1% BSA for 1 h at room temperature. The nucleus was stained with DAPI for 5 min, and the cells were finally mounted with Prolong Diamond Antifade Mountant (Life Technology) and examined on a Leica or Zeiss confocal microscope. The primary antibodies were used as follows: rabbit anti-HA (1:1000, Sigma), mouse anti-Myc (1:1000, MBL), and mouse anti-Flag (1:1000, MBL).
Transmission electron microscopy
HEK293T cells were transfected with the indicated plasmids. At 36 h post-transfection, the culture medium was discarded, and then the cells were fixed with a buffer (3% paraformaldehyde and 1.5% glutaraldehyde in a 0.1 mol/L sodium phosphate buffer (pH 7.4)) for 1 h at room temperature. Subsequently, the cells were harvested and subjected to gradient centrifugation at 1000 ×g, 3,000 ×g, 6,000 ×g, and 12,000 ×g. Then, the cells were fixed with 1% osmium tetroxide at 4 °C for 1 h under dark condition, followed by being incubated with 2% uranyl acetate overnight. After being dehydrated with sequential washes in 50%, 75%, 95%, and 100% ethanol, the cells were mounted with epoxy resin and then incubated overnight at 37 °C or 65 °C for 48–72 h and thinly sliced. The samples were absorbed on uncoated 200-mesh copper grids and stained with uranyl acetate and lead citrate, before being finally analyzed via transmission electron microscopy (JEOL, JEM-1400 plus).
Live cell imaging
A previous study showed that the viral proteins of Ebola fused to larger fluorescent tags were only fully functional if the respective wild type proteins are also expressed (Takamatsu et al., 2018). Therefore, GFP-NP, GFP-NPW699A, or mCherry-VP40 were co-expressed with the corresponding wild type protein. For live-cell imaging, HeLa cells were grown in 20 mm dishes and transfected with the indicated plasmids. At 24 h post-transfection, live-cell time-lapse experiment images were recorded with a Zeiss LSM 800 fluorescence microscope. Pictures were taken every 120 s and processed with Zeiss Microscope Imaging Software.
Cytosol and plasma membrane protein fractionation
HEK293T cells were transfected with the indicated plasmids. The cells were harvested at 24 h post-transfection and washed with cold PBS and then collected via low-speed centrifugation. The cytosol, organelle membrane, and plasma membrane protein fractions were isolated sequentially using the MinuteTM plasma membrane protein isolation and cell fractionation kit (INVENT) according to the manufacturer’s instructions. The cytosol and plasma membrane fractions were subjected to WB analysis. GAPDH and sodium potassium ATPase were used as controls for the cytosol and plasma membrane, respectively.
Minigenome assay
HEK293T cells, cultured in a 12-well plate, were transfected with the minigenome assay system: 125 ng pCAGGS-Myc-NP, 125 ng pCAGGS-HA-VP35, 225 ng pCAGGS-Flag-VP30, 1 μg pCAGGS-L, 250 ng pCAGGS-T7 polymerase, 250 ng pBS-Rluc (minigenome, encoding Renilla luciferase), and 25 ng pCAGGS-Luc2 (encoding firefly luciferase as a transfection control). Total DNA levels were kept constant by transfection with pCAGGS. At 30 h post-transfection, cells were lysed in 150 μL 1× lysis buffer, and 20 μL aliquots of lysates were analyzed with the Dual-Glo Luciferase Assay System (Promega) according to the manufacturer’s instructions. A portion of the cell lysate was subjected to WB analysis as described above. Data are presented as a relative scale, with the Renilla luciferase activity normalized to the firefly luciferase values.
In vivo RNA immunoprecipitation
HEK293T cells were grown in 6 cm dishes and transfected with 1 μg pCAGGS-Myc-NP/4 μg pCAGGS-Myc-NPL692A/2 μg pCAGGS-Myc-NPP697A/2 μg pCAGGS-Myc-NPP698A/4 μg pCAGGS-Myc-NPW699A, 1 μg pCAGGS-HAVP35, 1.8 μg pCAGGS-FlagVP30, 8 μg pCAGGS-L, 2 μg pCAGGS-T7, and 2 μg pBS-Rluc. In a separate experiment, 4 μg pCAGGS-VP40-Flag was co-expressed. pCAGGS was then transfected to keep the total DNA levels constant. After transfection for 30 h, the cells were lysed with 500 μL RNase-free NP-40 buffer (50 mmol/L Tris-HCl (pH 7.4), 50 mmol/L NaCl, 50 mmol/L NaF, 0.5% NP40) on ice for 20 min and then centrifuged at 12,000 ×g at 4 °C for 20 min. The supernatants were collected and divided into three parts. One portion (8%) was subjected to WB analysis to detect the expression of plasmids; another portion (12%) was resuspended in 1 mL TRIzol (Ambion, Invitrogen) to determine the total cellular minigenome RNA. The remaining cell lysates were subjected to immunoprecipitation with anti-Myc antibodies for 4 h at 4 °C. The beads were collected by centrifugation at 5,000 rpm for 2 min at 4 °C and washed 3 times with NP-40 buffer. The collected beads were finally resuspended in 1 mL NP-40 buffer and divided into two parts. One portion (6%) was collected and subjected to WB analysis to detect the amount of precipitated proteins. The remaining beads were resuspended in 1 mL TRIzol. The total cellular minigenome RNA and immunoprecipitated RNA were extracted following the manufacturer’s directions. Each sample of RNA was eluted in 20–30 μL RNase-free water. The RNA concentration was measured with a NanoDrop spectrophotometer (Promega).
RNA quantification
The cellular and immunoprecipitated RNA of the EBOV minigenome were quantified by quantitative reverse transcription PCR (RT-qPCR). A total of 500 ng of RNA was used for the reverse transcription of genomic viral RNA (vRNA), using the kit following the manufacturer’s instructions (Fermentas, Waltham, MA, USA), followed by a qPCR assay to quantify RNA abundance. In each case, data are represented as a relative scale, with the immunoprecipitated vRNA abundance normalized to the total cellular vRNA values. The values were expressed as a percentage, where 100% was taken as abundance of immunoprecipitated Myc-NP. The specific primer for reverse transcription of the vRNA was 5′-GACAAATTGCTCGGAATCACAAAATTCC-3′ (reverse primer). The primers for the Renilla luciferase reporter for qPCR were as follows: 5′-GGTAGGCGTTGTAGTTGCG-3′ (forward primer); 5′-TACGAGCA CCAAGACAAGA-3′ (reverse primer).
Statistical analysis
Data are expressed as the mean ± standard deviation (SD). The significance of the variability between different groups was determined by two-way ANOVA tests of variance using the GraphPad Prism software (version 5.0). P < 0.05 was considered statistically significant, and P > 0.05 was considered statistically non-significant.
Abbreviations
- coIP:
-
coimmunoprecipitation
- COPII:
-
coat protein complex II
- EBOV:
-
Ebola virus
- F:
-
fusion protein
- GP:
-
glycoprotein
- HN:
-
hemagglutinin-neuraminidase
- HPIV1:
-
human parainfluenza virus type 1
- HPIV3:
-
human parainfluenza virus type 3
- hRSV:
-
human respiratory syncytial virus
- IBs:
-
inclusion bodies
- IF:
-
immunofluorescence
- IP-RT-qPCR:
-
immunoprecipitation quantitative reverse transcription PCR
- LLOV:
-
Lloviu virus
- M:
-
matrix protein
- MARV:
-
Marburg virus
- MeV:
-
Measles virus
- NC:
-
nucleocapsid
- NiV:
-
Nipah virus
- NNSVs:
-
non-segmented negative strand RNA viruses
- NP:
-
nucleoprotein
- NSVs:
-
negative strand RNA viruses
- P:
-
phosphoprotein
- PM:
-
plasma membrane
- PP2A-B56:
-
protein phosphatase 2A, regulatory subunit B56 family
- RNP:
-
ribonucleoprotein
- Sec24C:
-
secretory COPII component 24C
- SeV:
-
Sendai virus
- SV5:
-
paramyxovirus simian virus 5
- TEM:
-
transmission electron microscopy
- VLP:
-
virus like particle
- vRNA:
-
genomic viral RNA
- VSV:
-
vesicular stomatitis virus
- WT:
-
wild type
References
Aruna A, Mbala P, Minikulu L, Mukadi D, Bulemfu D, Edidi F, Bulabula J, Tshapenda G, Nsio J, Kitenge R et al (2019) Ebola virus disease outbreak: Democratic Republic of the Congo, August 2018–November 2019. MMWR Morb Mortal Wkly Rep 68:1162–1165
Batra J, Hultquist JF, Liu D, Shtanko O, Von Dollen J, Satkamp L, Jang GM, Luthra P, Schwarz TM, Small GI et al (2018) Protein interaction mapping identifies RBBP6 as a negative regulator of ebola virus replication. Cell 175(1917–1930):e1913
Bharat TA, Noda T, Riches JD, Kraehling V, Kolesnikova L, Becker S, Kawaoka Y, Briggs JA (2012) Structural dissection of Ebola virus and its assembly determinants using cryo-electron tomography. Proc Natl Acad Sci USA 109:4275–4280
Bornholdt ZA, Noda T, Abelson DM, Halfmann P, Wood MR, Kawaoka Y, Saphire EO (2013) Structural rearrangement of ebola virus VP40 begets multiple functions in the virus life cycle. Cell 154:763–774
Buchholz CJ, Spehner D, Drillien R, Neubert WJ, Homann HE (1993) The conserved N-terminal region of Sendai virus nucleocapsid protein NP is required for nucleocapsid assembly. J Virol 67:5803–5812
Coronel EC, Murti KG, Takimoto T, Portner A (1999) Human parainfluenza virus type 1 matrix and nucleoprotein genes transiently expressed in mammalian cells induce the release of virus-like particles containing nucleocapsid-like structures. J Virol 73:7035–7038
Coronel EC, Takimoto T, Murti KG, Varich N, Portner A (2001) Nucleocapsid incorporation into parainfluenza virus is regulated by specific interaction with matrix protein. J Virol 75:1117–1123
De BP, Thornton GB, Luk D, Banerjee AK (1982) Purified matrix protein of vesicular stomatitis virus blocks viral transcription in vitro. Proc Natl Acad Sci USA 79:7137–7141
Dong S, Yang P, Li G, Liu B, Wang W, Liu X, Xia B, Yang C, Lou Z, Guo Y et al (2015) Insight into the Ebola virus nucleocapsid assembly mechanism: crystal structure of Ebola virus nucleoprotein core domain at 1.8 A resolution. Protein Cell 6:351–362
Dziubanska PJ, Derewenda U, Ellena JF, Engel DA, Derewenda ZS (2014) The structure of the C-terminal domain of the Zaire ebolavirus nucleoprotein. Acta Crystallogr D 70:2420–2429
Garcia-Barreno B, Delgado T, Melero JA (1996) Identification of protein regions involved in the interaction of human respiratory syncytial virus phosphoprotein and nucleoprotein: significance for nucleocapsid assembly and formation of cytoplasmic inclusions. J Virol 70:801–808
Groseth A, Charton JE, Sauerborn M, Feldmann F, Jones SM, Hoenen T, Feldmann H (2009) The Ebola virus ribonucleoprotein complex: a novel VP30-L interaction identified. Virus Res 140:8–14
Hartlieb B, Muziol T, Weissenhorn W, Becker S (2007) Crystal structure of the C-terminal domain of Ebola virus VP30 reveals a role in transcription and nucleocapsid association. Proc Natl Acad Sci USA 104:624–629
Hartman AL, Towner JS, Nichol ST (2010) Ebola and marburg hemorrhagic fever. Clin Lab Med 30:161–177
Heinrich BS, Cureton DK, Rahmeh AA, Whelan SP (2010) Protein expression redirects vesicular stomatitis virus RNA synthesis to cytoplasmic inclusions. PLoS Pathog 6:e1000958
Hertz EPT, Kruse T, Davey NE, Lopez-Mendez B, Sigurethsson JO, Montoya G, Olsen JV, Nilsson J (2016) A conserved motif provides binding specificity to the PP2A-B56 phosphatase. Mol Cell 63:686–695
Hoenen T, Jung S, Herwig A, Groseth A, Becker S (2010) Both matrix proteins of Ebola virus contribute to the regulation of viral genome replication and transcription. Virology 403:56–66
Hoenen T, Shabman RS, Groseth A, Herwig A, Weber M, Schudt G, Dolnik O, Basler CF, Becker S, Feldmann H (2012) Inclusion bodies are a site of ebolavirus replication. J Virol 86:11779–11788
Hoenen T, Groseth A, Feldmann H (2019) Therapeutic strategies to target the Ebola virus life cycle. Nat Rev Microbiol 17:593–606
Huang Y, Xu L, Sun Y, Nabel GJ (2002) The assembly of Ebola virus nucleocapsid requires virion-associated proteins 35 and 24 and posttranslational modification of nucleoprotein. Mol Cell 10:307–316
Iwasaki M, Takeda M, Shirogane Y, Nakatsu Y, Nakamura T, Yanagi Y (2009) The matrix protein of measles virus regulates viral RNA synthesis and assembly by interacting with the nucleocapsid protein. J Virol 83:10374–10383
Jasenosky LD, Neumann G, Lukashevich I, Kawaoka Y (2001) Ebola virus VP40-induced particle formation and association with the lipid bilayer. J Virol 75:5205–5214
Jasenosky LD, Neumann G, Kawaoka Y (2010) Minigenome-based reporter system suitable for high-throughput screening of compounds able to inhibit Ebolavirus replication and/or transcription. Antimicrob Agents Chemother 54:3007–3010
Johnson RF, McCarthy SE, Godlewski PJ, Harty RN (2006) Ebola virus VP35-VP40 interaction is sufficient for packaging 3E-5E minigenome RNA into virus-like particles. J Virol 80:5135–5144
Justice PA, Sun W, Li Y, Ye Z, Grigera PR, Wagner RR (1995) Membrane vesiculation function and exocytosis of wild-type and mutant matrix proteins of vesicular stomatitis virus. J Virol 69:3156–3160
Kirchdoerfer RN, Abelson DM, Li S, Wood MR, Saphire EO (2015) Assembly of the ebola virus nucleoprotein from a chaperoned VP35 complex. Cell Rep 12:140–149
Kirchdoerfer RN, Moyer CL, Abelson DM, Saphire EO (2016) The ebola virus VP30-NP interaction is a regulator of viral RNA synthesis. PLoS Pathog 12:e1005937
Kirchdoerfer RN, Wasserman H, Amarasinghe GK, Saphire EO (2017) Filovirus structural biology: the molecules in the machine. Curr Top Microbiol Immunol 411:381–417
Kruse T, Biedenkopf N, Hertz EPT, Dietzel E, Stalmann G, Lopez-Mendez B, Davey NE, Nilsson J, Becker S (2018) The ebola virus nucleoprotein recruits the host PP2A-B56 phosphatase to activate transcriptional support activity of VP30. Mol Cell 69(136–145):e136
Leung DW, Borek D, Luthra P, Binning JM, Anantpadma M, Liu G, Harvey IB, Su Z, Endlich-Frazier A, Pan J et al (2015) An intrinsically disordered peptide from ebola virus VP35 controls viral RNA synthesis by modulating nucleoprotein-RNA interactions. Cell Rep 11:376–389
Li Y, Luo L, Schubert M, Wagner RR, Kang CY (1993) Viral liposomes released from insect cells infected with recombinant baculovirus expressing the matrix protein of vesicular stomatitis virus. J Virol 67:4415–4420
Li D, Jans DA, Bardin PG, Meanger J, Mills J, Ghildyal R (2008) Association of respiratory syncytial virus M protein with viral nucleocapsids is mediated by the M2-1 protein. J Virol 82:8863–8870
Licata JM, Johnson RF, Han Z, Harty RN (2004) Contribution of ebola virus glycoprotein, nucleoprotein, and VP24 to budding of VP40 virus-like particles. J Virol 78:7344–7351
Longhi S, Receveur-Brechot V, Karlin D, Johansson K, Darbon H, Bhella D, Yeo R, Finet S, Canard B (2003) The C-terminal domain of the measles virus nucleoprotein is intrinsically disordered and folds upon binding to the C-terminal moiety of the phosphoprotein. J Biol Chem 278:18638–18648
Martin B, Hoenen T, Canard B, Decroly E (2016) Filovirus proteins for antiviral drug discovery: a structure/function analysis of surface glycoproteins and virus entry. Antiviral Res 135:1–14
Messaoudi I, Amarasinghe GK, Basler CF (2015) Filovirus pathogenesis and immune evasion: insights from Ebola virus and Marburg virus. Nat Rev Microbiol 13:663–676
Misasi J, Sullivan NJ (2014) Camouflage and misdirection: the full-on assault of Ebola virus disease. Cell 159:477–486
Modrof J, Muhlberger E, Klenk HD, Becker S (2002) Phosphorylation of VP30 impairs ebola virus transcription. J Biol Chem 277:33099–33104
Muhlberger E, Lotfering B, Klenk HD, Becker S (1998) Three of the four nucleocapsid proteins of Marburg virus, NP, VP35, and L, are sufficient to mediate replication and transcription of Marburg virus-specific monocistronic minigenomes. J Virol 72:8756–8764
Nanbo A, Watanabe S, Halfmann P, Kawaoka Y (2013) The spatio-temporal distribution dynamics of Ebola virus proteins and RNA in infected cells. Sci Rep 3:1206
Negredo A, Palacios G, Vazquez-Moron S, Gonzalez F, Dopazo H, Molero F, Juste J, Quetglas J, Savji N, de la Cruz Martinez M et al (2011) Discovery of an ebolavirus-like filovirus in europe. PLoS Pathog 7:e1002304
Nelson EV, Schmidt KM, Deflube LR, Doganay S, Banadyga L, Olejnik J, Hume AJ, Ryabchikova E, Ebihara H, Kedersha N et al (2016) Ebola virus does not induce stress granule formation during infection and sequesters stress granule proteins within viral inclusions. J Virol 90:7268–7284
Noda T, Halfmann P, Sagara H, Kawaoka Y (2007a) Regions in Ebola virus VP24 that are important for nucleocapsid formation. J Infect Dis 196(Suppl 2):S247–250
Noda T, Watanabe S, Sagara H, Kawaoka Y (2007b) Mapping of the VP40-binding regions of the nucleoprotein of Ebola virus. J Virol 81:3554–3562
Noda T, Hagiwara K, Sagara H, Kawaoka Y (2010) Characterization of the Ebola virus nucleoprotein-RNA complex. J Gen Virol 91:1478–1483
Noton SL, Simpson-Holley M, Medcalf E, Wise HM, Hutchinson EC, McCauley JW, Digard P (2009) Studies of an influenza A virus temperature-sensitive mutant identify a late role for NP in the formation of infectious virions. J Virol 83:562–571
Patch JR, Crameri G, Wang LF, Eaton BT, Broder CC (2007) Quantitative analysis of Nipah virus proteins released as virus-like particles reveals central role for the matrix protein. Virol J 4:1
Richard CA, Rincheval V, Lassoued S, Fix J, Cardone C, Esneau C, Nekhai S, Galloux M, Rameix-Welti MA, Sizun C et al (2018) RSV hijacks cellular protein phosphatase 1 to regulate M2-1 phosphorylation and viral transcription. PLoS Pathog 14:e1006920
Runkler N, Pohl C, Schneider-Schaulies S, Klenk H-D, Maisner A (2007) Measles virus nucleocapsid transport to the plasma membrane requires stable expression and surface accumulation of the viral matrix protein. Cell Microbiol 9:1203–1214
Sakaguchi T, Uchiyama T, Fujii Y, Kiyotani K, Kato A, Nagai Y, Kawai A, Yoshida T (1999) Double-layered membrane vesicles released from mammalian cells infected with Sendai virus expressing the matrix protein of vesicular stomatitis virus. Virology 263:230–243
Schmitt AP, Leser GP, Waning DL, Lamb RA (2002) Requirements for budding of paramyxovirus simian virus 5 virus-like particles. J Virol 76:3952–3964
Schudt G, Dolnik O, Kolesnikova L, Biedenkopf N, Herwig A, Becker S (2015) Transport of Ebolavirus Nucleocapsids Is Dependent on Actin Polymerization: Live-Cell Imaging Analysis of Ebolavirus-Infected Cells. J Infect Dis 212(Suppl 2):S160–166
Shi W, Huang Y, Sutton-Smith M, Tissot B, Panico M, Morris HR, Dell A, Haslam SM, Boyington J, Graham BS et al (2008) A filovirus-unique region of Ebola virus nucleoprotein confers aberrant migration and mediates its incorporation into virions. J Virol 82:6190–6199
Su Z, Wu C, Shi L, Luthra P, Pintilie GD, Johnson B, Porter JR, Ge P, Chen M, Liu G et al (2018) Electron cryo-microscopy structure of ebola virus nucleoprotein reveals a mechanism for nucleocapsid-like assembly. Cell 172(966–978):e912
Sugahara F, Uchiyama T, Watanabe H, Shimazu Y, Kuwayama M, Fujii Y, Kiyotani K, Adachi A, Kohno N, Yoshida T et al (2004) Paramyxovirus Sendai virus-like particle formation by expression of multiple viral proteins and acceleration of its release by C protein. Virology 325:1–10
Sugita Y, Matsunami H, Kawaoka Y, Noda T, Wolf M (2018) Cryo-EM structure of the Ebola virus nucleoprotein-RNA complex at 3.6 A resolution. Nature 563:137–140
Takamatsu Y, Kolesnikova L, Becker S (2018) Ebola virus proteins NP, VP35, and VP24 are essential and sufficient to mediate nucleocapsid transport. Proc Natl Acad Sci USA 115:1075–1080
Timmins J, Scianimanico S, Schoehn G, Weissenhorn W (2001) Vesicular release of ebola virus matrix protein VP40. Virology 283:1–6
Trunschke M, Conrad D, Enterlein S, Olejnik J, Brauburger K, Muhlberger E (2013) The L-VP35 and L–L interaction domains reside in the amino terminus of the Ebola virus L protein and are potential targets for antivirals. Virology 441:135–145
Vetter P, Fischer WA 2nd, Schibler M, Jacobs M, Bausch DG, Kaiser L (2016) Ebola virus shedding and transmission: review of current evidence. J Infect Dis 214:S177–S184
Watanabe S, Noda T, Kawaoka Y (2006) Functional mapping of the nucleoprotein of Ebola virus. J Virol 80:3743–3751
Weik M, Modrof J, Klenk HD, Becker S, Muhlberger E (2002) Ebola virus VP30-mediated transcription is regulated by RNA secondary structure formation. J Virol 76:8532–8539
Xu Y, Xing Y, Chen Y, Chao Y, Lin Z, Fan E, Yu JW, Strack S, Jeffrey PD, Shi Y (2006) Structure of the protein phosphatase 2A holoenzyme. Cell 127:1239–1251
Xu W, Luthra P, Wu C, Batra J, Leung DW, Basler CF, Amarasinghe GK (2017) Ebola virus VP30 and nucleoprotein interactions modulate viral RNA synthesis. Nat Commun 8:15576
Yamayoshi S, Noda T, Ebihara H, Goto H, Morikawa Y, Lukashevich IS, Neumann G, Feldmann H, Kawaoka Y (2008) Ebola virus matrix protein VP40 uses the COPII transport system for its intracellular transport. Cell Host Microbe 3:168–177
Yang XL, Tan CW, Anderson DE, Jiang RD, Li B, Zhang W, Zhu Y, Lim XF, Zhou P, Liu XL et al (2019) Author correction: characterization of a filovirus (Mengla virus) from rousettus bats in China. Nat Microbiol 4:543
Zhang S, Chen L, Zhang G, Yan Q, Yang X, Ding B, Tang Q, Sun S, Hu Z, Chen M (2013) An amino acid of human parainfluenza virus type 3 nucleoprotein is critical for template function and cytoplasmic inclusion body formation. J Virol 87:12457–12470
Zhang G, Zhang S, Ding B, Yang X, Chen L, Yan Q, Jiang Y, Zhong Y, Chen M (2014) A leucine residue in the C terminus of human parainfluenza virus type 3 matrix protein is essential for efficient virus-like particle and virion release. J Virol 88:13173–13188
Zhang G, Zhong Y, Qin Y, Chen M (2015) Interaction of human parainfluenza virus type 3 nucleoprotein with matrix protein mediates internal viral protein assembly. J Virol 90:2306–2315
Zhang S, Jiang Y, Cheng Q, Zhong Y, Qin Y, Chen M (2017) Inclusion body fusion of human parainfluenza virus type 3 regulated by acetylated alpha-tubulin enhances viral replication. J Virol 91:e01802
Zhang S, Cheng Q, Luo C, Qin Y, Chen M (2018) Human parainfluenza virus type 3 matrix protein reduces viral RNA synthesis of HPIV3 by regulating inclusion body formation. Viruses 10:125
ACKNOWLEDGMENTS
We acknowledge Dr. Bo Zhang (Wuhan Institute of Virology) and Dr. Heinz Feldmann (National Institutes of Health, Hamilton, Montana, USA) for providing the Ebola cDNA and minigenome assay system, respectively. This research is supported by grants from the National Natural Science Foundation of China (81825015), National Key R&D Program of China (2017YFA0505801), National Science and Technology Major Project (2018ZX10101004), National Natural Science Foundation of China (81871650 and 31630086), The Natural Science Foundation of Hubei Province Innovation Group (2017CFA022), and Advanced Customer Cultivation Project of Wuhan National Biosafety Laboratory (2019ACCP-MS06)
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Linjuan Wu, Dongning Jin, Dan Wang, Xuping Jing, Peng Gong, Yali Qin and Mingzhou Chen declare that they have no conflict of interest.
This article does not contain any studies with human or animal subjects performed by the any of the authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary material 2 (AVI 18436 kb)
Supplementary material 3 (AVI 13210 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, L., Jin, D., Wang, D. et al. The two-stage interaction of Ebola virus VP40 with nucleoprotein results in a switch from viral RNA synthesis to virion assembly/budding. Protein Cell 13, 120–140 (2022). https://doi.org/10.1007/s13238-020-00764-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13238-020-00764-0