Abstract
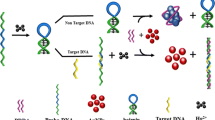
In this study, we introduced a simple method for detecting nucleoside triphosphates (NTPs) using controlled transcription-driven light-up aptamer amplification. Based on the concentration of target NTP, the light-up aptamer was amplified using a DNA template encoding the Broccoli aptamer sequence through an incomplete transcription mixture (without each NTP). The Broccoli aptamer associated with DFHBI-1T produced significantly enhanced fluorescence signals that could sensitively detect NTPs in a label-free manner. In addition, the proposed assay was useful for quantitatively detecting NTPs in serum. Thus, we expect that this method has great potential for NTP analysis in bioassays and biological researches.





Similar content being viewed by others
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Acin-Perez, R., Gatti, D.L., Bai, Y., Manfredi, G.: Protein phosphorylation and prevention of cytochrome oxidase inhibition by ATP: coupled mechanisms of energy metabolism regulation. Cell Metab.Metab. 13, 712–719 (2011)
Gourine, A.V., Llaudet, E., Dale, N., Spyer, K.M.: ATP is a mediator of chemosensory transduction in the central nervous system. Nature 436, 108–111 (2005)
Kerr, C., Szmacinski, H., Fisher, M.L., Nance, B., Lakowicz, J.R., Akbar, A., Keillor, J.W., Lok Wong, T., Godoy-Ruiz, R., Toth, E.A., Weber, D.J., Eckert, R.L.: Transamidase site-targeted agents alter the conformation of the transglutaminase cancer stem cell survival protein to reduce GTP binding activity and cancer stem cell survival. Oncogene 36, 2981–2990 (2017)
Goldsmith, D.J., Carrey, E.A., Edbury, S.M., Marinaki, A.M., Simmonds, H.A.: GTP concentrations are elevated in erythrocytes of renal transplant recipients when conventional immunosuppression is replaced by the inosine monophosphate dehydrogenase inhibitor mycophenolic acid mofetil (MMF). Nucleosides Nucleotides Nucl Acids 23, 1407–1409 (2004)
Fernadez, F., Shridas, P., Jiang, S., Aebi, M., Waechter, C.J.: Expression and characterization of a human cDNA that complements the temperature-sensitive defect in dolichol kinase activity in the yeast sec59-1 mutant: the enzymatic phosphorylation of dolichol and diacylglycerol are catalyzed by separate CTP-mediated kinase activities in Saccharomyces cerevisiae. Glycobiology 12, 555–562 (2002)
Cornell, R.B., Northwood, I.C.: Regulation of CTP: phosphocholine ctidiylyltrnsferase by amphitropism and relocalization. Trends Biochem. Sci.Biochem. Sci. 25, 441–447 (2000)
Lecca, D., Ceruti, S.: Uracil nucleotides: from metabolic intermediates to neuroprotection and neuroinflammation. Biochem. Pharmacol.. Pharmacol. 75, 1869–1881 (2008)
Anderson, C.M., Parkinson, F.E.: Potential signalling roles for UTP and UDP: sources, regulation and release of uracil nucleotides. Trends Pharmacol. Sci.Pharmacol. Sci. 18, 387–392 (1997)
Bradbury, D.A., Simmons, T.D., Slater, K.J., Crouch, S.P.: Measurement of the ADP:ATP ratio in human leukaemic cell lines can be used as an indicator of cell viability, necrosis and apoptosis. J. Immunol. Methods 240, 79–92 (2000)
Bianchi-Smiraglia, A., Bagati, A., Fink, E.E., Moparthy, S., Wawrzyniak, J.A., Marvin, E.K., Battaglia, S., Jowdy, P., Kolesnikova, M., Foley, C.E., Berman, A.E., Kozlova, N.I., Lipchick, B.C., Paul-Rosner, L.M., Bshara, W., Ackroyd, J.J., Shewach, D.S., Nikiforov, M.A.: Microphthalmia-associated transcription factor suppresses invasion by reducing intracellular GTP pools. Oncogene 36, 84–96 (2017)
Huang, R., Zhou, P.K.: DNA damage repair: historical perspectives, mechanistic pathways and clinical translation for targeted cancer therapy. Signal Transduct. Target. Ther.Transduct. Target. Ther. 6, 254 (2021)
Kepp, O., Bezu, L., Yamazaki, T., Di Virgillio, F., Smyth, M.J., Kroemer, G., Galluzzi, L.: ATP and cancer immunisurveillance. EMBO J. 40(13), e108130 (2021)
Gibson, J.J., Edwards, T.W.D., Birks, S.J., St Amour, N.A., Buhay, W.M., McEachern, P., Wolfe, B.B., Peters, D.L.: Progress in isotope tracer hydrology in Canada. Hydrol. Process. Process 19, 303–327 (2005)
Huang, D., Zhang, Y., Chen, X.: Analysis of intracellular nucleoside triphosphate levels in normal and tumor cell lines by high-performance liquid chromatography. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 784, 101–109 (2003)
Cohen, S., Megherbi, M., Jordheim, L.P., Lefebvre, I., Perigaud, C., Dumontet, C., Guitton, J.: Simultaneous analysis of eight nucleoside triphosphates in cell lines by liquid chromatography coupled with tandem mass spectrometry. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 877, 3831–3840 (2009)
Marion, D.: An introduction to biological NMR spectroscopy. Mol. Cell. Proteom. 12, 3006–3025 (2013)
Ouellet, J.: RNA fluorescence with light-up aptamers. Front. Chem. 4, 29 (2016)
Paige, J.S., Karen, Y.W., Samie, T.J.: RNA mimics of green fluorescent protein. Science 333, 642–646 (2011)
Han, K.Y., Leslie, B.J., Fei, J., Zhang, J., Ha, T.: Understanding the photophysics of the spinach-DFHBI RNA aptamer-fluorogen complex to improve live-cell RNA imaging. JACS 130, 19033–19038 (2013)
Kartje, Z.J., Janis H.I., Mukhopadhyay, S., Gagnon, K.T.: Revisiting T7 RNA polymerase transcription in vitro with the Broccoli RNA aptamer as a simplified real-time fluorescent reporter. J. Biol. Chem. 296 (2021)
Hong, Y., Kim, D.E., Park, Y.J., Kim, D.M., Byun, J.Y., Shin, Y.B.: MicroRNA detection using light-up aptamer amplification based on nuclease protection transcription. Chem. Commun. (Camb.)Commun. (Camb.) 58, 2359–2362 (2022)
Moroney, S.E., Piccirilli, J.A.: Abortive products as initiation nucleotides during transcription by T7 RNA polymerase. Biochemistry 30(42), 10343–10349 (1991)
Gong, P., Martin, C.T.: Mechanism of instability in abortive cycling by T7 RNA polymerase. J. Biol. Chem. 281(33), 23533–23544 (2006)
Dousis, A., Ravichandran, K., Hobert, E.M., Moore, M.J., Rabideau, A.E.: An engineered T7 RNA polymerase that produces mRNA free of immunostimnlatory byproducts. Nat. Biotechnol.Biotechnol. 41(4), 560–568 (2023)
Kennedy, W.P., Momand, J.R., Yin, Y.W.: Mechanism for De novo RNA synthesis and initiating nucleotide specificity by T7 RNA polymerase. J. Mol. Biol. 370, 256–268 (2007)
Kuzmine, I., Gottlieb, P.A., Martin, C.T.: Binding of the priming nucleotide in the initiation of transcription by T7 RNA polymerase. J. Biol. Chem. 278, 2819–2823 (2003)
Jia, Y., Patel, S.S.: Kinetic mechanism of GTP binding and RNA synthesis during transcription initiation by bacteriophage T7 RNA polymerase. J. Biol. Chem. 272, 30147–30153 (1997)
Pal, S., Dasgupta, D.: Differentail screening calorimetric approach to study the effect of melting region upon transcritpion initiation by T7 RNA polymerase and role of high affinity GTP binding. J. Biomol. Struct. Dyn.Biomol. Struct. Dyn. 31(3), 288–298 (2013)
Traut, T.W.: Physiological concentrations of purines and pyrimidines. Mol. Cell. Biochem.Biochem. 140, 1–22 (1994)
Maity, D., Li, M., Ehlers, M., Gigante, A., Schmuck, C.: Correction: a metal-free fluorescence turn-on molecular probe for detection of nucleoside triphosphates. Chem. Commun. (Camb.)Commun. (Camb) 53, 12588 (2017)
Mittal, L.S., Sharma, P., Kaur, N., Singh, P.: A perylenediimide based 'on-off’ chemosensor for the detection of nucleoside triphosphates: an efficient ensemble for monitoring alkaline phosphatase activity. Anal. Methods 11, 5320–5327 (2019)
Ding, Y., Jia, Q., Wen, Y., Liu, W., Ge, J., Wu, J., Zhang, H., Wang, P.: New detection method for nucleoside triphosphates based on carbon dots: the distance-dependent singlet oxygen trapping. Anal. Chim. Acta 1031, 145–151 (2018)
Wang, Z., Zhou, X., Han, J., Xie, G., Liu, J.: DNA coated CoZn-ZIF metal-organic frameworks for fluorescent sensing guanosine triphosphate and discrimination of nucleoside triphosphates. Anal. Chim. Acta 1207, 339806 (2022)
Liu, Z., Chen, S., Liu, B., Wu, J., Zhou, Y., He, L., Ding, J., Liu, J.: Intracellular detection of ATP using an aptamer beacon covalently linked to graphene oxide resisting nonspecific probe displacement. Anal. Chem. 86, 12229–12235 (2014)
Lee, J., Park, C., Kim, Y., Park, S.: Signal enhancement in ATP bioluminescence to detect bacerial pahogens via heat treatment. BioChip J. J. 11, 287–293 (2017)
Acknowledgements
This research was supported by the National Research Foundation of Korea (NRF-2021R1C1C1009105) and the National Research Council of Science and Technology (NST) (CRC22021-500).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, DG., Lim, HJ., Lee, HY. et al. A Controlled Transcription-Driven Light-Up Aptamer Amplification for Nucleoside Triphosphate Detection. BioChip J 17, 487–495 (2023). https://doi.org/10.1007/s13206-023-00124-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13206-023-00124-0