Abstract
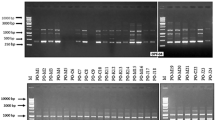
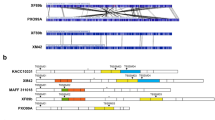
Brown spot disease, caused by Bipolaris oryzae, is one of the several disastrous diseases affecting rice. The brown spot fungus illustrates substantial pathogenic and genetic variability. To the best of our knowledge, extensive analysis utilizing specific SSR primers for B. oryzae genome is quite inadequate for the population structure and genetic diversity of Indian B. oryzae isolates. A total of 84 brown spot isolates were collected from rice-cultivating areas across southern and eastern Indian states, viz., Tamil Nadu, Andhra Pradesh, Odisha and Chhattisgarh. The pathogenicity and virulence characteristics of these isolates were assessed with the susceptible cultivar CR Dhan 201. Twelve genome-specific SSR markers of B. oryzae warranted the investigation of the population structure and genetic diversity among the isolates. These isolates were categorized based on their disease grade as highly virulent isolates (4 nos.), virulent isolates (8 nos.), moderately virulent isolates (47 nos.) and less virulent isolates (25 nos.). PCR amplification and DNA sequencing confirmed the isolates to be B. oryzae. PCR amplification and DNA sequencing confirmed the isolates to be B. oryzae. The SSR markers produced a total of 35 alleles with 1 to 4 alleles per locus with a gene diversity ranging between 0.00 and 0.687 and a major allele frequency variation of 0.425–0.975. The PIC value ranged from 0.00 to 0.638 having a mean value of 0.34. Cluster analysis technique was applied to group the brown spot isolates into four distinct clusters. Principal coordinate and structure analysis identified two genetic clusters of B. oryzae isolates for individual states with some degree of distinctness complying with their virulence. Analysis of molecular variance revealed more genetic variation within populations and less among populations. The study outcome would expedite the comprehension of genetic diversity of B. oryzae across the southern and eastern states of India. Furthermore, we anticipate its guidance in the development of more effective disease management strategies as well as in the generation of novel resistant varieties through marker-assisted breeding.





Similar content being viewed by others
References
Ahmadpour A, Castell-Miller C, Javan-Nikkhah M, Naghavi MR, Dehkaei FP, Leng Y, Puri KD, Zhong S (2018) Population structure, genetic diversity, and sexual state of the rice brown spot pathogen Bipolaris oryzae from three Asian countries. Plant Pathol 67:181–192. https://doi.org/10.1111/ppa.12714
Archana B, Kini KR, Prakash HS (2014) Genetic diversity and population structure among isolates of the brown spot fungus, Bipolaris oryzae, as revealed by inter-simple sequence repeats (ISSR). Afr J Biotechnol 13:238–244. https://doi.org/10.5897/AJB2013.12063
Ashfaq B, Arshad HMI, Atiq M, Yousaf S, Saleem K, Arshad A (2021) Biochemical profiling of resistant phenotypes against Bipolaris oryzae causing brown spot disease in rice. Front Agron 3:675895. https://doi.org/10.3389/fagro.2021.675895
Baranwal MK, Kotasthane A, Magculia N, Mukherjee PK, Savary S, Sharma AK, Singh HB, Singh US, Sparks AH, Variar M, Zaidi N (2013) A review on crop losses, epidemiology and disease. Eur J Plant Pathol 136:443–457
Boka A, Bouet A, Tiendrebeogo A, Kassankogno AI, Ouedraogo I, Nda GN, Denezon OD, Adiko A (2018) Pathogenic variability of Bipolaris oryzae causing leaf spot disease of rice in West Africa. Int J Phytopathol 7:103–110. https://doi.org/10.33687/phytopath.007.03.2643
Burgos MRG, Katimbang MLB, Paz MD, Beligan GA, Goodwin PH, Ona IP, Mauleon RP, Ardales EY, Cruz CV (2013) Genotypic variability and aggressiveness of Bipolaris oryzae in the Philippines. Eur J Plant Pathol 137:415–429
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Fajolu OL, Wadl PA, Vu AL, Gwinn KD, Scheffler BE, Trigiano RN, Ownley BH (2013) Development and characterization of simple sequence repeats for Bipolaris sorokiniana and cross transferability to related species. Mycologia 105:1164–1173. https://doi.org/10.3852/12-210
Gomez KA, Gomez AA (1984) Statistical procedure for agricultural research. Wiley, New York
Gupta V, Shamas N, Razdan VK, Sharma BC, Sharma R, Kaur K, Singh I, John D, Kumar A (2013) Foliar application of fungicides for the management of brown spot disease in rice (Oryza sativa L.) caused by Bipolaris oryzae. Afr J Agric Res 8:3303–3309
Harish S, Kumar DS, Radjacommare R, Ebenezar EG, Seetharaman K (2008) Use of plant extracts and biocontrol agents for the management of brown spot disease in rice. Biol Control 53:555–567. https://doi.org/10.1007/s10526-007-9098-9
Hau FC, Rush MC (1980) A system for inducing sporulation of Bipolaris oryzae. Plant Dis 64:788–789. https://doi.org/10.1094/PD-64-788
Hughes KW, Petersen RH, Lodge DJ, Bergemann SE, Baumgartner K, Tulloss RE, Lickey E, Cifuentes J (2013) Evolutionary consequences of putative intra-and interspecific hybridization in agaric fungi. Mycologia 105:1577–1594. https://doi.org/10.3852/13-041
IRRI (2013) Standard evaluation system (SES) of rice (revised), 5th edn. Philippines, Manila
Jaiganesh V, Kannan C (2019) Studies on the cultural characters and pathogenicity studies of brown leaf spot of rice caused by Helminthosporium oryzae. Plant Arch 19:585–587
Johnson DR, Percich JA (1992) Wild rice domestication, fungal brown spot disease, and the future of commercial production in Minnesota. Plant Dis 76:1193–1198. https://doi.org/10.1094/PD-76-1193
Kamal MM, Mia MAT (2009) Diversity and pathogenicity of the rice brown spot pathogen, Bipolaris oryzae (Breda de Haan) Shoem. in Bangladesh assessed by genetic fingerprint analysis. Bangl J Bot 38:119–125. https://doi.org/10.3329/bjb.v38i2.5135
Kandan A, Akhtar J, Singh B, Dixit D, Chand D, Roy A, Rajkumar S, Agarwal PC (2015) Molecular diversity of Bipolaris oryzae infecting Oryza sativa in India. Phytoparasitica 43:5–14. https://doi.org/10.1007/s12600-014-0430-5
Krupinsky JM, Berdahl JD, Schoch CL, Rossman AY (2004) Leaf spot on switch grass (Panicum virgatum), symptoms of a new disease caused by Bipolaris oryzae. Can J Plant Pathol 26:371–378. https://doi.org/10.1080/07060660409507155
Kumar A, Solanki IS, Akhtar J, Gupta V (2016) Morpho-molecular diversity of Bipolaris oryzae causing brown spot of paddy. Indian J Agric Sci 86:615–620
Kumari S, Kumar A, Rani S (2015) Morphological characterization of Bipolaris oryzae causing brown spot of paddy in Bihar. Int Educ Res J 1:185–187
Lee CH, Hou HH, Jong SJ (1984) Cytology of Helminthosporium oryzae. Mycopathologia 87:23–27. https://doi.org/10.1007/BF00436621
Librado P, Rozas J (2009) DnaSP v5: software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452. https://doi.org/10.1093/bioinformatics/btp187
Liu K, Muse SV (2005) PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21:2128–2129
Mew TW, Gonzales P (2002) A handbook of rice seedborne fungi. International Rice Research Institute/Science Publisher, Los Banos
Milgroom MG (2015) Recombination and randomly mating populations. In: Milgroom MG (ed) Population biology of plant pathogens: genetics, ecology, and evolution. American Phytopathological Society, St Paul, pp 147–184
Motlagh MRS, Anvari M (2010) Genetic variation in a population of Bipolaris oryzae based on RAPD-PCR in north of Iran. Afr J Biotechnol 9:5800–5804
Nazari S, Javan-Nikkhah M, Fotouhifar KB, Khosravi V, Alizadeh A (2015) Bipolaris species associated with rice plant: pathogenicity and genetic diversity of Bipolaris oryzae using rep-PCR in Mazandaran province of Iran. J Crop Prot 4:497–508
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York. https://doi.org/10.7312/nei-92038
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research an update. Bioinformatics 28:2537–2539. https://doi.org/10.1093/bioinformatics/bts460
Prabhukarthikeyan SR, Yadav MK, Anandan A, Aravindan S, Keerthana U, Raghu S, Baite MS, Parameswaran C, Panneerselvam P, Rath PC (2019) Bio-protection of brown spot disease of rice and insight into the molecular basis of interaction between Oryza sativa, Bipolaris oryzae and Bacillus amyloliquefaciens. Biol Control 137:104018. https://doi.org/10.1016/j.biocontrol.2019.104018
Pritchard JX, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Spring O, Thines M (2010) Molecular techniques for classification and diagnosis of plant pathogenic Oomycota. In: Gherbawy Y, Voigt K (eds) Molecular identification of fungi. Springer, Berlin. https://doi.org/10.1007/978-3-642-05042-8-2
Sultana S, Adhikary SK, Islam MM, Rahman SMM (2018) Evaluation of pathogenic variability based on leaf blotch disease development components of Bipolaris sorokiniana in Triticum aestivum and agroclimatic origin. Plant Pathol J 34:93–103. https://doi.org/10.5423/PPJ.OA.08.2017.0175
Surendhar M, Anbuselvam Y, Ivin J (2021) Status of rice brown spot (Helminthosporium oryza) management in India: a review. Agric Rev. https://doi.org/10.18805/ag.R-2111
Webster RK, Gunnell PS (1992) Compendium of rice diseases. The American Phytopathological Society, St. Paul
Yadav MK, Aravindan S, Raghu S, Prabhukarthikeyan SR, Keerthana U, Ngangkham U, Pramesh D, Banerjee A, Adak T, Kar MK, Parameswaran C, Deshmukh R, Tiwari JK, Mohanty MR, Rath PC (2019) Assessment of genetic diversity and population structure of Magnaporthe oryzae causing rice blast disease using SSR markers. Physiol Mol Plant Pathol 106:157–165. https://doi.org/10.1016/j.pmpp.2019.02.004
Brun S, Silar P (2010) In: Evolutionary biology-concepts, molecular and morphological evolution. In: Pontarotti P (ed). Springer, Berlin, pp 317–328
Lee FN (1992) Brown spot. In: Webster RK, Gunnell PS (eds) Compendium of rice diseases. APS, St. Paul, pp 14–17
Ou SH (1985) Fungus diseases-foliage diseases. In: Rice diseases, 2nd edn. Commonwealth Mycological Institute, Kew, pp 109–201
Perrier X, Jacquemoud–Collet JP (2006) DARwin software. http://darwin.cirad.fr/darwin
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W, Fungal Barcoding Consortium (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. PNAS 109:6241–6246
Umbrey Y, Divya M, Das T, Das S, Mahapatra S (2021) Isolation and identification of seed borne mycoflora associated with popular rice cultivars in North East India. J Cereal Res 13(Spl-1):43–50. https://doi.org/10.25174/2582-2675/2021/112852
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: PCR protocols. A guide to methods and applications. Academic Press, San Diego, pp 315–322. https://doi.org/10.1016/b978-0-12-372180-8.50042-1
Acknowledgements
The authors gratefully acknowledge ICAR-National Rice Research Institute, Cuttack, India for providing the necessary facilities.
Author information
Authors and Affiliations
Contributions
UK and MP contributed to conceptualization, methodology and writing––original draft; SRP was involved in writing––review and editing; RN and MKY performed data analysis and data curation; CP contributed to software and validation; MSB and SR carried out survey and methodology; MGR and SH carried out survey and processed samples; PP and PCR contributed to resources and data validation.
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Keerthana, U., Phalguni, M., Prabhukarthikeyan, S.R. et al. Elucidation of the population structure and genetic diversity of Bipolaris oryzae associated with rice brown spot disease using SSR markers. 3 Biotech 12, 281 (2022). https://doi.org/10.1007/s13205-022-03347-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03347-4