Abstract
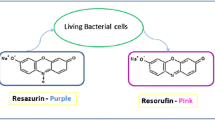
A new resazurin-based assay was evaluated and optimized using a microplate (384-well) format for high-throughput screening of antibacterial molecules against Klebsiella pneumoniae. Growth of the bacteria in 384-well plates was more effectively measured and had a > sixfold higher signal-to-background ratio using the resazurin-based assay compared with absorbance measurements at 600 nm. Determination of minimum inhibitory concentrations of the antibiotics revealed that the optimized assay quantitatively measured antibacterial activity of various antibiotics. An edge effect observed in the initial assay was significantly reduced using a 1-h incubation of the bacteria-containing plates at room temperature. There was an approximately 10% decrease in signal variability between the edge and the middle wells along with improvement in the assay robustness (Z′ = 0.99). This optimized resazurin-based assay is an efficient, inexpensive, and robust assay that can quantitatively measure antibacterial activity using a high-throughput screening system to assess a large number of compounds for discovery of new antibiotics against K. pneumoniae.




Similar content being viewed by others
References
Bogdanovich T, Adams-Haduch JM, Tian GB, Nguyen MH, Kwak EJ, Muto CA, Doi Y (2011) Colistin-resistant, Klebsiella pneumoniae carbapenemase (KPC)-producing Klebsiella pneumoniae belonging to the international epidemic clone ST258. Clin Infect Dis Off Publ Infect Dis Soc Am 53:373–376. https://doi.org/10.1093/cid/cir401
Butler MS, Blaskovich MA, Cooper MA (2016) Antibiotics in the clinical pipeline at the end of 2015. J Antibiot. https://doi.org/10.1038/ja.2016.72
Dostal SM, Fang Y, Guerrette JC, Scanlon TC, Griswold KE (2015) Genetically enhanced lysozyme evades a pathogen derived inhibitory protein. ACS Chem Biol 10:1110–1117. https://doi.org/10.1021/cb500976y
Elemam A, Rahimian J, Mandell W (2009) Infection with panresistant Klebsiella pneumoniae: a report of 2 cases and a brief review of the literature. Clin Infect Dis Off Publ Infect Dis Soc Am 49:271–274. https://doi.org/10.1086/600042
Franzblau SG et al (1998) Rapid, low-technology MIC determination with clinical Mycobacterium tuberculosis isolates by using the microplate Alamar Blue assay. J Clin Microbiol 36:362–366
Hanson M, Jordan LD, Shipelskiy Y, Newton SM, Klebba PE (2016) High-throughput screening assay for inhibitors of TonB-dependent iron transport. J Biomol Screen 21:316–322. https://doi.org/10.1177/1087057115613788
Kirchner S, Fothergill JL, Wright EA, James CE, Mowat E, Winstanley C (2012) Use of artificial sputum medium to test antibiotic efficacy against Pseudomonas aeruginosa in conditions more relevant to the cystic fibrosis lung Journal of visualized experiments. JoVE. https://doi.org/10.3791/3857
Lee J, Patel G, Huprikar S, Calfee DP, Jenkins SG (2009) Decreased susceptibility to polymyxin B during treatment for carbapenem-resistant Klebsiella pneumoniae infection. J Clin Microbiol 47:1611–1612. https://doi.org/10.1128/JCM.02466-08
Livermore DM, Mushtaq S, Warner M, Miossec C, Woodford N (2008) NXL104 combinations versus Enterobacteriaceae with CTX-M extended-spectrum beta-lactamases and carbapenemases. J Antimicrob Chemother 62:1053–1056. https://doi.org/10.1093/jac/dkn320
Lundholt BK, Scudder KM, Pagliaro L (2003) A simple technique for reducing edge effect in cell-based assays. J Biomol Screen 8:566–570. https://doi.org/10.1177/1087057103256465
Marchaim D et al (2011) Outbreak of colistin-resistant, carbapenem-resistant Klebsiella pneumoniae in metropolitan Detroit. Mich Antimicrob Agents Chemother 55:593–599. https://doi.org/10.1128/AAC.01020-10
Nairn BL et al (2017) Fluorescence high-throughput screening for inhibitors of TonB action. J Bacteriol. https://doi.org/10.1128/JB.00889-16
Palomino JC, Portaels F (1999) Simple procedure for drug susceptibility testing of Mycobacterium tuberculosis using a commercial colorimetic assay. Eur J Clin Microbiol Infect Dis Off Publ Eur Soc Clin Microbiol 18:380–383
Petrosillo N, Ioannidou E, Falagas ME (2008) Colistin monotherapy vs. combination therapy: evidence from microbiological, animal and clinical studies. Clin Microbiol Infect Off Publ Eur Soc Clin Microbiol Infect Dis 14:816–827. https://doi.org/10.1111/j.1469-0691.2008.02061.x
Ruzin A, Visalli MA, Keeney D, Bradford PA (2005) Influence of transcriptional activator RamA on expression of multidrug efflux pump AcrAB and tigecycline susceptibility in Klebsiella pneumoniae. Antimicrob Agents Chemother 49:1017–1022. https://doi.org/10.1128/AAC.49.3.1017-1022.2005
Scanlon TC, Dostal SM, Griswold KE (2014) A high-throughput screen for antibiotic drug discovery. Biotechnol Bioeng 111:232–243. https://doi.org/10.1002/bit.25019
Stachyra T, Levasseur P, Pechereau MC, Girard AM, Claudon M, Miossec C, Black MT (2009) In vitro activity of the {beta}-lactamase inhibitor NXL104 against KPC-2 carbapenemase and Enterobacteriaceae expressing KPC carbapenemases. J Antimicrob Chemother 64:326–329. https://doi.org/10.1093/jac/dkp197
Van den Driessche F, Rigole P, Brackman G, Coenye T (2014) Optimization of resazurin-based viability staining for quantification of microbial biofilms. J Microbiol Methods 98:31–34. https://doi.org/10.1016/j.mimet.2013.12.011
Zetts R (2016) Antibiotics currently in clinical development. The Pew Charitable Trusts. http://www.pewtrusts.org/en/multimedia/data-visualizations/2014/antibiotics-currently-in-clinical-development
Acknowledgements
We thank Kevin Pethe for invaluable advice on this research. This work was supported by National Research Foundation of Korea (NRF) grants funded by the Korean government (MSIP) (2012-00011) (NRF-2014K1A4A7A01074645), Gyeonggi-do, and KISTI.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
No conflict of interest declared.
Rights and permissions
About this article
Cite this article
Kim, H.J., Jang, S. Optimization of a resazurin-based microplate assay for large-scale compound screenings against Klebsiella pneumoniae . 3 Biotech 8, 3 (2018). https://doi.org/10.1007/s13205-017-1034-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-017-1034-9