Abstract
Combating antibiotic resistance requires discovery of novel antimicrobials effective against resistant bacteria. Herein, we present for the first time, pGLO plasmid transformed Escherichia coli HB 101 K 12 as novel model for screening of nanomaterial-based antimicrobial agents against β-lactamase resistance. E. coli HB 101 was transformed by pGLO plasmid in the presence of calcium chloride (50 mM; pH 6.1) aided by heat shock (0–42–0 °C). The transformed bacteria were grown on Luria–Bertani agar containing ampicillin (amp) and arabinose (ara). The transformed culture was able to grow in the presence of ampicillin and also exhibited fluorescence under UV light. Both untransformed and transformed bacteria were used for screening citrate-mediated nanosilver (CNS), aloin-mediated nanosilver (ANS), 11-α-keto-boswellic acid (AKBA)-mediated nanosilver (BNS); nanozinc oxide, nanomanganese oxide (NMO) and phytochemicals such as aloin and AKBA. Minimum inhibitory concentrations (MIC) were obtained by microplate method using ρ-iodo nitro tetrazolium indicator. All the compounds were effective against transformed bacteria except NMO and AKBA. Transformed bacteria exhibited reverse cross resistance against aloin. ANS showed the highest antibacterial activity with a MIC of 0.32 ppm followed by BNS (10.32 ppm), CNS (20.64 ppm) and NZO (34.83 ppm). Thus, pGLO plasmid can be used to induce resistance against β-lactam antibiotics and the model can be used for rapid screening of new antibacterial agents effective against resistant bacteria.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Beta-lactam antibiotics account for almost 50 % of global use of antibiotics (Walsh 2003; Elander 2003). Such an extensive use of β-lactam antibiotics has led to the emergence of resistance globally (Thomson and Bonomo 2005; Rice 2012). The transfer of resistance genes is mostly mediated by plasmids and transposons (Bennett 2008; Touchon et al. 2012) and occurs by conjugation, transduction or transformation. Resistance to β-lactam antibiotics occurs by different mechanisms, (Wilke et al. 2005; Fisher et al. 2005) of which β-lactamase mediated resistance is important in Gram-negative organisms. (Li et al. 1994; Poole 1994) Multidrug-resistant bacteria producing β-lactamases exhibit resistance towards broad range of β-lactam antibiotics, including third-generation cephalosporins and pose serious challenge to clinicians and limits treatment options (Jacoby and Price 2005; Paterson and Bonomo 2005).
Effective tackling of antimicrobial resistance requires better diagnostic techniques and novel drugs (Khanal et al. 2013). Traditionally, the discovery of new antimicrobial agents effective against resistant bacteria involves the use of pathogenic resistant strains isolated from clinical setting. Further, the presence of resistance genes needs to be confirmed through PCR or other molecular techniques. Ideally, a bacterial model for screening new compounds effective against resistant forms should be non-pathogenic and allow for rapid high throughput screening. We propose pGLO plasmid transformed Escherichia coli HB 101 K 12 as a novel model for screening candidate drug molecules effective against β-lactam antibiotics. The model has the advantages of being non-pathogenic and provides visual confirmation for the presence of resistance genes. Further, as both resistant and non-resistant strains of the same bacteria are available, the pattern of susceptibility can be studied.
pGLO plasmid consists of genes encoding for either kanamycin resistance or ampicillin resistance, green fluorescent protein (GFP) genes and arabinose operon repressor (Bassitu 2011). The resistance to ampicillin is conferred through the production of β-lactamase enzyme, which destroys β-lactam antibiotics. GFP is a fairly small protein (27 kDa) isolated from the jellyfish, Aequorea victoria and requires no cofactors or substrates for its expression (Chalfie et al. 1994). GFP emits green fluorescence at 509 nm providing visual confirmation with a hand held ultraviolet (UV) lamp or non-invasively with UV spectrometry. The modified GFP gene is quite stable and is mainly used for educational purposes to demonstrate genetic engineering to students (Mosher 2002). Several applications for pGLO plasmid and GFP were reported. GFP tagging for fluorescence-based random mutagenesis of IS2 transposase for structure–function analysis was reported (Lewis et al. 2011). A high throughput microfluidic biosensor using E. coli HB 101 and pGLO plasmid for the detection of specific substrates and for screening of antibiotic drugs was developed (Sun et al. 2011). pGLO plasmid was used for evaluating survival of Salmonella spp. and Yersinia enterocolitica bacteria in pig slurry treated with urea and ammonia (Bolton et al. 2012) and to engineer a sensitive biosensor to screen for enzyme activity in E. coli producing 3, 4 dihydroxy benzoate (Jha et al. 2014). In the recent years, several reports have been cited in the literature on antimicrobial properties of a variety of nanoscale materials against an array of pathogens (Prabha shetty et al. 2014; Supraja et al. 2015).
Materials and methods
Chemicals
Luria–Bertani (LB) agar, LB broth, Calcium chloride (50 mM; pH 6.1), Ampicillin (Amp) and L (+) Arabinose (Ara) were obtained from Bio-Rad, USA. Mueller–Hinton (MH) broth, ρ-iodo nitro tetrazolium (INT) and dimethyl sulfoxide (DMSO) were obtained from Hi-Media, India. Aloin was from Sigma-Aldrich, USA, and Acetyl-11-α-keto-β-boswellic acid (AKBA) was obtained from Natural Remedies, India.
Preparation of the nanoparticles used in this study
Silver nitrate (>99 %) and trisodium citrate dihydrate (99.0 %) were purchased from Sigma-Aldrich. Sodium citrate and silver nitrate (5:1) were mixed in a conical flask and aged for about 2 h. The solution was heated to 100 °C on slow heating. During the heating process, 2–3 drops of 0.01 M sodium borohydrate was added to the solution. Then it has been observed that the colour of the solution was changed gradually to yellow within a few minutes, indicating the formation of Ag nanoparticles. The solution was kept boiling for an additional 6 min and was allowed to cool. Nanozinc oxide and nanomanganese oxide particles were prepared using oxalate decomposition method (Prasad et al. 2012). Aloin-mediated nanosilver particles and AKBA-mediated nanosilver particles were prepared by using the method described by Chaitanya et al.
Transformation of Escherichia coli
The process of transformation of E. coli HB 101 K 12 using pGLO plasmid was carried out as per manufacturer’s instructions. Briefly, two to four colonies of the bacteria were suspended in 250 µL of 50 mM CaCl2. 10 µL of pGLO plasmid was added and incubated on ice for 10 min. Heat shock was provided by quickly transferring to a water bath at 42 °C for exactly 50 s and then rapidly transferring onto ice. After an incubation of 2 min, 250 µL of LB broth was added and the culture was incubated overnight at 37 °C. 100 µL of the culture was spread each on LB/Amp/Ara agar, LB/Amp and LB agar and incubated at 37 °C. The procedure was repeated for E. coli culture without the addition of pGLO plasmid to obtain negative controls.
Transformation efficacy
The transformation efficacy was calculated by counting the number of colonies on LB/Amp/Ara plates for the transformed culture using the following formula:
Minimum inhibitory concentration by microdilution method
The minimum inhibitory concentration (MIC) of various compounds was evaluated as per Clinical and Laboratory Standards Institute (CLSI 2006). A twofold dilution of CNS, ANS, BNS and ampicillin was made in 100 µL of Mueller–Hinton (MH) broth in a microplate. NMO, NZO, Aloin and AKBA were dissolved in DMSO and similar two-fold dilutions were carried out in MH broth. To each well, 50 µL of 1:10 diluted 0.5 Mc Farland units of bacterial suspension was added to provide a final concentration of 5 × 105 cfu/mL per well. Positive and negative controls for culture and broth were also maintained. The plates were covered with parafilm to prevent drying and incubated at 37 °C for 18 h. One hour before the completion of incubation, 50 µL of p-nitro iodo tetrazolium (INT) (2 mg/mL in distilled water) (Eloff 1998) was added to each well and the plates were incubated at 37 °C for another hour. The minimum inhibition concentration was defined as the minimum concentration of the compound, which inhibited visible growth of bacteria, evidenced by lack of development of any colour.
Dynamic light scattering (particle size) and zeta potential analysis
Dynamic light scattering (DLS) technique is one of the widely accepted techniques to measure the hydrodynamic diameter (HDD) of the particles in a hydrosol. The particle size measurements were carried out using Nanopartica SZ-100 (HORIBA). Zeta potential was also measured using the same instrument using electrical conducting cell.
Results and discussion
Particle size and zeta potential analysis
Particle size and zeta potential measurements were carried out to reveal the hydrodynamic diameter and electrostatic interaction of the prepared nanomaterials with their surrounding environment. The hydrodynamic diameter (HDD) of ANS (Fig. 1), BNS (Fig. 3) and CNS (Fig. 4) was recorded as 142.7, 19.6, 48.2 nm, respectively, which indicating the influence of bio-reducing agent on the size of the formed nanoparticles. Further, the zeta potentials of ANS and CNS particles were found to be −5.8 mV (Fig. 2) and −81 mV (Fig. 5), respectively, indicating the greater stability of CNS over ANS.
pGLO transformation
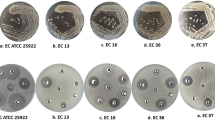
pGLO plasmid transformed E. coli exhibited an even lawn of growth on LB agar whereas 16 colonies were visible on LB/Amp and LB/Amp/Ara plates. A transformation efficacy of 100 cells/μg of pGLO plasmid was obtained. The colonies only on LB/Am/Ara plates showed fluorescence under long UV light (Fig. 6). The non-transformed E. coli showed growth only on LB agar.
Minimum inhibitory concentration
The MIC (ppm) of CNS, ANS, BNS, aloin, AKBA, nanomanganese, nanozinc oxide (NZnO), aloin, AKBA and standard ampicillin was determined for transformed and non-transformed bacteria (Table 1). Transformed E. coli was susceptible to CNS (20.64), ANS (0.32), BNS (10.32), NZO (34.83) and aloin (2.58); whereas resistance was observed against ampicillin (>666.7 ppm), NMO (>79.33) and AKBA (>73.33). Non-transformed E. coli was susceptible to all the compounds except nano manganese and AKBA. However, transformed E. coli (2.58) was more susceptible to aloin than untransformed E. coli (5.17).
The uptake of pGLO plasmid is enhanced in the presence of CaCl2 aided by heat shock, which increases the competence of the bacterial cells to take up extraneous genetic material (Cohen et al. 1972; Bergmans et al. 1981). The competence of the bacteria is also reported to be increased by other methods such as electroporation, (Dower et al. 1998) plasmid artificial modification (Yasui et al. 2009) and micro-shock waves (Divya et al. 2011). Transformation with pGLO plasmid expresses β-lactamase enzyme. Hence, visible growth was observed on LB/Amp and LB/Amp/Ara agar plates only in +pGLO group (Fig. 6). However, fluorescence was not detected in +pGLO colonies on LB/AMP agar (Fig. 6) due to the absence of arabinose, which is required for switching on GFP expression. GFP is expressed only in the presence of sugar arabinose, as the genes ara A and D are replaced with GFP genes. In the LB/AMP/ARA agar, due to the presence of arabinose, fluorescence was exhibited by +pGLO colonies confirming transformation.
In microdilution method for MIC determination, MH broth is recommended as the medium of choice for susceptibility testing for aerobic or facultative organisms. The broth demonstrates good batch-to-batch reproducibility, is low in sulphonamide, trimethoprim, and tetracycline inhibitors and yields satisfactory growth of most pathogens and could be supplemented to support the growth of fastidious bacteria (Ericsson and Sherris 1971). The resistance of +pGLO E. coli to ampicillin is due to the presence of β-lactamase gene, which produces β-lactamase that cleaves the amide bond of β-lactam ring rendering ampicillin ineffective (Livermore 1995; Poole 2004). Among the compounds screened, ANS showed highest antibacterial activity against resistant bacteria followed by CNS and BNS. Similar observations of potent antibacterial activity of ANS against Staphylococcus aureus with an MIC of 21.8 ng/mL were reported (Kumar et al. 2013, 2014). Several authors evaluated the antibacterial activity of citrate mediated silver nanoparticles against E. coli O157:H7 and observed a MIC of 12.43 ppm; (Petrus et al. 2011); E. coli (ATCC8739) with an MIC of 160 ppm (El-Kheshen and El-Rab 2012).
Apart from nanosilver particles, NZnO also exhibited antibacterial activity against transformed and non-transformed bacteria. Nanozinc oxide was shown to have a MIC of 0.1 ppm against E. coli K88 strain (Wang et al. 2012). Liu et al. (2009) observed complete inhibition of microbial growth at a concentration of 12 mmol/L with nanozinc oxide against E. coli O157:H7. Nano manganese oxide on the other hand was not effective against both transformed and non-transformed E. coli. However, earlier (Al-Hazmi et al. 2012) nano magnesium oxide wires were shown to possess bacteriostatic effect on E. coli and Bacillus spp. at concentrations above 100 ppm. As low concentration of MNO (79.33 ppm) was used in this study, the antibacterial activity of MnO could not be detected.
Aloin exhibited higher antibacterial activity against both transformed and non-transformed bacteria. Similar results have been reported previously (Tian et al. 2003; Minale et al. 2014) against E. coli. Aloin had an MIC of 10 ppm against E. coli CD/99/1, K88, K99, LT 37, ROW 7/12, 3:37 C, 306, and 872 strains. In this study, transformed E. coli exhibited enhanced susceptibility to aloin than non-transformed E. coli. The increased susceptibility could be explained by ‘negative cross-resistance’ or ‘collateral sensitivity’, where the induction of resistance to one compound enhances the toxicity to other compounds (Li et al. 2002; Palmer et al. 2010). Previously, similar phenomenon was observed through the increased susceptibility of E. coli to fusaric acid consequent to development of resistance against tetracyclines by modification of efflux pumps (Bochner et al. 1980).
Another phytochemical, AKBA showed no antibacterial activity against both transformed and non-transformed E. coli. This is due to the presence of lipophilic outer membrane in Gram-negative bacteria, which acts as a hydrophilic permeability barrier for the entry of hydrophobic compounds such as AKBA (Raja et al. 2011; Hancock 1997; Helander et al. 1998). Similar observations were made in studies dealing with antibacterial activity of other hydrophobic compounds against Gram-negative bacteria (Trombetta et al. 2005; Gallucci et al. 2009).
Conclusion
Antibiotic resistance is an expanding menace and this needs immediate attention by the scientific community. Nanoscale materials are the new generation of materials which exhibit novel antimicrobial properties, noble nanoscale materials in particular, against an array of microbes. Consequently, methods need to be developed for the rapid assessment of antimicrobial efficacy of materials with different size, shape and surface coatings. We reported for the first time that pGLO plasmid transformed E. coli HB 101 K 12 serves as a novel model for screening of new antimicrobial compounds including nanomaterial based antimicrobials which were effective against β-lactam resistance. The model has the advantages of being non-pathogenic and provides visual confirmation for the presence of resistance genes.
References
Al-Hazmi F, Alnowaiser F, Al-Ghamdi AA, Al-Ghamdi AA, Aly MM, Reem M et al (2012) A new large-scale synthesis of magnesium oxide nanowires: structural and antibacterial properties. Superlattices Microstruct 52:200–209
Bassitu EA (2011) pGLO mutagenesis: a laboratory procedure in molecular biology for biology students. Biochem Mol Biol Educ 2011(39):432–439
Bennett PM (2008) Plasmid encoded antibiotic resistance: acquisition and transfer of antibiotic resistance genes in bacteria. Br J Pharmacol 153:S347–S357
Bergmans HEN, van Die IM, Hoekstra WPM (1981) Transformation in E. coli: stages in the process. J Bacteriol 146:564–570
Bochner BR, Huang HC, Schieven GL, Ames BN (1980) Positive selection for loss of tetracycline resistance. J Bacteriol 143:926–933
Bolton DJ, Ivory C, McDowell DA (2012) The effect of urea and ammonia treatments on the survival of Salmonella spp. and Yersinia enterocolitica in pig slurry. J Appl Microbiol 114:134–140
Chalfie M, Tu Y, Euskirchen G, Ward W, Prasher D (1994) GFP as a marker for gene expression. Science 263:802–805
CLSI. Clinical and Laboratory Standards Institute, USA (2006) Methods for Dilution antimicrobial susceptibility tests for bacteria that grow aerobically; approved standard. Seventh Edition. CLSI document M7-A7
Cohen SN, Chang ACY, Hsu L (1972) Non-chromosomal antibiotic resistance in bacteria: genetic transformation of Escherichia coli by R-Factor DNA. Proc Natl Acad Sci USA 69:2110–2114
Divya PG, Anish RV, Jagadesh G, Chakravortty D (2011) Bacterial transformation using micro-shock waves. Anal Biochem 419:219–301
Dower WJ, Miller J, Ragsdale CW (1998) High efficiency transformation of E. coli by high voltage electroporation. Nucl Acids Res 16:6127–6145
Elander RP (2003) Industrial production of beta-lactam antibiotics. Appl Microbiol Biotechnol 61:385–392
El-Kheshen AA, El-Rab SFG (2012) Effect of reducing and protecting agents on size of silver nanoparticles and their anti-bacterial activity. Der Pharm Chem 4:53–65
Eloff JN (1998) A sensitive and quick microplate method to determine the minimal inhibitory concentration of plant extracts for bacteria. Plant Med 64:711–713
Ericsson HM, Sherris JC (1971) Antibiotic sensitivity testing. Report of an international collaborative study. Acta Pathol Microbiol Scand B Microbiol Immunol 217:1–90
Fisher JF, Meroueh SO, Mobashery S (2005) Bacterial resistance to betalactam antibiotics: compelling opportunism, compelling opportunity. Chem Rev 105:395–424
Gallucci MN, Oliva M, Casero C, Dambolena J, Luna A, Zygadlo J et al (2009) Antimicrobial combined action of terpenes against the food-borne microorganisms Escherichia coli, Staphylococcus aureus and Bacillus cereus. Flavour Fragr J 24:348–354
Hancock RE (1997) The bacterial outer membrane as a drug barrier. Trends Microbiol 5:37–42
Helander IM, Alakomi HL, Latva-Kala K, Mattila-Sandholm T, Pol I, Smid EJ et al (1998) Characterization of the action of selected essential oil components on Gram-negative bacteria. J Agric Food Chem 46:3590–3595
Jacoby GA, Price LSM (2005) The new beta-lactamases. N Engl J Med 352:380–391
Jha RK, Kern TL, Fox DT, Strauss (2014) Engineering an Acinetobacter regulon for biosensing and high-throughput enzyme screening in E. coli via flow cytometry. Nucl Acids Res 42:8150–8160
Khanal S, Joshi DR, Bhatta DR, Devakota U, Pokhrel BM (2013) β-Lactamase-producing multidrug-resistant bacterial pathogens from tracheal aspirates of intensive care unit patients at national institute of neurological and allied sciences, Nepal. ISRN Microbiol. Article ID 847569, pp 5. http://dx.doi.org/10.1155/2013/847569
Kumar TVC, Muralidhar Y, Prasad PE, Prasad TN, Raj MA (2013) Evaluation of therapeutic potential of nanosilver particles synthesised using aloin in experimental murine mastitis model. IET Nanobiotechnol 7:78–82
Kumar TVC, Prasad TN, Adilaxmamma K, Alpha Raj M, Muralidhar Y, Prasad PE (2014) Novel synthesis of nanosilver particles using plant active principle aloin and evaluation of their cytotoxic effect against Staphylococcus aureus. Asian Pac J Trop Dis 4(Suppl 1):S92–S96
Lewis LA, Astatke M, Umekubo PT, Alvi S, Saby R, Afrose J (2011) Soluble expression, purification and characterization of the full length IS2 Transposase. Mob DNA 2:14
Li XZ, Ma D, Livermore DM, Nikaido H (1994) Role of efflux pump(s) in intrinsic resistance of Pseudomonas aeruginosa: active efflux as a contributing factor to beta-lactam resistance. Antimicrob Agents Chemother 38:1742–1752
Li Q, Lee JY, Castillo R, Hixon MS, Pujol C, Doppalapudi VR et al (2002) NB2001, a novel antibacterial agent with broad-spectrum activity and enhanced potency against beta-lactamase-producing strains. Antimicrob Agents Chemother 46:1262–1268
Liu Y, He L, Mustapha A, Li H, Hu ZQ, Kin M (2009) Antibacterial activities of zinc oxide nanoparticles against Escherichia coli O157:h7. J Appl Microbiol 107:1193–1201
Livermore DM (1995) Beta-Lactamases in laboratory and clinical resistance. Clin Microbiol Rev 8:557–584
Minale G, Bisrat D, Asres K, Mazumder A (2014) In vitro antimicrobial activities of anthrones from the leaf latex of Aloe sinana Reynolds. Int J Green Pharm 8:7–12
Mosher R (2002) Using pGLO to demonstrate the effects of catabolite repression on gene expression in E coli. Bioscience 28:17–23
Palmer AC, Angelino E, Kishony R (2010) Chemical decay of an antibiotic inverts selection for resistance. Nat Chem Biol 6:105–107
Paterson DL, Bonomo RA (2005) Extended-spectrum β-lactamases: a clinical update. Clin Microbiol Rev 18:657–686
Petrus EM, Tinakumari S, Chai LC, Ubong A, Tunung R, Elexson N et al (2011) A study on the minimum inhibitory concentration and minimum bactericidal concentration of Nano Colloidal Silver on food-borne pathogens. Int Food Res J 18:55–66
Poole K (1994) Bacterial multidrug resistance: emphasis on efflux mechanisms and Pseudomonas aeruginosa. J Antimicrob Chemother 34:453–456
Poole K (2004) Resistance to beta-lactam antibiotics. Cell Mol Life Sci 61:2200–2223
Prabha S, Supraja N, Garud M, Prasad TNVKV (2014) Synthesis, characterization and antimicrobial activity of Alstonia scholaris bark-extract mediated silver nanoparticles. J Nanostruct Chem 4:161–170
Prasad TNVKV, Sudhakar P, Sreenivasulu Y, Latha P, Munaswamy V, Raja Reddy K, Sreeprasad TS, Sajanlal PR, Pradeep T (2012) Effect of nanoscale zinc oxide particles on the germination, growth and yield of peanut. J Plant Nutri 35(6):905–927
Raja AF, Ali F, Khan IA, Shawl AS, Arora DS, Shah BA et al (2011) Antistaphylococcal and biofilm inhibitory activities of acetyl-11-keto-β-boswellic acid from Boswellia serrata. BMC Microbiol 11:54
Rice LB (2012) Mechanisms of resistance and clinical relevance of resistance to beta-lactams, glycopeptides, and fluoroquinolones. Mayo Clin Proc 87:198–208
Sun P, Liu Y, Sha J, Zhang Z, Tu Q, Chen P et al (2011) High-throughput microfluidic system for long-term bacterial colony monitoring and antibiotic testing in zero-flow environments. Biosens Bioelectron 26:1993–1999
Supraja N, Prasad TNVKV, Giridhara Krishna T, David E (2015) Synthesis, characterization and evaluation of the efficacy of Boswellia ovalifoliolata stem bark extract mediated zinc nanoparticles. Appl Nanosci. doi:10.1007/s13204-015-0472-0
Thomson JM, Bonomo RA (2005) The threat of antibiotic resistance in Gram-negative pathogenic bacteria: beta-lactams in peril! Curr Opin Microbiol 8:518–524
Tian B, Hua YJ, Ma XQ, Wang GL (2003) Relationship between antibacterial activity of aloe and its anthaquinone compounds. Zhongguo Zhong Yao Za Zhi 28:1034–1037
Touchon M, Charpentier S, Pognard D, Picard B, Arlet G, Rocha EP et al (2012) Antibiotic resistance plasmids spread among natural isolates of Escherichia coli in spite of CRISPR elements. Microbiology 158:2997–3004
Trombetta D, Castelli F, Sarpietro MG, Venuti V, Cristani M, Daniele C (2005) Mechanisms of antibacterial action of three monoterpenes. Antimicrob Agents Chemother 49:2474–2478
Walsh C (2003) Where will new antibiotics come from? Nat Rev Microbiol 1:65–70
Wang C, Liu LL, Zhang AT, Xie P, Lu JJ, Zou XT (2012) Antibacterial effects of zinc oxide nanoparticles on Escherichia coli K 88. Afr J Biotech 11:10248–10254
Wilke MS, Lovering AL, Strynadka NC (2005) Beta-lactam antibiotic resistance: a current structural perspective. Curr Opin Microbiol 8:525–533
Yasui K, Kano Y, Tanaka K, Watanbe K, Shimizu-Kadota M, Yoshikawa H et al (2009) Improvement of bacterial transformation efficiency using plasmid artificial modification. Nucl Acids Res 37:e3
Acknowledgments
Authors are thankful to the authorities of Acharya N G Ranga Agricultural University for providing research facilities at Institute of Frontier Technology, Regional Agricultural Research Station, Tirupati, to carryout the part of the research work.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Raj, M.A., Muralidhar, Y., Sravanthi, M. et al. First report on rapid screening of nanomaterial-based antimicrobial agents against β-lactamase resistance using pGLO plasmid transformed Escherichia coli HB 101 K-12. Appl Nanosci 6, 941–949 (2016). https://doi.org/10.1007/s13204-015-0506-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13204-015-0506-7