Abstract
Background
Pancreatic cancer presents extremely poor prognosis due to the difficulty of early diagnosis, low resection rate, and high rates of recurrence and metastasis. Immune checkpoint blockades have been widely used in many cancer types but showed limited efficacy in pancreatic cancer. The current study aimed to evaluate the landscape of tumor microenvironment (TME) of pancreatic cancer and identify the potential markers of prognosis and immunotherapy efficacy which might contribute to improve the targeted therapy strategy and efficacy in pancreatic cancer in the context of predictive, preventive, and personalized medicine (PPPM).
Methods
In the current study, a total of 382 pancreatic samples from the datasets of Gene Expression Omnibus (GEO) and The Cancer Genome Atlas (TCGA) were selected. LM22 gene signature matrix was applied to quantify the fraction of immune cells based on “CIBERSORT” algorithm. Weighted Gene Co-expression Network Analysis (WGCNA) and Molecular Complex Detection (MCODE) algorithm was applied to confirm the hub-network of immune-resistance phenotype. A nomogram model based on COX and Logistic regression was constructed to evaluate the prognostic value and the predictive value of hub-gene in immune-response. The role of transmembrane protein 92 (TMEM92) in regulating cell proliferation was evaluated by MTS assay. Western blot and Real-time PCR were applied to assess the biological effects of PD-L1 inhibition by TMEM92. Moreover, the effect of TMEM92 in immunotherapy was evaluated with PBMC co-culture and by MTS assay.
Results
Two tumor-infiltrating immune cell (TIIC) phenotypes were identified and a weighted gene co-expression network was constructed to confirm the 167 gene signatures correlated with immune-resistance TIIC subtype. TMEM92 was further identified as a core gene of 167 gene signature network based on MCODE algorithm. High TMEM92 expression was significantly correlated with unfavorable prognosis, characterizing by immune resistance. A nomogram model and external validation confirmed that TMEM92 was an independent prognostic factor in pancreatic cancer. An elevated tumor mutation burden (TMB), mostly is consistent with commonly mutations of KRAS and TP53, was found in the high TMEM92 group. The predictive role of TMEM92 in immunotherapeutic response was also confirmed by IMvigor210 datasets. In addition, the specific biological roles of TMEM92 in cancer was explored in vitro. The results showed that abnormal overexpression of TMEM92 was significantly associated with the poor survival rate of pancreatic cancer. Moreover, we demonstrated that TMEM92 inhibit tumour immune responses of the anti-PD-1 antibody with PBMC co-culture.
Conclusion
The current study explored for the first time the immune-resistance phenotype of pancreatic cancer and identified TMEM92 as an innovative marker in predicting clinical outcomes and immunotherapeutic efficacy. These findings not only help to recognize high-risk and immune-resistance population which could be supplied targeted prevention, but also provide personalized medical services by intervening TMEM92 function to improve the prognosis of pancreatic cancer. In addition, the biological role of TMEM92 might reveal the potential molecular mechanisms of pancreatic cancer and lead to a novel sight for development of a PPPM approach for pancreatic cancer management.






Similar content being viewed by others
Data availability
The datasets generated and/or analyzed during the current study are available in the TCGA, GEO, and IMvigor210 repository: TCGA (https://portal.gdc.cancer.gov); GEO (GSE57495: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE57495; GSE62452:https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE62452; GSE85916: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE85916); IMvigor210 (http://research-pub.gene.com/IMvigor210CoreBiologies).
Code availability
All software applications used are included in this article.
Abbreviations
- TME:
-
Tumor microenvironment
- TIIC:
-
Tumor-infiltrating immune cells
- WGCNA:
-
Weighted gene co-expression network analysis
- MCODE:
-
Molecular complex detection
- GEO:
-
Gene Expression Omnibus
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- PD-1:
-
Programmed cell death protein 1
- PD-L1:
-
Programmed cell death ligand 1
- TMEM92:
-
Transmembrane protein 92
- TMB:
-
Tumor mutation burden
References
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics. CA Cancer J Clin. 2021;71(1):7–33.
Park W, Chawla A, O’Reilly EM. Pancreatic cancer: a review. JAMA. 2021;326(9):851–62.
Grech G, Zhan X, Yoo BC, Bubnov R, Hagan S, Danesi R, et al. EPMA position paper in cancer: current overview and future perspectives. EPMA J. 2015;6(1):9.
Nagaraju GP, Malla RR, Basha R, Motofei IG. Contemporary clinical trials in pancreatic cancer immunotherapy targeting PD-1 and PD-L1. Semin Cancer Biol. 2021; S1044–579X(21)00270–4.
Uhlitz F, Zamarin D. Rejuvenating dysfunctional T cells in ovarian cancer: CD28 is the license to kill. Cancer Cell. 2021;S1535–6108(21):00562–6.
Zhang B, Vogelzang A, Miyajima M, Sugiura Y, Wu Y, Chamoto K, et al. B cell-derived GABA elicits IL-10+ macrophages to limit anti-tumour immunity. Nature. 2021;599(7885):471–6.
Au L, Hatipoglu E, Robert de Massy M, Litchfield K, Beattie G, Rowan A, et al. TRACERx Renal Consortium. Determinants of anti-PD-1 response and resistance in clear cell renal cell carcinoma. Cancer Cell. 2021;39(11):1497–518.
Nguyen NT, Huang K, Zeng H, Jing J, Wang R, Fang S, et al. Nano-optogenetic engineering of CAR T cells for precision immunotherapy with enhanced safety. Nat Nanotechnol. 2021;16(12):1424–34.
Scheiner B, Pomej K, Kirstein MM, Hucke F, Finkelmeier F, Waidmann O, et al. Prognosis of patients with hepatocellular carcinoma treated with immunotherapy - development and validation of the CRAFITY score. J Hepatol. 2021;76(2):353–63.
Ribas A, Wolchok JD. Cancer immunotherapy using checkpoint blockade. Science. 2018;359(6382):1350–5.
Mazur PK, Siveke JT. Genetically engineered mouse models of pancreatic cancer: unravelling tumour biology and progressing translational oncology. Gut. 2012;61(10):1488–500.
Stromnes IM, Hulbert A, Pierce RH, Greenberg PD, Hingorani SR. T-cell localization, activation, and clonal expansion in human pancreatic ductal adenocarcinoma. Cancer Immunol Res. 2017;5(11):978–91.
Jiang ZB, Huang JM, Xie YJ, Zhang YZ, Chang C, Lai HL, et al. Evodiamine suppresses non-small cell lung cancer by elevating CD8+ T cells and downregulating the MUC1-C/PD-L1 axis. J Exp Clin Cancer Res. 2020;39(1):249.
Schmit K, Michiels C. TMEM proteins in cancer: a review. Front Pharmacol. 2018;9:1345.
Koteluk O, Bielicka A, Lemańska Ż, Jóźwiak K, Klawiter W, Mackiewicz A, et al. The landscape of transmembrane protein family members in head and neck cancers: their biological role and diagnostic utility. Cancers (Basel). 2021;13(19):4737.
Carroll J, He J, Ding S, Fearnley IM, Walker JE. TMEM70 and TMEM242 help to assemble the rotor ring of human ATP synthase and interact with assembly factors for complex I. Proc Natl Acad Sci U S A. 2021;118(13):e2100558118.
Luo S, Wang X, Bai M, Jiang W, Zhang Z, Chen Y, et al. The conserved autoimmune-disease risk gene TMEM39A regulates lysosome dynamics. Proc Natl Acad Sci U S A. 2021;118(6):e2011379118.
Duan J, Qian Y, Fu X, Chen M, Liu K, Liu H, et al. TMEM106C contributes to the malignant characteristics and poor prognosis of hepatocellular carcinoma. Aging (Albany NY). 2021;13(4):5585–606.
Rao J, Wu X, Zhou X, Deng R, Ma Y. TMEM205 is an independent prognostic factor and is associated with immune cell infiltrates in hepatocellular carcinoma. Front Genet. 2020;11:575776.
Listing H, Mardin WA, Wohlfromm S, Mees ST, Haier J. MiR-23a/-24-induced gene silencing results in mesothelial cell integration of pancreatic cancer. Br J Cancer. 2015;112(1):131–9.
Yu G, Wang LG, Han Y, He QY. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS. 2012;16(5):284–7.
Zhang S, Zang D, Cheng Y, Li Z, Yang B, Guo T. Identification of key gene and pathways for the prediction of peritoneal metastasis of gastric cancer by co-expression analysis. J Cancer. 2020;11(10):3041–51.
Hazra A, Gogtay N. Biostatistics series module 3: comparing groups: numerical variables. Indian J Dermatol. 2016;61(3):251–60.
Camp RL, Dolled-Filhart M, Rimm DL. X-tile: a new bio-informatics tool for biomarker assessment and outcome-based cut-point optimization. Clin Cancer Res. 2004;10(21):7252–9.
Xu L, Zhang Y, Liu J, Qu J, Hu X, Zhang F, et al. TRAIL-activated EGFR by Cbl-b-regulated EGFR redistribution in lipid rafts antagonises TRAIL-induced apoptosis in gastric cancer cells. Eur J Cancer. 2012;48(17):3288–99.
Zang D, Zhang C, Li C, Fan Y, Li Z, Hou K, et al. LPPR4 promotes peritoneal metastasis via Sp1/integrin α/FAK signaling in gastric cancer. Am J Cancer Res. 2020;10(3):1026–44.
Marcq E, Audenaerde JRV, Waele J, Jacobs J, Loenhout JV, Cavents G, et al. Building a bridge between chemotherapy and immunotherapy in malignant pleural mesothelioma: investigating the effect of chemotherapy on immune checkpoint expression. Int J Mol Sci. 2019;20(17):4182.
Marcq E, Van Audenaerde JRM, De Waele J, Merlin C, Pauwels P, van Meerbeeck JP, et al. The search for an interesting partner to combine with PD-L1 blockade in mesothelioma: focus on Tim-3 and LAG-3. Cancers. 2021;13(2):282.
Tanaka A, Sakaguchi S. Regulatory T cells in cancer immunotherapy. CELL RES. 2017;27(1):109–18.
Jairath NK, Farha MW, Jairath R, Harms PW, Tsoi LC, Tejasvi T. Prognostic value of intratumoral lymphocyte-to-monocyte ratio and M0 macrophage enrichment in tumor immune microenvironment of melanoma. Melanoma Manag. 2020; 7(4):Mmt51.
Hugo W, Zaretsky JM, Sun L, Song C, Moreno BH, Hu-Lieskovan S, et al. Genomic and transcriptomic features of response to anti-PD-1 therapy in metastatic melanoma. Cell. 2016;165(1):35–44.
Ayers M, Lunceford J, Nebozhyn M, Murphy E, Loboda A, Kaufman DR, et al. IFN-γ-related mRNA profile predicts clinical response to PD-1 blockade. J Clin Invest. 2017;127(8):2930–40.
Snyder A, Makarov V, Merghoub T, Yuan J, Zaretsky JM, Desrichard A, et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med. 2014;371(23):2189–99.
Rizvi NA, Hellmann MD, Snyder A, Kvistborg P, Makarov V, Havel JJ, et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science (New York, NY). 2015;348(6230):124–8.
Mayakonda A, Lin DC, Assenov Y, Plass C, Koeffler HP. Maftools: efficient and comprehensive analysis of somatic variants in cancer. Genome Res. 2018;28(11):1747–56.
Kleeff J, Korc M, Apte M, La Vecchia C, Johnson CD, Biankin AV, et al. Pancreatic cancer. Nat Rev Dis Primers. 2016;2:16022.
Dreyer SB, Chang DK, Bailey P, Biankin AV. Pancreatic cancer genomes: implications for clinical management and therapeutic development. Clin Cancer Res. 2017;23(7):1638–46.
Golubnitschaja O, Baban B, Boniolo G, Wang W, Bubnov R, Kapalla M, et al. Medicine in the early twenty-first century: paradigm and anticipation - EPMA position paper 2016. EPMA J. 2016;7(1):23.
Foley K, Kim V, Jaffee E, Zheng L. Current progress in immunotherapy for pancreatic cancer. Cancer Lett. 2016;381(1):244–51.
Pico de Coaña Y, Choudhury A, Kiessling R. Checkpoint blockade for cancer therapy: revitalizing a suppressed immune system. Trends Mol Med. 2015;21(8):482–91.
Morrison AH, Byrne KT, Vonderheide RH. Immunotherapy and prevention of pancreatic cancer. Trends Cancer. 2018;4(6):418–28.
Ino Y, Yamazaki-Itoh R, Shimada K, Iwasaki M, Kosuge T, Kanai Y, et al. Immune cell infiltration as an indicator of the immune microenvironment of pancreatic cancer. Br J Cancer. 2013;108(4):914–23.
Mahajan UM, Langhoff E, Goni E, Costello E, Greenhalf W, Halloran C, et al. Immune cell and stromal signature associated with progression-free survival of patients with resected pancreatic ductal adenocarcinoma. Gastroenterology. 2018;155(5):1625–39.
Stromnes IM, Hulbert A, Pierce RH, Greenberg PD, Hingorani SR. T-cell localization, activation, and clonal expansion in human pancreatic ductal adenocarcinoma. Cancer Immunol Res. 2017;5(11):978–91.
Zhang X, He Y, Jiang Y, Bao Y, Chen Q, Xie D, et al. TMEM229A suppresses non-small cell lung cancer progression via inactivating the ERK pathway. Oncol Rep. 2021;46(2):176.
Shiraishi T, Ikeda K, Tsukada Y, Nishizawa Y, Sasaki T, Ito M, et al. High expression of TMEM180, a novel tumour marker, is associated with poor survival in stage III colorectal cancer. BMC Cancer. 2021;21(1):302.
Lin MZ, Teng LL, Sun XL, Zhang LP, Chen F, Yu LJ. Transmembrane protein 92 performs a tumor-promoting function in breast carcinoma by contributing to the cell growth, invasion, migration and epithelial-mesenchymal transition. Tissue Cell. 2020;67:101415.
Buscail L, Bournet B, Cordelier P. Role of oncogenic KRAS in the diagnosis, prognosis and treatment of pancreatic cancer. Nat Rev Gastroenterol Hepatol. 2020;17(3):153–68.
Li J, Byrne KT, Yan F, Yamazoe T, Chen Z, Baslan T, et al. Tumor cell-intrinsic factors underlie heterogeneity of immune cell infiltration and response to immunotherapy. Immunity. 2018;49(1):178–93.
Zhang X, Shi M, Chen T, Zhang B. Characterization of the immune cell infiltration landscape in head and neck squamous cell carcinoma to aid immunotherapy. Mol Ther Nucleic Acids. 2020;22:298–309.
Zeng D, Li M, Zhou R, Zhang J, Sun H, Shi M, et al. Tumor microenvironment characterization in gastric cancer identifies prognostic and immunotherapeutically relevant gene signatures. Cancer Immunol Res. 2019;7(5):737–50.
Funding
This work was supported by Science and Technology Plan Project of Liaoning Province (2021-BS-102); National Natural Science Foundation of China (81801661).
Author information
Authors and Affiliations
Contributions
Guan Wang and Simeng Zhang designed the experiments. Guan Wang, Simeng Zhang, and Qiaoyun Chu wrote the manuscript. Mengzhu Lv and Ce Li performed the experiments. Simeng Zhang and Xing Wan performed the bioinformatics analysis. All authors contributed to the study design and data interpretation and have reviewed the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Data retrieved from the GEO and TCGA controlled-access database was collected using tumors from patients who provided informed consent based on guidelines laid out by the GEO and TCGA Ethics, Law and Policy Group. This study was approved by the Human Ethics Review Committee of China Medical University, and all procedures were conducted in accordance with ethical principles (protocol #: 2015PS63K).
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13167_2022_287_MOESM1_ESM.jpg
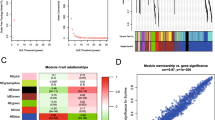
Supplementary file1 (JPG 3538 KB) Supplementary Figure 1. (A-D) Consensus matrixes of 382 pancreatic cancer samples for each k (k = 2–5) in classifying TIIC clusters, displaying the clustering stability using 1000 iterations of hierarchical clustering. (E) 2 pairs of gene modules were merged according to their similarity base on the threshold (Red line). (F) The genedendrogram was constructed by hierarchical clustering base on dissTOM of genes. (G) The correlation of epigengenes in green module with TIIC cluster trait. (H-I) Probability of survival at 1 (H) and 3 (I) years in the total cohort.
Rights and permissions
About this article
Cite this article
Zhang, S., Wan, X., Lv, M. et al. TMEM92 acts as an immune-resistance and prognostic marker in pancreatic cancer from the perspective of predictive, preventive, and personalized medicine. EPMA Journal 13, 519–534 (2022). https://doi.org/10.1007/s13167-022-00287-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13167-022-00287-0