Abstract
Oceanic islands have long been considered engines of differentiation and speciation for terrestrial organisms. Here we investigated colonisation and radiation processes in the Madeira Archipelago and the Canary Islands of the Andrena wollastoni group of bees (subgenus Micrandrena), which comprises six endemic species and five endemic subspecies on the islands. Mitochondrial COI sequences support the monophyly of the four species of the Canary Islands and the two species of the Madeira Archipelago and suggest a relatively young age for all taxa. The data do not support a simple stepping-stone model (eastern-western colonisation from the mainland, with splitting into new taxa), but suggest Andrena gomerensis (extant on La Gomera and La Palma) or its ancestor as the basal lineage from which all other taxa evolved. Andrena lineolata (Tene-rife) or its putative ancestor (A. gomerensis) is sister to A. dourada (Porto Santo), A. catula (Gran Canaria), and A. acuta (also Tenerife). Andrena dourada (Porto Santo) and A. wollastoni (Madeira Island) are sister species. Morphologically and morphometrically defined subspecies were not distinguishable with COI DNA sequences. Colonisation likely led from the Canary Islands to the Madeira Archipelago and not from the mainland directly to the latter.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oceanic islands represent excellent systems to answer questions of biogeography, phylogeography, and evolutionary biology. Compared to isolated oceanic islands, archipelagos are much more complex, e.g. due to the mostly different ages of the islands and their relative proximity to each other and to the mainland (MacArthur & Wilson, 1967). Island radiations of animal and plant species provide the means to study the effects of ecological and evolutionary forces and to infer colonisation history, leading to currently observed patterns of differentiation (Juan et al., 2000). The variety of observed phenomena in island radiations is extraordinarily diverse. In terms of colonisation alone, possibilities include (a) single colonisation of an ancestor followed by radiation, (b) multiple independent colonisations, (c) back colonisation to the mainland from island archipelagos, (d) additional colonisations and back colonisations within an archipelago (Kim et al., 2008), and (e) colonisations between archipelagos (Gillespie et al., 2008).
The Atlantic archipelagos of Madeira and the Canary Islands are excellent models for questions of island biogeo-graphy. In contrast to the Hawaiian Archipelago or the Galápagos Islands, the Canary Islands are relatively close to the mainland, providing a prolific source of colonisation events (e.g., Fuerteventura, eastern Canary Islands: 96 km to the west coast of Africa). In addition, both the Hawaiian Archipelago (oldest island 6 Ma) and the Galápagos Islands (oldest island about 4 Ma) are much younger than Porto Santo of the Madeira Archipelago (14.3 Ma) or Fuerteventura of the Canary Islands (20.2 Ma).
Patterns of colonisation of the Madeira Archipelago and the Canary Islands have been inferred for a variety of vascular plants (e.g., Böhle et al., 1996; Francisco-Ortega et al., 1996; Francisco-Ortega et al., 1997; Kim et al., 1996; Juan et al., 2000; Francisco-Ortega et al., 2002; Mort et al., 2002; Percy & Cronk, 2002) and vertebrates (Cano et al., 1984; Gonzales et al., 1996; Carranza et al., 1999; Barahona et al., 2000; Brehm et al., 2003; Maca-Meyer et al., 2003), as well as many invertebrates such as Gastropoda (Henriquez et al., 1993), Araneae (Wunderlich, 1991), Diplopoda (Enghoff, 1992), Collembola (Fjellberg, 1992), Hemiptera (Lindberg, 1953), and especially Coleoptera (Dajoz, 1977; Lindberg et al., 1958; Machado et al., 2017). Many of these taxa have undergone radiations associated with island colonisation. Vertebrate examples on the Canary Islands include lizards (Brown & Pestano, 1998; Cox et al., 2010; Thorpe et al., 1994). Invertebrate studies have addressed, e.g., weevils (Machado et al., 2017), tenebrionid beetles (Juan et al., 1996a; Rees et al., 2001), carabid beetles (Emerson et al., 1999, 2000a), grasshoppers (Hochkirch & Görzig, 2009; Hochkirch & Husemann, 2008; Husemann et al., 2014), mites (Salomone et al., 2002), spiders (Arnedo et al., 2001), cockroaches (Oromi et al., 1991), and molluscs (Abreu & Teixeira, 2008). It is striking that to date no studies have focused on the radiation of endemic wild bees in the Macaronesian islands, though they play a key role in the pollination of many of the islands’ endemic plant species (summarised in Kratochwil & Schwabe, 2018a, Kratochwil et al., 2019, Kratochwil & Schwabe, 2020). The Madeira Archipelago is characterised by 20 wild bee species with eight endemic species and one endemic subspecies (Kratochwil et al., 2018). The Canary Islands are home to 124 wild bee species, including 64 endemic species and 25 subspecies, 24 of which are endemic (Hohmann et al., 1993; Kratochwil et al., 2018; Kratochwil & Schwabe, 2018b).
Worldwide, numerous studies of the wild bee faunas of islands have been published. Oceanic volcanic islands mostly have small numbers of wild bee species, e.g., on the Galápagos Islands, only three wild bee species exist (including two probably introduced species; Gonzales et al., 2010; Rasmussen et al., 2012). An exception are islands with rapid diversification after a colonisation event, e.g., bees of the genus Hylaeus on Hawaii, which have undergone an impressive radiation in which 63 species derived from a single ancestor (Magnacca & King, 2013). Studies on oceanic islands with a focus on the radiation of taxonomically related taxa within the Anthophila using molecular genetic methods are rare (e.g., Hawaiian Hylaeus; Magnacca & Danforth, 2006, 2007), and none at all exists for the Madeira Archipelago and the Canary Islands. Exceptions are those that focus on the honey bee Apis mellifera (De la Rúa et al., 1998, 2001) and the bumble bee Bombus terrestris (Widmer et al., 1998).
The Madeira Archipelago and the Canary Islands do not have many wild bee taxa that show obvious patterns of radiation. Local assemblages are often characterised by the presence of various unrelated genera. In the case of the Canary Islands, this may be partly due to the favourable possibilities for colonisation from the mainland, which could have limited opportunities for in situ radiation of taxa. This idea is supported by the taxonomic distribution of species on the Madeira Archipelago, where many genera are represented by only a few species (Kratochwil et al., 2018). However, one genus has undergone a radiation on the archipelagos: mining bees of the genus Andrena, subgenus Micrandrena, namely, in the Andrena wollastoni group, including A. lineolata. The group has not been covered by the worldwide molecular analysis of the Andreninae by Pisanty et al. (2021).
The Andrena wollastoni group has recently been taxonomically investigated using qualitative morphological and morphometric traits (Kratochwil, 2020). All taxa of this group of small bees are endemic to the Madeira Archipelago and the Canary Islands (body length in females: 6.8–7.6 mm; in males: 5.5–6.4 mm) and are characterised by special habitat preferences (Kratochwil & Schwabe, 2020; Text S1). The group is thought to consist of two species on the Madeira Archipelago (A. wollastoni Cockerell, 1922, Madeira Island; A. dourada Kratochwil & Scheuchl, 2013, Porto Santo) and four species on the Canary Islands (A. acuta Warncke, 1968, Tenerife; A. catula Warncke, 1968, Gran Canaria; A. gomerensis, Warncke, 1993, La Gomera, La Palma; and A. lineolata Warncke, 1968, Tene-rife). Andrena gomerensis and A. acuta are further differentiated into subspecies (La Gomera: A. g. gomerensis Warncke, 1993; La Palma: A. g. palmae Kratochwil, 2020; Tenerife, Anaga region: A. a. acuta Warncke, 1968, Tenerife, Teno region: A. a. tenoensis Kratochwil, 2020, Tenerife, Dorsal Rift region: A. a. wildpreti Kratochwil, 2020).
The hypothetical ancestor of the A. wollastoni group is assumed to be related to A. tiaretta Warncke, 1974 or one of its ancestors (Text S2). Andrena tiaretta is distributed in southern Spain, Morocco, and Algeria. This hypothesis is specified in Kratochwil & Scheuchl (2013), Kratochwil (2015), and Kratochwil (2020). Andrena tiaretta morphologically shares the highest similarity with A. lineolata (Kratochwil, 2020). No other Micrandrena species occurring in Morocco today shows even approximate morphological similarity to Andrena tiaretta (Lhomme et al., 2020). The taxonomic position of the A. wollastoni group is described in the supplemental material (Text S2). To date, any phylogenetic hypotheses of the group were solely built on distribution and morphological data. However, homology of morphological characters is in many cases difficult to assess and genetic data may provide a more neutral, additional data source of information with which to infer phylogentic relationships and colonisation patterns.
We here present the first molecular study of Canarian and Madeiran wild bees. We use mitochondrial COI sequences together with qualitative morphological and morphometric data of the Andrena wollastoni group of the Madeira Archipelago and the Canary Islands to reconstruct their biogeographic history and derive colonisation and speciation scenarios. In doing so, we also test whether the results of the classification into different taxa by means of morphological and morphometric analyses published in Kratochwil (2020) can also be supported by molecular genetic markers.
Materials and methods
Study area
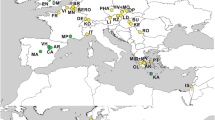
The distribution of the studied taxa covers the Madeira Archipelago and the Canary Islands with the exceptions of Lanzarote and Fuerteventura (Fig. 1).
Location of the Azores, the Archipelago of Madeira, the Selvagens Islands, the Canary Islands, and Cape Verde (a). Close view to the islands of the Madeira Archipelago (b) and the Western Canary Islands (below, right) (c). Tenerife is characterised by the regions of Anaga, Teno, Las Cañadas/Teide, and Dorsal Rift. The taxa of the A. wollastoni group (Kratochwil, 2020) and the centres of their distribution areas are indicated. Modified according to Aguiar & Karsholt (2006). The colours of the islands correspond to those used to represent species in Fig. 2. Distances from the mainland: 737 km from Porto Santo (Madeira Archipelago) to Cape Sim (Morocco); 96 km from Fuerteventura (Canary Islands) to Stafford Point (Western Sahara)
The age of the Madeiran Archipelago ranges from 5.2 to 14.3 Ma (Galopim De Carvalho & Brandão, 1991; Geldmacher et al., 2000; Schmincke, 1998). Geologically, Porto Santo is the oldest island of the Madeiran Achipelago (14.3 Ma). Much younger are Madeira Island (5.2 Ma) and Desertas (5.07 Ma). Morphospecies of the Andrena wollastoni group occur on Porto Santo and Madeira Island.
Due to the large number of islands differing in geological history and geomorphology, the Canary Islands are highly diverse. Fuerteventura and Lanzarote are the oldest islands (20.2 Ma and 15.5 Ma years; Carracedo, 2011) but are not known to have bees of the Andrena wollastoni group. Species of the A. wollastoni group occur on Gran Canaria (age 14.6 Ma), La Gomera (9.4 Ma), La Palma (1.72 Ma), and Tenerife (age between 6 to 11.5 Ma; Carracedo, 2011). The geological and geomorphological structure of Tenerife is highly complex. Features of current geological structures are mainly based on three previously separated volcanic islands with different ages (Anaga area, age 6 Ma; Teno area, 8 Ma; Roque del Conde Adeje area, 11.5 Ma), brought together by various volcanic-type processes. These, over long periods, volcanic activities have markedly affected Tenerife. The volcanic events that led to the connection of its regions Teno and Anaga began around 2 Ma and resulted in the formation of the volcanic structures of Las Cañadas and later Pico Teide (Ancochea et al., 1990). The area between Anaga and Las Cañadas, in contrast, has been filled by coastal and lacustrine sediments since the last 0.9 Ma (Dorsal Rift region = Cordillera Dorsal; Ancochea et al., 1990).
Sampling of specimens for molecular analyses
Specimens were collected from the Madeira Archipelago and the Canary Islands between 2016 and 2019 (Andrena wollastoni: A. F. Aguiar; all other taxa: A. Kratochwil and A. Schwabe). Collecting permits were issued by the governments of the Canary Islands and authorities of the Madeira Archipelago. Specimens were mostly caught on flowers with a small net or with small glass tubes, and then immediately transferred into a cool box. They were then stored individually in glass vials at 4 °C in 99% ethanol until further processing. Care was taken to ensure that at least three populations with at least four specimens per species and island were sampled at a greater distance (usually 10–20 km) from each other. Specimens that were collected a few hundred meters apart were considered members of the same population. A total of 143 specimens were collected (134 females, 9 males): Andrena acuta acuta (n = 4), A. a. tenoensis (n = 16), A. a. wildpreti (n = 12), A. catula (n = 13), A. dourada (n = 21), A. g. gomerensis (n = 24), A. g. palmae (n = 14), A. lineolata (n = 22), and A. wollastoni (n = 17). The specimens and populations per taxon are listed in Table S1.
Species and subspecies concept
The taxonomic basis of this publication is the revision of the A. wollastoni group in Kratochwil (2020). All existing types of the updated subspecies of Warncke were studied, and the types of existing species (A. lineolata, A. tiaretta) analysed. In the case of A. wollastoni, it was necessary to describe lectotypes (Kratochwil, 2018). Since no molecular genetic analysis was performed in Kratochwil (2020), which is the subject of this publication, the characterisation of the taxa (species and subspecies) corresponds to the morpho-species concept following the code of the ICZN (1999). For the molecular analyses, we analysed species and subspecies to test for molecular genetic differentiation among all taxa defined in Kratochwil (2020). We therefore treated the subspecies hypothetically as rankless morphotaxa.
Molecular analyses
DNA was extracted using a high salt protocol (Paxton et al., 1996) from adult bees, including both males and females that were all morphologically determined to species and subspecific levels (Table S1). The universal animal barcoding fragment of the cytochrome oxidase I (COI) gene was sequenced using standard protocols recommended by BOLD (http://www.barcodinglife.org) and oligonucleotide primers LCO/HCO (Folmer et al., 1994). We used the standard barcoding gene in our study as it has proven to be highly informative in previous studies, especially for bees (Schmidt et al., 2015). However, it is nevertheless important to recognise the limitations of this marker type in resolving phylogenies as problems may arise from lack of variation, hybridisation, incomplete lineage sorting, and other processes related to the maternal inheritance of mitochondrial DNA. DNA sequences were used to interrogate NCBI’s database using BLAST (https://blast.ncbi.nlm.nih.gov/Blast.cgi). All 140 sequences were devoid of stop codons, gave good query coverage (> 90%) and sequence identity (> 90%) to other Micrandrena species (e.g., Andrena enslinella Stoeckhert, 1924), though sequence identity was < 97% for all specimens, suggesting that databases currently lack reference sequences for the focal taxa.
Sequences were cleaned and trimmed using Geneious v. 6.1.2. (Kearse et al., 2012) and were aligned using the MUSCLE algorithm. In addition to our 140 sequences of the Andrena wollastoni group, we added eight reference sequences, two each from Andrena enslinella Stoeckhert, 1924; A. minutuloides Perkins, 1914; A. semilaevis Pérez, 1903; and A. subopaca Nylander, 1848, retrieved from the NCBI (GenBank) database as outgroups. The final alignment was unambiguous, devoid of INDELS, and, after trimming, the final dataset comprised 148 sequences of 533 bp length.
Phylogenetic analyses
We estimated the most suitable substitution model using MrModeltest v. 2.3 (Nylander, 2004). The most suitable model was determined as GTR + G based on the AIC. In order to adjust for rate heterogeneity, we used the reversible jump model with Gamma rate parameter in MrBayes v. 3.2.6 (Ronquist & Huelsenbeck, 2003). We ran the simulation for 10 million generations, sampling every 1000 generations for a total of 10,000 final sampled trees. Convergence was confirmed with average split frequencies being below 0.01. Trees were visualised with FigTree v. 1.4 (Rambaut, 2009).
Further, we ran *BEAST in BEAST v. 1.8.2. (Drummond et al., 2012) with a strict molecular clock to generate a species tree with a rough estimate for the timing of divergence. As we deal with very young divergences, we consider the strict clock a valid approach (Hein et al., 2005). We first generated the input file for BEAST using BEAUti v. 1.8.2. For *BEAST we grouped species and subspecies as units. As no bee specific rate is available, we used the rate proposed by Machado et al. (2017) for Macaronesian flightless weevils (genus Laparocerus; 225 taxa studied; confirmation of monophyly of 36 species and subspecies of the Madeira Archipelago, and 196 species and subspecies of the Canary Islands). The rate proposed by Machado et al. (2017) is intermediate between those proposed by Brower (1994) and Papadopoulou et al. (2010) and hence represents a conservative estimate. The Yule process was used as a tree prior with a piecewise linear and constant root as population size model as no coalescent models are available with *BEAST; a random tree was used as starting tree. The ploidy type was chosen as mitochondrial. We ran the analysis for 300 million generations, sampling every 30,000 iterations. Effective sampling size (ESS) values were above 100 for all parameters and above 200 for most parameters, suggesting the analyses converged. Trees were summarised with TreeAnnotator v. 1.8.2 and visualised with FigTree. We further calculated a Median Joining network using Popart (Leigh & Bryant, 2015).
Morphometric analyses
For the morphometric analyses, 279 females were available: Andrena acuta acuta (n = 15), A. a. tenoensis (n = 22), A. a. wildpreti (n = 13), A. catula (n = 50), A. dourada (n = 46), A. g. gomerensis (n = 40), A. g. palmae (n = 31), A. lineolata (n = 13), and A. wollastoni (n = 49). The analysed specimens are deposited in the following collections (with acronyms of the depository): A. Kratochwil (KR; n = 204); Upper Austrian State Museum Linz, Austria (OLML: n = 37); Überseemuseum Bremen, Germany (UMBB; n = 35); and H. R Schwenninger, Stuttgart, Germany (HS: n = 3). The specimens are individually characterised by an identity code (ID-No) with further information (locality, altitude above sea level, latitude and longitude coordinates, date of collection, collector and further remarks) (see Table S2). Only females were considered since a preliminary study revealed that morphometric analysis provided clearer differences between taxa in females (AK, unpubl. data).
Eighteen parameters were investigated (in alphabetical order; in brackets, abbreviation, and magnification used for measurement): body length (BL, 16.25); from antennal base to tip of the pygidium, clypeus length (CL, 100 ×), facial fovea maximal length (FVL, 100 ×), facial fovea maximal width (FVW, 100 ×), flagellomere length 1–3 (each flagellomere separately, FL1, FL2, FL3, 100 × ; measured on ventral surfaces of the flagellomeres with the antenna stretched forward), head length (HL, 40 × ; from the top of the vertex to the lower margin of the clypeus excluding the process of the labrum), head width (HW, 40 ×), labrum process width (apical process width) at the top (LPW, 100 ×), mesosomal width (MSW, 40 × ; between outer rims of the tegulae), metasomal width (MTW, 40 ×), maximum width of terga from dorsal view, ocelloccipital distance (OCD, 62.5 ×), ocellocular distance (OOD, 62.5 ×), postocular distance (POD, 62.5 ×), propodeum basal area length (PBAL, 100×), pterostigma length (PSL, 100×), wing length (WL, 16.25 ×), and length of the forewing including the tegula. The morphological and morphometric studies were carried out with a stereo microscope (Wild M3Z modular stereomicroscope, Heerbrugg, Switzerland, with a 25 × eyepiece with micrometer giving 16.25 × , 40 × , 62.5 × , and 100 × magnification).
Kratochwil (2020) used statistical methods for a morphometric analysis of the taxa of the A. wollastoni group according to Baur and Leuenberger (2011). These methods include univariate analysis, calculation of Pearson product-moment correlation coefficients, multivariate ratio analysis, principal component analysis, linear discriminant analysis, and allometry ratio spectrum. Of particular importance is the analysis of whether or not individual morphometric measures depend on individual body size. In Kratochwil (2020), further tools of correlation and eigenvalue analyses were developed. The morphometric results are presented in Kratochwil (2020) in 50 boxplots. In total, 225 significance tests were performed. Furthermore, the shape PC1/shape PC2 clusters and the best plotted ratio of morphometric features were presented in 60 diagrams. We summarise these results in one diagram using the following method: Pearson product-moment correlation coefficients were calculated for all measurements with the function ‘cor ()’ in R (R Core Team, 2016) using default settings. Ten of our parameters with the highest correlation coefficients were used for further analysis (BL, FL2, FL3, HL, HW, MSW, MTW, OOD, PSL, WL). The morphometric data of these parameters are presented in Table S3. For further analysis, the data were log10-transformed. A linear discriminant analysis (LDA) in R was then used to detect linear combinations of the original variables (10 morphometric parameters) that provided the best possible separation between the groups (taxa) in our dataset, including highest eigenvalues. Linear discriminant analysis was carried out using the ‘lda()’ function of the R package MASS (Venables & Ripley, 2002). LDA determines group means and computes, for each specimen, the probability of belonging to different groups. A specimen is then assigned to the group with the highest probability score. A plot of LDA scores was generated with a confidence ellipse around individuals of each taxon corresponding to the 95% confidence level, and further adjusted in EazyDraw (2020).
Qualitative morphological analyses
A total of 40 qualitative (non-meristic) parameters (colour, structure, pubescence) were measured in the same individuals (Table S4). An explanation of the parameter groups is given in Table S5, whilst the data matrix is shown in Table S6. All parameters are explained in detail in Kratochwil (2020). Principal component analysis was used to explore non-meristic character variation across individuals using the function ‘prcomp’ from R’s stats package (R Core Team, 2016). The various assumptions of the PCA approach were satisfied by the data. The results were visualised in the R package ggbiplot (Vu, 2011) and adjusted in EazyDraw (2020).
Combined analyses
In addition, we correlated the genetic and the qualitative morphological and morphometric results to understand their relationships, following the synthetic approach of Baur et al. (2014). We generated estimates of genetic differentiation between species as ΦST with DnaSP v. 6.12.03 (Rozas et al., 2017); subspecies were grouped together within species as phylogenetic analyses already suggested no genetic differences between them. Differentiation in qualitative morphological and morphometric traits were calculated as square Euclidian and Mahalanobis distances, respectively, with PAST v. 4.03 (Hammer-Muntz et al., 2001). The different distance matrices were correlated with PAST.
Results
Molecular analyses
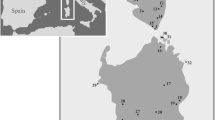
We generated a total of 140 ingroup sequences and included eight outgroup sequences. The total alignment was 533 bp in length. The haplotype network showed little overlap between the island populations (Fig. 2). Only La Gomera and La Palma shared similar haplotypes on the Canary Islands; Madeira and Porto Santo also shared haplotypes. Individuals from La Gomera and La Palma were very distinct from all others (22 mutations). Further, Andrena lineolata from Tenerife was quite divergent from all other populations (11 mutations to the closest other haplotype from A. dourada and A. wollastoni, a maximum of 33 mutations to A. gomerensis). Andrena dourada (Porto Santo) took a central place in the haplotype network and is connected to most other species. Andrena dourada and A. wollastoni (Madeira Island) have private haplotypes, but also share two haplotypes. In turn, A. catula (Gran Canary) and A. acuta (Tene-rife) only differ by one mutation, but do not share any haplotypes. Andrena acuta is likely derived from an A. catula haplotype. Subspecies differentiation, as is found within A. acuta and within A. gomerensis using morphometric traits (Kratochwil, 2020), is not reflected in the haplotype analysis.
Median Joining network of Andrena species (Micrandrena) of the Canary Islands and the Madeira Archipelago. Circle size is relative to number of haplotype copies present in dataset. A branch re-presents a single nucleotide change (mutation); bars on branches re-present inferred missing haplotypes (single nucleotide changes). LG La Gomera, LP La Palma. The colours correspond to those used to represent the location of species on islands in Fig. 1
The phylogeny generated with MrBayes did not resolve all species (Fig. S1). As in the haplotype network, subspecies of Andrena gomerensis from La Gomera and La Palma could not be distinguished. The populations of both islands took the basal position in the ingroup and are the sister group to all other populations (PP: 1.00). Next, Andrena lineolata from Tenerife split off (PP: 1.00). The other populations were not well resolved and represent the sister group to A. lineolata. Andrena acuta (Tenerife) and A. catula (Gran Canary) grouped together with high support (PP: 0.99) but did not form monophyletic clusters. Andrena dourada (Porto Santo) and A. wollastoni (Madeira Island) were admixed.
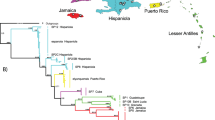
The analysis with *BEAST provided overall similar results in terms of the relationships (Fig. 3). Again, Andrena gomerensis from La Palma and La Gomera grouped together with high support (PP: 1.0) and represented the basal lineage within the ingroup. Next, Andrena lineolata from Tenerife split off (PP: 1.0) and is the sister group to all other species. Andrena catula (Gran Canary) is the sister of A. acuta (Tenerife; PP: 0.94); the group of A. catula and A. acuta is the sister group to the species from the Madeira Archipelago (PP: 1.00). Andrena dourada is sister of A. wollastoni (PP: 0.97). The molecular clock analysis dates the basal split within the ingroup at about 2.18 Ma (1.28–3.11; 95% confidence intervals), when A. gomerensis split off from all others. Andrena lineolata split off the other populations at about 0.89 (0.41–1.41) Ma, whereas the species of the Madeira Archipelago and A. catula and A. acuta diverged about 0.27 (0.11–0.5) Ma. Andrena catula and A. acuta split about 0.14 (0.04–0.28) Ma. The species from the Madeira Archipelago diverged within the last 60 (10–160) Ka. Andrena wollastoni is the likely most recent species.
Dated species tree demonstrating the phylogenetic relationships of the different island populations calculated with *BEAST compared to the outgroup species Andrena enslinella, A. subopaca, A. minutuloides, and A. semilaevis (all Micrandrena); dating is based on a strict molecular clock with the rate proposed by Machado et al. (2017). Branch support above 0.95 is shown with red circles; posterior probabilities above 0.90 are shown with yellow circles. Times are given in million years. Blue bars represent 95% confidence intervals
Morphometric analyses
The LDA analysis, which includes ten different morphometric parameters (BL, FL2, FL3, HL, HW, MSW, MTW, OOD, PSL, WL), differentiates to a greater or lesser extent among the studied species (Fig. 4). Among the characters, HW and WL have the strongest influence on clustering, BL, FL2, and FL3 the lowest. The parameters HL, MSW, MTW, and OOD have an intermediate position.
Andrena gomerensis is morphometrically well differentiated from A. lineolata (Fig. 4). Andrena wollastoni, the likely most recently differentiated species, is morphometrically in the LDA analysis distinguishable from all others (Fig. 4). Smaller differences exist between A. gomerensis and A. catula, as well as between A. dourada and A. acuta (Fig. 4). Andrena lineolata represents a well-defined morphometric cluster compared to all other species, with very limited overlap with A. catula and A. dourada (Fig. 4). Andrena dourada shows an almost complete separation from A. wollastoni. Andrena dourada and A. catula, as well as A. dourada and A. acuta show no clear differentiation (Fig. 4).
Qualitative morphological analyses
The PCA analysis of 40 morphological parameters (species-specific colours, structures, pubescence of different body parts) documents that the species are well differentiated and is in accord with the molecular results. The PCA shows a central position for Andrena gomerensis (Fig. 5) and a peripheral position for A. lineolata. The decisive morphological factors are shown in the vector diagram (Fig. 5, numbers 1–10). Andrena catula and A. dourada show high morphological similarity and are clearly differentiated from A. acuta and A. wollastoni, which themselves exhibit higher morphological similarity.
Principal component analysis of the morphological data. The colours of the specimen labels correspond to those used in Figs. 1, 2 (explained variance PC1: 58.8%, PC2: 19.3%) The black arrows plotted in the centre (biplot function) characterise for each parameter the direction of increasing values. The length of the arrow indicates how accurately the parameter explains the distribution of the data
Comparative analyses
Genetic differentiation (ΦST) was generally high and varied between 1.0 between Andrena gomerensis and A. lineolata and 0.42 between A. wollastoni and A. dourada. Euclidian distances for morphometric traits varied between 9.33 between A. lineolata and A. acuta and 3.46 between A. acuta and A. gomerensis. Squared Mahalanobis distances for morphometric traits differed between 82.31 between A. gomerensis and A. wollastoni and 6.7 between A. catula and A. acuta. Both qualitative morphological and morphometric data showed limited correlation with genetic distance (r2 = 0.12 and r2 = 0.18, respectively; Fig. 6).
Discussion
We here provide the first molecular and taxonomic comparison of a group of bee species of the Madeira Archipelago and the Canary Islands using molecular, qualitative morphological, and morphometric data. Based on molecular clock estimates, the group seems to be young, with a common origin within the last two Ma; based on the (limited) genetic data, most, but not all, species are monophyletic, yet all are morphologically well defined. Our data suggest a complex colonisation scenario deviating from classic island hopping. We show that while the different types of data largely concur, the combination of all data provides the most complete picture of the group.
Colonisation patterns of Macaronesian Islands
For most animal groups studied in Macaronesia, taxa have generally only been found to have colonised a single archipelago, e.g., lizards (Cox et al., 2010), beetles (Emerson et al., 2000b, 2000c; Contreras-Díaz et al., 2007; Mas-Peinado et al., 2018; Rangel Lopez et al., 2018) and grasshoppers (Hochkirch & Görzig, 2009). The wild bees of the Andrena wollastoni group, on the other hand, occur with their representatives on two archipelagos (Madeira Archipelago and Canary Islands). Yet, separate colonisation of the two archipelagos from the mainland seems unlikely for the species of the Andrena wollastoni group due to the molecular results; the populations of the Madeira Archipelago are nested within the Canary Island populations.
It is remarkable that the molecular results presented here provide evidence of colonisation between the two archipelagos for the tiny bees of the Andrena wollastoni species group. Indeed, our results suggest that the sister group of all other Macaronesian species is found on La Gomera or La Palma (Canary Islands). Already Warncke (1968) considered all taxa of the Andrena wollastoni group, except A. lineolata, to be subspecies of A. wollastoni (Kratochwil, 2020); he hypothesised, but without any supporting data, (a) either the former existence of A. wollastoni on the African continent with later colonisation of the Madeira Archipelago and the Canary Islands (followed by diversification at the subspecies level) or (b) the speciation of A. wollastoni after colonisation, followed by secondary colonisation between both archipelagos, similar to our results presented here.
Colonisation models
Many colonisations within the Canary Islands follow a simple stepping-stone model (Emerson, 2002) from east to west according to the prevailing ocean currents and wind direction. This is documented for grasshoppers (Hochkirch & Görzig, 2009; López et al., 2007) and for Canarian beetles (Emerson et al., 2000c; Juan et al., 1995; Machado et al., 2017; Rees et al., 2001). In our case, successful colonisation of one island (La Gomera) seems to have been followed by subsequent radiation on the complete archipelago (Fig. 7). In Chalcides skinks (Brown & Pestano, 1998), La Gomera is also the centre of origin from which Gran Canaria and Tenerife were directly colonised.
Such colonisation patterns may have various causes. The islands of a volcanic archipelago did not all arise at the same point in time and have not exhibited equal patterns of volcanic activity. Therefore, the possible time windows for colonisations not only depend on the age of the islands but also on the habitat conditions for the coloniser (Text S1) and local extinction promoted by volcanism.
Andrena gomerensis (La Gomera, La Palma): Following our molecular clock analyses, the basal split of Andrena gomerensis (La Gomera, La Palma) is the basis for the development of all other species (2.18 Ma; Pleistocene, Gelasian) (Fig. 7). This is currently the oldest evidence of a wild bee species in the Canary Islands. It is unlikely that the ancestor of A. gomerensis colonised La Gomera directly from the mainland NW Africa. Fuerteventura and/or Lanzarote, as well as Gran Canaria, may have served as stepping-stone islands. Fuerteventura and Lanzarote are currently not inhabited by bee species of the A. wollastoni group, which may be explained by changes in the climate from more humid conditions at the time of the possible colonisation (Alonso-Zarza & Silva, 2002) to the currently extremely hot infracanarian zone (with the exception of very small subhumid summit areas of the mountains); the infracanarian zone is not a suitable habitat for this bee group (Text S1 and Kratochwil & Schwabe, 2020).
Lanzarote and Fuerteventura are also characterised by marked, recent volcanism that devastated large parts of the islands (Pallarés Padilla, 2007) and Fuerteventura had frequent periods of volcanic activity, with the most recent ones dated from about 6000 years ago (Carracedo, 2011). A reason for the apparent lack of early successful colonisation of the Andrena wollastoni group on Gran Canaria may be the island’s very long and violent volcanic activity. The marked volcanic activity in the area of the Roque Nublo complex on Gran Canaria destroyed large parts of the island 3–3.5 Ma ago (Pérez-Torrado & Mangas, 1994; Marrero, 2004). During the Quaternary, volcanic activity, which continued through to 3000 ybp, was mainly restricted to the northeast of the island (Funck et al., 1996). Extinctions or reduction in species numbers or range reductions of species as a probable consequence of volcanism have been suggested for insects, reptiles and also plants (Juan et al., 1995, Marrero & Francisco-Ortega, 2001, Emerson, 2003, Marrero, 2004, Machado et al., 2017).
The environmental conditions were completely different on La Gomera, where before 4–2 Ma, there is evidence of only insignificant, very local, and sporadic volcanic activity. In the last two million years, volcanic activity has been non-existent (Ancochea et al., 2006). The prevailing habitat conditions were likely favourable to the settlement and long-term establishment of Andrena gomerensis.
The populations of both islands (La Gomera, age 9.4 Ma; La Palma, age 1.72 Ma) could be separated at the subspecies level by morphometric methods (A. g. gomerensis, A. g. palmae; Kratochwil, 2020), but not with mDNA molecular methods. Due to the young age of La Palma, the colonisation can be hypothesised as a more recent one, as supported by the genetic data. The distance between both islands is only 58 km.
We hypothesise that A. gomerensis colonised Tenerife from La Gomera (distance about 30 km) (Fig. 7). On Tene-rife, the former palaeo-islands Anaga and Teno merged around 2 Ma ago by volcanism, forming Las Cañadas and later El Teide of Tenerife. The area between the regions Anaga and Las Cañadas had been filled by lacustrine and coastal material over the past 0.9 Ma (Ancochea et al., 1990). The young geomorphological structures of Las Cañadas were formed less than 200,000 years ago. According to our results, Andrena lineolata (Tenerife) split off at about 0.89 Ma (Pleistocene, Calabrian). We hypothesise that A. lineolata colonised primarily the lower zones and shifted later to the mountain areas. Today A. lineolata is limited to the Las Cañadas/Teide area, 2000 up to around 3000 m a.s.l., an extreme environment in the subalpine shrub zone with temperature extremes and stormy conditions.
Andrena catula (Gran Canaria) probably evolved from A. gomerensis or its ancestor, more likely through dispersal from La Gomera (direct line 125 km) than from La Palma (Fig. 7). Long-term establishment of a species of the A. wollastoni group on Gran Canaria from the mainland was probably not possible due to the long-lasting volcanic activity on that island (see above).
Andrena dourada (Porto Santo): The long colonisation distance of Porto Santo from La Palma (approximately 480 km) or La Gomera (approximately 540 km) seems astonishing (Fig. 7). Yet the most remote oceanic island archipelago Hawaii, 3200 km from the nearest land mass or island archipelago, has been colonised naturally by small insects, including those of the genus Hylaeus (Anthophila, Colletidae; Magnacca & Danforth, 2006), which are of similar body size to the species of the A. wollastoni-group studied here. Dispersal from La Gomera to Porto Santo is therefore plausible for members of the A. wollastoni group.
Violent sandstorms from the Sahara region can reach the Canary Islands, where they can cause great damage. Satellite images show that such winds can also reach the Madeira Archipelago via the Canary Islands. Impressive satellite images of such ‘Calima’ events exist (e.g., https://de.wikipedia.org/wiki/Calima#/media/Datei:Sahara_Canary_Islands.jpg). Such events could easily lead to aerial drifting of small animal species between the two archipelagos (e.g., Odonata; Weihrauch, 2011). It might be assumed that the decisive factor for successful long-distance colonisation is not so much the colonisation event per se as the ability to establish a new population once having reached a distant island, which depends on preexisting habitats for nesting and the availability of food at the recipient island.
Volcanic activity ended on Porto Santo over 8 million years ago (Cameron et al., 2006). The proposed split of Andrena dourada (Porto Santo) (0.27 Ma) falls at the Middle Pleistocene (Chibanian). Porto Santo is today an example of an old, already strongly eroded island with a mainly Mediterranean xeric oceanic climate; only small parts are exposed to subhumidic influence (Kratochwil & Schwabe, 2018b). We hypothesise that, in the time window of and after the split, the habitat conditions on Porto Santo were much more favourable for A. dourada or its ancestor than today. The island was less eroded, had more humid-subhumid parts and had larger and higher mountain areas, including sites with Laurisilva vegetation (Press & Short, 1994; Silva, 2003). We propose that in the Upper/Late Pleistocene, A. dourada or its ancestor colonised Madeira Island (Fig. 7).
The region of Madeira Island closest to Porto Santo (about 40 km) is a small peninsula (Ponta de São Lourenço). This was volcanically active in two time-windows: > 5.2–4.0 Ma and 2.4–0.9 Ma (Klügel et al., 2009).
Andrena wollastoni (Madeira Island) seemingly arose from Andrena dourada (Porto Santo) or its ancestor. Andrena wollastoni is a supergeneralist, nowadays found in large numbers on Madeira Island.
The current distance between Gran Canaria and Tene-rife is 62 km. Colonisation of Tenerife probably took place by individuals of Andrena catula from Gran Canaria about 140,000 years ago (Fig. 7), at a time when volcanic activity of the Canary Islands was much lower than earlier, at the end of the Pleistocene. On lower altitudes of Tenerife, a second endemic A. wollastoni-group species, Andrena acuta, is nowadays found in regions of the thermo-infracanarian and thermo-mesocanarian zones.
As the colonisation by A. acuta possibly occurred very recently, it would have been at a time when the former palaeo-islands were already connected. Machado (1976) and Machado et al. (2017) assumed that ‘inland islands’ covered with vegetation and surrounded by lava flows (‘kipuka scenario’) played a role in the separation of populations which were able to develop in isolation. ‘Kipukas’ means remnants of the original habitats in the area surrounded by lava flows (Vandergast & Gillespie, 2004).
Comparison between molecular, morphometric, und qualitative morphological results
The molecular, morphometric, and qualitative morphological analyses show mostly consistent results, but are only moderately correlated and differences are also apparent. All species can be separated as monophyletic group arising from Andrena gomerensis or its ancestor using all three methods. Although this is a very young radiation, the younger species can also be differentiated from each other with the exception of Andrena dourada and A. wollastoni.
As already documented in Kratochwil (2020), a taxon-taxon comparison using PCA and LDA shows that morphometry clearly separates all taxa. This method provides the highest resolution. In Kratochwil (2020) the morphometric results are presented in 50 boxplots comparing all taxa including 225 significance tests. In addition, a pairwise taxa comparison was performed and the shape PC1/shape PC2 clusters were visualised in 30 diagrams together with diagrams of the most differentiating ratios of morphometric features. Figure 4 sums this data up in a single diagram. However, it does not provide a morphometric comparison of all taxa in one diagram due to the compression of axes. The LDA, with all taxa considered, documents this through the loss of resolution. Nevertheless, the differences are clear at the 95% level and the species can be morphometrically related to each other, corresponding to the molecular analysis.
The large morphological divergence between A. gomerensis and A. lineolata can be explained by a longtime separation of the two taxa, as also supported by the sequence divergence between them. On the other hand, the morphometric analysis differentiated the two closely related species A. wollastoni and A. dourada much better than the molecular genetic data. This may suggest that morphology is under high selection, driving divergence. The weak correlation of morphological and morphometric data may also point towards selection on morphological traits (e.g., Saether et al., 2007). It may, however, also simply point to a high resolution of the morphological data or, conversely, the low resolution of COI gene sequences. However, generally caution is required when analysing qualitative morphological parameters as convergent feature complexes may also occur. In combination with the molecular results, the close morphological relationship of A. wollastoni and A. acuta (both are the youngest taxa) suggests that some morphological similarities (especially the presence of darker facial hairs, and equally darker scopa hairs) may be the result of convergence, while that of A. catula and A. dourada is more likely to be due to common ancestry.
The molecular relationships between island taxa using mitochondrial sequences do not always correspond to morphological classifications. According to Juan et al. (1996b), a discrepancy between mtDNA and morphological differentiation may result from higher morphological variability of island taxa compared to their related mainland species. One reason might be a higher evolutionary rate of morphological traits compared to molecular traits, which might be a reaction to increased selection under new environmental conditions. On the other hand, the effects of drift may be high in small founder populations and hence lead to faster shifts in traits.
Data availability
All sequence data generated during the current study are available in the NCBI Genbank under the accessions MZ868018-MZ868157; all morphological and morphometric data generated in this study is available in the supplemental information of this article.
References
Abreu, C., & Teixeira, D. (2008). The molluscs (Mollusca) of the Madeira and Selvagens archipelagos. In Borges P. A. V., Abreu C., Aguiar A. M. F., Carvalho P., Jardim R., Melo I., Oliveira P., Sérgio C., Serrano A. R. M., & P. Vieira (Eds.) A list of the terrestrial fungi, flora and fauna of Madeira and Selvagens archipelagos (pp. 227–244). Funchal and Angra do Heroísmo: Direcção Regional do Ambiente da Madeira and Universidade dos Açores.
Aguiar, A. M. F., & Karsholt, O. (2006). Lepidoptera: Systematic catalogue of the entomofauna of the Madeira Archipelago and Selvagens Islands. Boletim do Museu Municipal do Funchal (Historia Natural), 9, 5–140.
Alonso-Zarza, A. M., & Silva, P.-G. (2002). Quaternary laminar calcretes with bee nests: Evidences of smallscale climatic fluctuations, Eastern Canary Islands, Spain. Palaeogeography, Palaeoclimatology, Palaeoecology, 178, 119–135.
Ancochea, E., Füster, J., Ibarrola, E., Cendrero, A., Coello, J., Hernan, F., Cantagrel, J. M., & Jamond, C. (1990). Volcanic evolution of the island of Tenerife (Canary Islands) in the light of new K-Ar data. Journal of Volcanology and Geothermology Research, 44, 231–249.
Ancochea, E., Hernán, F., Huertas, M. J., Brändle, J. L., & Herrera, R. (2006). A new chronostratigraphical and evolutionary model for La Gomera: Implications for the overall evolution of the Canarian Archipelago. Journal of Volcanology and Geothermology Research, 157, 271–293.
Arnedo, M. A., Oromi, P., & Ribera, C. (2001). Radiation of the spider genus Dysdera (Araneae, Dysderidae) in the Canary Islands: cladistic assessment based on multiple data sets. Cladistics, 17, 313–353.
Barahona, F., Evans, S., Mateo, J. A., García-Márquez, M., & López-Jurado, L. F. (2000). Endemism, gigantism and extinction in island lizards: The genus Gallotia on the Canary Islands. Journal of Zoology London, 250, 373–388.
Baur, H., & Leuenberger, C. (2011). Analysis of ratios in multivariate morphometry. Systematic Biology, 60, 813–825.
Baur, H., Kranz-Baltensperger, Y., Cruaud, A., Rasplius, J.-Y., Timokhov, A. V., & Gokhan, V. E. (2014). Morphometric analysis and taxonomic revision of Anisopteromalus Ruschka (Hymenoptera: Chalcidoidea: Pteromalidae) — An integrative approach. Systematic Entomology, 39, 691–709.
Böhle, U.-R., Hilger, H. H., & Martin, W. F. (1996). Island colonization and evolution of the insular woody habit in Echium L. (Boraginaceae). Proceedings of the National Academy of Science USA, 93, 11740–11745.
Brehm, A., Jesus, J., Spinola, H., Alves, C., Vicente, L., & Harris, D. J. (2003). Phylogeography of the Madeiran endemic lizard Lacerta dugesii inferred from mitochondrial DNA sequences. Molecular Phylogenetics and Evolution, 26, 222–230.
Brower, A. V. Z. (1994). Rapid morphological radiation and convergence among races of the butterfly Heliconius erato inferred from patterns of mitochondrial DNA Evolution. Proceedings of the National Academy of Science USA, 91, 6491–6495.
Brown, R. P., & Pestano, J. (1998). Phylogeography of skinks (Chalcides) in the Canary Islands inferred from mitochondrial DNA sequences. Molecular Ecology, 7, 1183–1191.
Cameron, R. A. D., Cook, L. M., Goodfriend, G. A., & Seddon, M. B. (2006). Fossil land snail faunas of Porto Santo, Madeiran Archipelago: Change and stasis in Pleistocne to recent times. Malacologica, 49(1), 25–59.
Cano, J., Baez, M., López-Jurado, L. F., & Ortega, G. (1984). Karyotype and chromosome structure in the lizard Gallotia galloti in the Canary Islands. Journal of Herpetology, 18, 344–349.
Carracedo, J. C. (2011). Geología de Canarias I. (Origen, evolución, edad y volcanismo). Editorial Rueda, Madrid.
Carranza, S., Arnold, E. N., Thomas, R. H., Mateo, J. A., & López-Jurado, L. F. (1999). Status of the extinct giant lacertid lizard Gallotia simonyi simonyi (Reptilia: Lacertidae) assessed using mtDNA sequences from museum specimens. Journal of Herpetology, 9, 83–86.
Contreras-Díaz, H. G., Moya, O., Oromí, P., & Juan, J. (2007). Evolution and diversification of the forest and hypogean ground-beetle genus Trechus in the Canary Islands. Molecular Phylogenetics and Evolution, 47, 687–699.
Cox, S. C., Carranza, S., & Brown, R. P. (2010). Divergence times and colonization of the Canary Islands by Gallotia lizards. Molecular Phylogenetics and Evolution, 56, 747–757.
Dajoz, R. (1977). Faune de l’Europe et du bassin méditerranéen, 8. Coléoptères Colydiidae et Anommatidae Paléarctiques, Masson, Paris.
De la Rúa, P., Serrano, J., & Galián, J. (1998). Mitochondrial DNA variability in the Canary Islands honeybees (Apis mellifera L.). Molecular Ecology, 7, 1543–1547.
De la Rúa, P., Galián, J., Serrano, J., & Moritz, R. F. A. (2001). Genetic structure and distinctness of Apis mellifera L. populations from the Canary Islands. Molecular Ecology, 10, 1733–1742.
Drummond, A. J., Suchard, M. A., Xie, D., & Rambaut, A. (2012). Bayesian phylogenetics with BEAUti and the BEAST 1.7. Molecular Biology and Evolution, 29, 1969–1973.
EazyDraw. (2020). Vector drawing software. Vers. 9.8.1. Dekorra Optics LLC, Wisconsin, USA.
Emerson, B. C. (2002). Evolution on oceanic islands: Molecular phylogenetic approaches to understanding pattern and process. Molecular Ecology, 11, 951–966.
Emerson, B. C. (2003). Genes, geology and biodiversity: Faunal and floral diversity on the island of Gran Canaria. Animal Biodiversity and Conservation, 26, 9–20.
Emerson, B. C., Oromí, P., & Hewitt, G. M. (1999). MtDNA phylogeography and recent intra-island diversification among Canary Island Calathus beetles. Molecular Phylogenetics and Evolution, 13, 149–158.
Emerson, B. C., Oromí, P., & Hewitt, G. M. (2000a). Interpreting colonization of the Calathus (Coleoptera: Carabidae) on the Canary Islands and Madeira through the application of the parametric bootstrap. Evolution, 54, 2081–2090.
Emerson, B. C., Oromí, P., & Hewitt, G. M. (2000b). Tracking colonization and diversification of insect lineages on islands: Mitochondrial DNA phylogeography of Tarphius canariensis (Coleoptera: Colydiidae) on the Canary Islands. Proceedings of the Royal Society of London B, Biological Science, 267, 2199–2205.
Emerson, B. C., Oromí, P., & Hewitt, G. M. (2000c). Colonization and diversification of the species Brachyderes rugatus (Coleoptera) on the Canary Islands: Evidence from mitochondrial DNA COII gene sequences. Evolution, 54, 911–923.
Enghoff, H. (1992). Macaronesian millipedes (Diplopoda) with emphasis on endemic species swarms on Madeira and the Canary Islands. Biological Journal of the Linnean Society, 46, 153–161.
Fjellberg, A. (1992). Revision of European and North African Folsomides Stach with special emphasis on the Canarian fauna (Collembola: Isotomidae). Insect Systematics and Evolution, 23, 453–473.
Folmer, O., Black, M., Hoeh, W., Lutz, R., & Vrigenhoek, R. (1994). DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Molecular Marine Biology and Biotechnology, 3, 294–299.
Francisco-Ortega, J., Jansen, R., & Santos-Guerra, A. (1996). Chloroplast DNA evidence of colonization, adaptive radiation, and hybridization in the evolution of the Macaronesian flora. Proceedings of the National Academy of Science USA, 93, 4085–4090.
Francisco-Ortega, J., Crawford, D. J., Santos-Guerra, A., & Jansen, R. K. (1997). Origin and evolution of Argyranthemum (Asteraceae: Anthemideae). In Givnish T. J. & K. J. Sytsma (Eds.) Molecular evolution and adaptive radiation (pp. 407–432). New York, Cambridge University Press.
Francisco-Ortega, J., Fuertes-Aguilar, J., Kim, S. C., Santos-Guerra, A., Crawford, D. J., & Jansen, R. K. (2002). Phylogeny of the Macaronesian endemic Crambe section Dendrocrambe (Brassicaceae) based on internal transcribed spacer sequences of nuclear ribosomal DNA. American Journal of Botany, 89, 1984–1990.
Funck, T., Dickmann, T., Rihm, R., Krastel, S., Lykke-Anderson, H., & Schmincke, H.-U. (1996). Reflection seismic investigations in the volcaniclastic apron of Gran Canaria and implications for its volcanic evolution. Geophysical Journal International, 125(2), 518–536.
Galopim de Carvalho, A. M., & Brandão, J. M. (1991). Geologia do Arquipélago da Madeira. Museo Nacional History in National University Lisboa 170 pp.
Geldmacher, J., van der Bogaard, P., & Schmincke, H.-U. (2000). The 40Ar/36Ar age dating of the Madeira Archipelago and hotspot track (eastern North Atlantic). Geochemistry Geophysics Geosystem, 1: Paper number 1999GC000018.
Gillespie, R. G., Claridge, E. M., & Goodacre, S. L. (2008). Biogeography of the fauna of French Polynesia: diversification within and between a series of hot spot archipelagos. Phil. Trans. Roy. Soc. London. Series B, Biological Science, 363(1508), 3335–3346.
Gonzalez, P., Pinto, F., Nogales, M., Jiménez, A., Hernández, A. M., & Cabrera, V. M. (1996). Phylogenetic relationships of the Canary Island endemic lizard genus Gallotia (Sauria: Lacertidae), inferred from mitochondrial DNA sequences. Molecular Phylogenetics and Evolution, 6, 63–71.
Gonzales, V. H., Koch, J. B., & Griswold, T. (2010). Anthidium vigintiduopunctatum Friese (Hymenoptera: Megachilidae): The elusive ‘dwarf bee’ of the Galápagos Archipelago? Biological Invasions, 12, 2381–2383.
Hein, J., Schierup, M. H., & Wiuf, C. (2005). Gene Genealogies, Variation and Evolution: A Primer in Coalescent Theory. Oxford University Press.
Hammer-Muntz, O., Harper, D., & Ryan, P. D. (2001). PAST: paleontological statistics software package for education and data analysis version 2.09.
Henriquez, F. C., Henriquez, F. C., Ibanez, M., & Alonso, M. R. (1993). Revision of the genus Napaeus Albers, 1850 (Gastropoda Pulmonata: Enidae). The problem of Napaeus (Napaeinus) nanodes (Shuttleworth, 1852) and description of five new species from its conchological group. Journal of Molluscan Studies, 59, 147–163.
Hochkirch, A., & Husemann, M. (2008). A review of the Canarian Sphingonotini with description of a new species from Fuerteventura (Orthoptera: Acrididae, Oedipodinae). Zoological Studies, 47, 495–506.
Hochkirch, A., & Görzig, Y. (2009). Colonization and speciation on volcanic islands: Phylogeography of the flightless grasshopper genus Arminda (Orthoptera, Acrididae) on the Canary Islands. Systematic Entomology, 34, 188–197.
Hohmann, H., La Roche, F., Ortega, G., & Barquin, J. (1993). Bienen, Wespen und Ameisen der Kanarischen Inseln (Insecta: Hymenoptera: Aculeata). Veröff. Überseemus. Bremen, Naturwiss, 12, 3–712.
Husemann, M., Deppermann, J., & Hochkirch, A. (2014). Multiple independent colonization of the Canary Islands by the winged grasshopper genus Sphingonotus Fieber, 1852. Molecular Phylogenetics and Evolution, 81, 174–181.
ICZN (International Commission on Zoological Nomenclature). (1999): International Code of Zoological Nomenclature. Fourth edition. International Trust for Zoological Nomenclature, London, xxix + 306 pp.
Juan, C., Oromí, P., & Hewitt, G. M. (1995). Mitochondrial DNA phylogeny and sequential colonization of Canary Islands by darkling beetles of the genus Pimelia (Tenebrionidae). Proceedings of the Royal Entomology Society of London B, 261, 173–180.
Juan, C., Ibrahim, K. M., Oromí, P. & Hewitt, G. M. (1996a). Mitochondrial DNA sequence variation and phylogeography of Pimelia darkling beetles on the island of Tenerife (Canary Islands). Heredity, 77, 589–598.
Juan, C., Oromí, P., & Hewitt, G. M. (1996b). Phylogeny of the genus Hegeter (Tenebrionidae, Coleoptera) and its colonization of the Canary Islands deduced from cytochrome oxidase I mitochondrial DNA sequences. Heredity, 76, 392–403.
Juan, C., Emerson, B. C., Oromí, P., & Hewitt, G. M. (2000). Colonization and diversification: toward a phylogeographic synthesis for the Canary Islands. Trends in Ecology and Evolution, 15, 104–109.
Kearse, M., Moir, R., Wilson, A., Stones-Havas, S., Cheung, M., Sturrock, S., Buxton, S., Cooper, A., Markowitz, S., Duran, C., Thierer, T., Ashton, B., Meintjes, P., & Drummond, A. (2012). Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics, 28, 1647–1649.
Kim, S.-C., Crawford, D. J., Francisco-Ortega, J., & Santos-Guerra, A. (1996). A common origin for woody Sonchus and five related genera in the Macaronesian islands: Molecular evidence for extensive radiation. Proceedings of the National Academy of Science USA, 93, 7743–7748.
Kim, S. C., McGowen, M. R., Lubinsky, P., Barber, J. C., Mort, M. E., & Santos-Guerra, A. (2008). Timing and tempo of early and successive adaptive radiations in Macaronesia. PloS One, 3, e2139.
Klügel, A., Schwarz, S., van den Bogaard, P., Hoernle, K., Wohlgemuth-Ueberwasser, C., & Köster, J. (2009). Structure and evolution of the volcanic rift zone at Ponta de São Lourenço, eastern Madeira. Bulletin of Volcanology, 71(6), 671–685.
Kratochwil, A. (2015). Revision of the Andrena (Micrandrena) tiaretta group: redescription of A. tiaretta Warncke (1974) and description of two new species (A. cyrenaica nov.sp. and A. orientalis nov.sp.) demarcating the central and eastern part of the range (Libya, Israel, Syria). Linzer biol. Beiträge, 47(2), 1403–1437.
Kratochwil, A. (2018). Type specimens of Andrena wollastoni Cockerell, 1922 (Hymenoptera, Anthophila): Deposition, evaluation and designation of a lectotype. Linzer biol. Beiträge, 50(1), 337–351.
Kratochwil, A. (2020). Revision of the Andrena wollastoni group (Hymenoptera, Anthophila, Andrenidae) from the Madeira Archipelago and the Canary Islands: Upgrading of three former subspecies and a description of three new subspecies. Linzer biol. Beiträge, 52(1), 161–244.
Kratochwil, A., & Scheuchl, E. (2013). Andrena (Micrandrena) dourada nov.sp. from Porto Santo, Madeira Archipelago, Portugal. Linzer biol. Beiträge, 45(1), 755–774.
Kratochwil, A., Smit, J., & Aguiar, A. (2018). Updated checklist of the wild bees of the Madeira Archipelago (Hymenoptera: Apoidea: Anthophila). Linzer biol. Beiträge, 50(2), 1213–1228.
Kratochwil, A., & Schwabe, A. (2018a). Wild bees (Anthophila) of Porto Santo (Madeira Archipelago) and their habitats: Species diversity, distribution patterns and bee-plant network. Linzer biol. Beiträge, 50(2), 1219–1247.
Kratochwil, A., & Schwabe, A. (2018b). Wild bees (Anthophila) of Macaronesia — Biogeographical and evolutionary aspects. Berichte Reinhold-Tüxen-Gessellschaft, 30, 149–162.
Kratochwil, A., Schwabe, A., Smit, J., & Aguiar, A. (2019). Wild bee-plant network of Madeira Island with a comparison to Porto Santo (Madeira Archipelago, Portugal). Bol. Mus. Mun. Funchal, 69, 13–32.
Kratochwil, A., & Schwabe, A. (2020). Flower-visiting behaviour and habitats of the taxa of the Andrena wollastoni group (Hymenoptera, Anthophila, Micrandrena) on the Canary Islands compared to the Madeira Archipelago. Linzer biol. Beiträge, 52(1), 309–326.
Leigh, J.W., & Bryant, D. (2015). PopART: Full-feature software for haplotype network construction. Methods in Ecology and Evolution, 6(9), 1110–1116.
Lhomme, P., Michez, D., Christmann, S., Scheuchl, E., El Abdouni, I., Hamroud, L., Ihsane, O., Sentil, A., Smaili, M. C., Schwarz, M., Dathe, H. H., Straka, J., Pauly, A., Schmid-Egger, C., Patiny, S., Terzo, M., Müller, A., Praz, C., Risch, S., Kasparek, M., Kuhlmann, M., Wood, T. J., Bogusch, P., Ascher, J., & Rasmont, P. (2020). The wild bees (Hymenoptera: Apoidea) of Morocco. Zootaxa, 4892(1), 1–159.
Lindberg, H. (1953). Hemiptera Insularum Canariensium. Commentationes Biologica, 14, 1–304.
Lindberg, H., Lindberg, H., & Roudier, A. (1958). Coleoptera Insularum canariensium. 1. Aglycyderidae and Curculionidae. Commentationes Biologica, 17, 1–97.
López, H., Contreras Díaz, H. G., Oromí, P., & Juan, C. (2007). Delimiting species boundaries for endangered Canary Island grasshoppers based on DNA sequence data. Conservation Gentics, 8, 587–598.
Maca-Meyer, N., Carranza, S., Rando, J. C., Arnold, E. N., & Cabrera, V. M. (2003). Status and relationships of the extinct giant Canary island lizard Gallotia goliath (Reptilia: Lacertidae), assessed using ancient mtDNA from its mummified remains. Biological Journal of the Linnean Society, 80, 659–670.
MacArthur, R. H., & Wilson, E. O. (1967). The theory of island biogeography. Princeton Univ. Press, Princeton.
Machado, A. (1976). Introduction to a faunal study of the Canary Islands’ laurisilva, with special reference to the ground-beetles (Coleoptera, Caraboidea). In Kunkel G. (Ed.) Biogeography and ecology in the Canary Islands (pp. 347–412). Dr. W. Junk b.v. Publishers, The Hague.
Machado, A., Rodríguez-Expósito, E., López, M., & Hernández, M. (2017). Phylogenetic analysis of the genus Laparocerus, with comments on colonisation and diversification in Macaronesia (Coleoptera, Curculionidae, Entiminae). ZooKeys, 651, 1–77.
Magnacca, K. N., & Danforth, B. N. (2006). Evolution and biogeography of the native Hawaiian Hylaeus bees (Hymenoptera: Colletidae). Cladistics, 22, 393–411.
Magnacca, K., & Danforth, B. N. (2007). Low nuclear DNA variation supports a recent origin of Hawaiian Hylaeus bees (Hymenoptera: Colletidae). Mol. Phyl. Evol., 43, 908–915.
Magnacca, K,. & King, C. (2013). Assessing the presence and distribution of 23 Hawaiian yellowed-faced bee species on lands adjacent to military installations on O’ahu and Hawai’i Island (26 pp. and appendix). Pacific Cooperative Studies Unit, University of Hawai’i at Manoa: Honolulu, HI, USA.
Marrero, Á. (2004). Procesos evolutivos en plantas insulares, el caso de Canarias. In Fernández-Palacios J.Mm, Morici C. (Eds.) Ecología insular/Island ecology (pp. 305–356). Excmo. Cabildo Insular de La Palma y Asociación Española de Ecología Terrestre, Santa Cruz de La Palma.
Marrero, Á., & Francisco-Ortega, J. (2001). 14. Evolución en islas: la metáfora espacio-tiempo-forma. In Fernández-Palacios J.M. & J.L. Martín Esquivel (Eds.) Naturaleza de las islas Canarias Ecología y conservación (pp. 133–140). Turquesa Ediciones, Santa Cruz de Tenerife.
Mas-Peinado, P., Buckley, D., Ruiz, J. L., & García-París, M. (2018). Recurrent diversification patterns and taxonomic complexity in morphologically conservative ancient lineages of Pimelia (Coleoptera: Tenebrionidae). Systematic Entomology, 43, 522–548.
Mort, M. E., Soltis, D. E., Soltis, P. S., Francisco-Ortega, J., & Santos-Guerra, A. (2002). Phylogenetics and evolution of the Macaronesian clade of Crassulaceae inferred from nuclear and chloroplast sequence data. Systematic Botany, 27, 271–288.
Nylander, J. A. A. (2004). MrModeltest v2Program distributed by the author. Uppsala University.
Oromí, P., Martín, J. L., Medina, A. L., & Izquierdo, I. (1991). The evolution of the hypogean fauna in the Canary Islands. In Dudley E.C. (Ed.) The unity of evolutionary biology (pp. 380–395). Vol. 2. Dioscorides Press, Portland, Oreg.
Papadopoulou, A., Anastasiou, I., & Vogler, A. P. (2010). Revisiting the insect mitochondrial molecular clock: The mid-Aegean trench calibration. Molecular Biology and Evolution, 27, 1659–1672.
Pallarés Padilla, A. (2007). Nuevas aportaciones al conocimiento de la erupción de Timanfaya, Lanzarote. Academia de Ciencias e Ingenierías de Lanzarote. Discursos Académicos, 22, 3–45.
Paxton, R. J., Thorén, P. A., Tengö, J., Estoup, A., & Pamilo, P. (1996). Mating structure and nestmate relatedness in a communal bee, Andrena jacobi (Hymenoptera: Andrenidae), using microsatellites. Molecular Ecology, 5, 511–519.
Percy, D. M., & Cronk, Q. C. B. (2002). Different fates of island brooms: Contrasting evolution in Adenocarpus, Genista, and Teline (Genisteae, Fabaceae) in the Canary Islands and Madeira. American Journal of Botany, 89, 854–864.
Pérez-Torrado, F. J., & Mangas, J. (1994). El estratovolcán Roque Nublo (Gran Canaria): estratigrafía y modelo evolutivo. In García A. & A. Felpeto (Eds.) In memoriam Dr José Luis Díez Gil (pp. 179–192). Cabildo Insular de Lanzarote; Serie Casa de los Volcanes nº 3, Arrecife
Pisanty, G., Richter, R., Martin, T., Dettman, J., & Cardinal, S. (2021). Molecular phylogeny, historical biogeography and revised classification of andrenine bees (Hymenoptera: Andrenidae). Molecular Phylogenetics and Evolution. https://doi.org/10.1016/j.ympev.2021.107151
Press, J. R., & Short, M. J. (1994). Flora of Madeira. The Natural History Museum London. St. Edmundsbury Press, Suffolk. 574 pp.
Rambaut, A. (2009). FigTree v.1.4.0 2006–2012. Available at http://tree.bio.ed.ac.uk/software/figtree/.
Rangel López, J. A., Husemann, M., Schmitt, T., Kramp, K., & Habel, J. C. (2018). Mountain barriers and trans-Saharan connections shape the genetic structure of Pimelia darkling beetles (Coleoptera: Tenebrionidae). Biological Journal of the Linnean Society, 124, 547–556.
Rasmussen, C., Carrion, A. L., Castro-Urgal, R., Chamorro, S., González, V. H., Griswold, T. L., Herrera, H. W., McMullen, C. K., Olesen, J. M. & Traveset, A. (2012). Megachile timberlakei Cockerell (Hymenoptera: Megachilidae): Yet another adventive bee species to the Galápagos Archipelago. The Pan-Pacific Entomologist, 88(l), 98–102.
R Core Team (2016). R: A language and environment for statistical computing. R Foundation for statistical computing, Vienna, Austria [WWW document]. R 3.6.0 GUI 1.70. https://www.R-project.org.
Rees, D. J., Emerson, B. C., Oromí, P., & Hewitt, G. M. (2001). Reconciling gene trees with organism history: The mtDNA phylogeography of three Nesotes species (Coleoptera: Tenebrionidae) on the western Canary Islands. Journal of Evolutionary Biology, 14, 139–147.
Ronquist, F., & Huelsenbeck, J. P. (2003). MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics, 19, 1572–1574.
Rozas, J., Ferrer-Mata, A., Sánchez-DelBarrio, J. C., Guirao-Rico, S., Librado, P., Ramos-Onsins, S. E., & Sánchez-Gracia, A. (2017). DnaSP 6: DNA Sequence polymorphism analysis of large datasets. Molecular Biology and Evolution, 34, 3299–3302. https://doi.org/10.1093/molbev/msx248
Saether, S. A., Fiske, P., Kålås, J. A., Kuresoo, A., Luigujõe, L., Piertney, S. B., Sahlman, T., & Höglund, J. (2007). Inferring local adaptation from QST-FST comparisons: neutral genetic and quantitative trait variation in European populations of great snipe. Journal of Evolutionary Biology, 20(4), 1563–76. https://doi.org/10.1111/j.1420-9101.2007.01328.x.
Salomone, N., Emerson, B. C., Hewitt, G. M., & Bernini, F. (2002). Phylogenetic relationships among the Canary Island Steganacaridae (Acari, Oribatida) inferred from mitochondrial DNA sequence data. Molecular Ecology, 11, 79–89.
Schmidt, S., Schmid-Egger, C., Morinére, J., Haszprunar, G., & Hebert, P. D. N. (2015). DNA barcoding largely supports 250 years of classical taxonomy: Identifications for Central European bees (Hymenoptera, Apoidea partim. Molecular Ecology Resources, 15, 985–1000.
Schmincke, H. U. (1998). Zeitliche, strukturelle und vulkanische Entwicklung der Kanarischen Inseln, der Selvagens-Inseln und des Madeira-Archipels. In Bischoff W. (Ed.) Die Reptilien der Kanarischen Inseln, der Selvagens-Inseln und des Madeira-Archipels (pp. 27–69). Aula, Wiesbaden.
Silva, J. B. P. (2003). Areia de Praia da Ilha do Porto Santo. Madeira Rochas, 344 pp.
Thorpe, R. S., McGregor, D., Cummings, A. M., & Jordan, W. C. (1994). DNA evolution and colonization sequence of island lizards in relation to geological history: mtDNA RFLP, cytochrome b, cytochrome oxidase, 12srRNA sequence and nuclear RAPD analysis. Evolution, 48, 230–240.
Vandergast, A. G., & Gillespie, R. G. (2004). Effects of natural forest fragmentation on a Hawaiian spider community. Environmental Entology, 33, 1296–1305.
Venables, W. N., & Ripley, B. D. (2002). MASS library of functions. Modern applied statistics with S. New York: Springer.
Vu, V. Q. (2011). ggbiplot: A ggplot2 based biplot. R package version 0.55.
Warncke, K. (1968). Zur Kenntnis der Bienengattung Andrena F. auf den Kanarischen Inseln. Notulae Entomologicae, 48, 63–80.
Weihrauch, F. (2011). A review of the distribution of Odonata in the Macaronesian Islands, with particular reference to the Ischnura puzzle. Journal of the British Dragonfly Society, 27(1), 28–46.
Widmer, A., Schmid-Hempel, P., Estoup, A., & Scholl, A. (1998). Population genetic structure and colonization history of Bombus terrestris s.l. (Hymenoptera: Apidae) from the Canary Islands and Madeira. Heredity, 81, 563–572.
Wunderlich, J. (1991). The spider fauna of the Macaronesian Islands. Taxonomy, ecology, biogeography and evolution. Journal of the British Dragonfly Society, Beiträge zur Araneologie, 1, 1–619.
Acknowledgements
Special thanks go to the facilities and authorities in the Madeira Archipelago and the governments of the Canary Islands for access and collection permits. We thank Fritz Gusenleitner (Biology Centre of the Upper Austrian Provincial Museum Linz) for his permission to use the Warncke collection for analysing specimens in Linz (Austria) and Volker Lohrmann and Herbert Hohmann for the possibility of analysing specimens from the museum collection of the Überseemuseum Bremen (Germany). For borrowing specimens from his private collection, we gratefully thank Hans Richard Schwenninger (Stuttgart, Germany). We are indebted to Hannes Baur (Natural History Museum Berne, Switzerland), who informed us about new tools for morphometric analysis. Erwin Scheuchl (Germany), in particular, inspired us in many discussions to see that it is an important task to analyse Micrandrena species. We thank Eoin Laws-Quinn for assistance with molecular genetic work in the laboratory.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kratochwil, A., Paxton, R.J., Schwabe, A. et al. Morphological and genetic data suggest a complex pattern of inter-island colonisation and differentiation for mining bees (Hymenoptera: Anthophila: Andrena) on the Macaronesian Islands. Org Divers Evol 22, 189–204 (2022). https://doi.org/10.1007/s13127-021-00513-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13127-021-00513-z