Abstract
Acute lymphoblastic leukemia (ALL) is the most common pediatric cancer with precursor B-cell ALL (pB-ALL) accounting for ~ 85% of the cases. Childhood pB-ALL development is influenced by genetic susceptibility and host immune responses. The role of the intestinal microbiome in leukemogenesis is gaining increasing attention since Vicente-Dueñas’ seminal work demonstrated that the gut microbiome is distinct in mice genetically predisposed to ALL and that the alteration of this microbiome by antibiotics is able to trigger pB-ALL in Pax5 heterozygous mice in the absence of infectious stimuli. In this review we provide an overview on novel insights on the role of the microbiome in normal and preleukemic hematopoiesis, inflammation, the effect of dysbiosis on hematopoietic stem cells and the emerging importance of the innate immune responses in the conversion from preleukemic to leukemic state in childhood ALL. Since antibiotics, which represent one of the most widely used medical interventions, alter the gut microbial composition and can cause a state of dysbiosis, this raises exciting epidemiological questions regarding the implications for antibiotic use in early life, especially in infants with a a preleukemic “first hit”. Sheading light through a rigorous study on this piece of the puzzle may have broad implications for clinical practice.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Genetic susceptibility and exogenous exposures
Childhood pB-ALL development is influenced by a combination of genetic susceptibility and exogenous factors.
Germline predisposition is an important risk factor underlying the development of pediatric ALL [1, 2], present in 3–10% of childhood ALL cases [3]. In pB-ALL the genetic susceptibility ranges from frequent but low-penetrant prenatal somatic chromosomal aberrations [4,5,6,7,8] and adverse combinations of low penetrant germline variations [2, 9], to rare but highly penetrant germline mutations [10].
Germline variations predisposing to ALL include constitutional syndromes like Down syndrome[11], Noonan syndrome[12], familial cancer syndromes, Li-Fraumeni syndrome[13], constitutional mismatch repair deficiency syndrome[14, 15]), PAX5 mutations[16, 17], ETV6 variants [18, 19] as well as genes harboring germline non-silent variants presumed to confer a risk of sporadic ALL like NBN, FLT3, SH2B3, and CREBBP [2, 10].
The most frequent fusion gene in pediatric ALL, resulting from t(12;21)(p13.2;q22.1), is the ETV6-RUNX1 fusion, which occurs in approximately 25% of childhood pB-ALL [3]. Pre-leukemic clones carrying ETV6-RUNX1 oncogenic lesions have been found in 1–5% of healthy newborn cord bloods [20, 21], with differences based mainly on different detection methods, with at an estimated frequency that was 100–500-fold greater than the risk of the corresponding leukemia [7, 20,21,22,23,24]. Prenatal low penetrant somatic chromosomal alterations in pB-ALL include the ETV6-RUNX1 fusion [4], high hyperdiploidy [5], the TCF3-PBX1 gene fusion [6, 25] and BCR-ABL1 fusions [8]. Why only a fraction of children born with these preleukemic “hits” develop full-blown leukemia is still unknown, highlighting the fact that a second oncogenic step is required for overt leukemia development.
Although childhood pB-ALL does not generally cluster geographically, various pathogens have been reported linked to pB-ALL in space–time clusters, such as peaks after seasonal influenza [26], adenoviral infection [27], and H1N1 infection clusters [28]. Greaves hypothesized that while microbial exposures earlier in life are protective, in their absence delayed exposure of an immature untrained immune system to infection results in altered immune responses triggering critical secondary mutations [20]. Epidemiological factors linked to a protective effect for ALL in children less than one year old include birth order [29], mode of delivery [30,31,32,33], breastfeeding [34], early day care attendance [35,36,37,38,39], early common infections [40] and animal contact [41], supporting the hypothesis that infections have a protective effect for ALL in children less than one year of age [42]. Concomitant with rising hygiene standards, the incidence of pB-ALL has increased in developed countries [43,44,45,46]. More hygienic environments may reduce the chance to properly train the immune system to prevent secondary mutations during later infections. Modulation of the innate and adaptive immune system by vaccines has been associated to protection from ALL, in particular early BCG vaccination before 3 months of age [47]. The Bacillus Calmette-Guérin (BCG) vaccine is a live attenuated tuberculosis vaccine that has the ability to induce non-specific cross-protection against pathogens that might be unrelated to the target disease. A meta-analysis of 12 studies observed that early vaccination (< 3 months of age) with the Bacillus Calmette–Guérin (BCG) vaccine results in statistically robust protection from ALL [47]. In line with this observation, differences in vaccination protocols of BCG vaccination in East and West Germany prior to reunification, with compulsory BCG vaccination in East but not in West Germany, correlated with a lower rate of childhood leukemia in East Germany before reunification, which increased to West German levels 8 years after reunification [48].
2 Insights from preclinical pB-ALL models
The “innate immune memory” or “trained immunity” immunologic principle highlights the responses of innate immune cells and their memory properties after immune stimuli [49]. Macrophages, monocytes and NK cells undergo metabolic and epigenetic modifications following exposure to infection or vaccination, providing innate immune cells with a memory, which subsequently modulates their response to a second infection exposure later in life [50]. The lasting immunologic memory is mediated by persistent epigenetic modifications in hematopoietic stem cells (HSC) and myeloid progenitors and depends on the transcription factor CCAAT/enhancer-binding protein b [51]. When cells of the hematopoietic niche are trained with beta-glucan and BCG, they generate accumulation of H3K4me3 and H3K27ac histone complexes, which are partially preserved when HSC differentiate along myeloid and lymphoid lineages [52]. A new mechanism of inherited trained immunity has been described in mice [53], conferring further protection against infections to the offsping. Innate host immune responses can influence the penetrance of B-ALL. The protective role of early immune training in B-ALL progression even in the presence of first hits was explored by Fidanza et al. in Eμ-ret and E2A-PBX1 transgenic murine B-ALL models [54]. Ex vivo stimulation of leukemia-initiating precursor B-cells derived from spleens of 4-week-old Eμ-ret mice with TLR7 ligand, TLR8 ligand or TLR9 ligand showed reduced viable cell recovery, while increased cell expansion following TLR3 stimulation was observed. TLR9 stimulation induced long-term control of preleukemia and established leukemia in the same Eμ-ret model. Innate immune cells, in particular natural killer cells and macrophages were critical in mediating these effects and protected these mice in remission from relapse after leukemia re-challenge. Treatment with IFN-α- or IFN-γ-neutralizing antibodies reversed these effects, implying that proliferation or regression of leukemia initiating cells is interferon-dependent [55].
Loss of immunosurveillance within the bone marrow microenvironment is a further mechanism leading to immune evasion of ALL blasts. The “infective lymphoid recovery hypothesis” described by Richardson et al. focuses on the leukemia-promoting effects of delayed recurrent infections in childhood, causing heat-shock responses and transient involution of the thymus and other lymphoid organs, contributing to a decline in antitumor immuno-surveillance and the maturation arrest of B-cells secondary to glucocorticoid release in response to infection. The release of pro-inflammatory (Th1) cytokines in response to infection promotes cell survival and a hypermutable state, whereas the release of Th2 cytokines and interleukin-7 (IL-7) stimulates immature B-cell proliferation, including preleukemic cells [56].
Inflammation impacts preleukemic, stromal cells and the immune system in the bone marrow. An inflammation-microenvironment—leukemia initiating cell triad has been proposed as a key pathogenic mechanism in leukemia pathobiology [57]. The putative pediatric pB-ALL initiating niche shows gene expression signatures of bone marrow mesenchymal stem cell (MSC) with pro-inflammatory and suppressor niches [58]. The transcriptional profiles of MSC reveal two characteristic signatures: a CXCL12-low inflammatory and leukemia expansion-like niche that likely supports leukemic burden and a CXCL11 high immune-suppressive and leukemia-initiating cell (SLIC)-like niche, where LICs are likely sustained. The pro-inflammatory signature includes a large set of chemokines involved in neutrophil recruitment, metalloproteinase functional activation and leukocyte migration as well as pro-inflammatory molecules. The immune suppressive signature shows TLR signaling, cytokine mediated signaling and a negative regulation of leukocyte proliferation signatures with high expression of chemokines CXCL10 and CXCL11 as well as suppressor molecules like indole amine 2,3-dioxygenase (IDO1) and galectin 9 (LGALS9). In ALL, pro-inflammatory cytokine signaling including IL6, TNFa and IL-1b is involved in promoting leukemia initiation by cooperating with mesenchymal stromal cell (MSC) niches in selection and induction of ETV6-RUNX1 pre-leukemic cells [59]. Key cells of the immune system that shape the leukemic microenvironment include immature myeloid populations, non-classical monocytes, macrophages, NK cells, and various subpopulations of T cells [60]. Immature myeloid populations involved in immunoregulation, referred to as myeloid-derived suppressor cells (MDSCs), include granulocytic (G) and monocytic (M) MSDCs [61, 62]. G-MDSCs inhibit the activity of T cells and natural killer (NK) cells through reactive oxygen species (ROS)-dependent mechanisms, and M-MDSCs through secretion of immunosuppressive cytokines (IL-10, TGF beta), arginase and nitric oxide. ALL blasts evade immunosurveillance escaping from NK cell recognition and lysis through downregulation of the ligands for NK cell-activating receptors (MICA/B, ULBP1, and NEC-2) [63], resist phagocytosis by macrophages by upregulating CD47 [64] and avoid a T cell response by triggering selected inhibitory checkpoints (TIM-3, CD200R) [65]. Increased Treg-cells have been linked with diminished potency of selected immunotherapies against pB-ALL, such as blinatumomab [66] or CAR-T cells [67]. The CXCL10/CXCL11/CXCR3 axis has been implicated in chemotherapy resistance and CNS infiltration in B-ALL [68]. Through single-cell transcriptome profiling Witkowski et al. elegantly compared the bone marrow immune microenvironment of healthy individuals and of B-ALL patients at diagnosis, remission, and relapse. The non-malignant immune bone marrow microenvironment in ALL patients is altered and shows enrichment with a non-classical monocyte subpopulation both at diagnosis and at relapse [69]. Differential gene expression analysis of leukemia-associated non-classical monocytes and their healthy counterpart show a leukemia-associated upregulation of genes involved in monocyte interactions with vascular endothelium during inflammation and vascular endothelial repair, such as PECAM1, CD44, ITGA4, CX3CR1, and TNFSF10, as well as significant downregulation of genes encoding subunits of HLA-DR. The differentiation of human classical monocytes toward the non-classical monocyte lineage is enhanced by the presence of human B-ALL. Acute leukemia interacts with the vascular endothelium leading to increased vascular permeability, and non-classical monocytes emerge in response to leukemia-induced inflammation to repair the damaged endothelium.
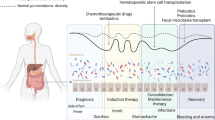
A further element gaining recognition in this puzzle is the role of the intestinal microbiome in pB-ALL [70, 71]. Within a human organism there are trillions of microbes that interact with the host constantly, mainly on the skin and on mucosal surfaces of the gastrointestinal tract. The gut microbiota, which mainly resides in the colon, develops during the first three years of life [72,73,74]. The structure of the gut, which contains a large mucosa surface consisting of a single epithelial cell layer made up of intestinal epithelial cells and intraepithelial lymphocytes, facilitates the interaction with the immune system. This gut-associated lymphoid tissue represents the largest component of the immune system in the body and influences immune responses locally and systemically. Bacterial fermentation of undigested dietary carbohydrates, mainly resistant starch and dietary fiber, produces short chain fatty acids (SCFA), primarily acetate, propionate, and butyrate, which are major players in the maintenance of gut physiology and integrity by promoting immune and metabolic homeostasis, with important anti-inflammatory effects [75], also influencing hematopoiesis through G protein–coupled receptors [76,77,78]. While alpha diversity is a measure of microbiome diversity applicable to a single sample, beta diversity is a measure of similarity or dissimilarity of two communities from different environments. Antibiotic-induced microbiota alterations have been shown to alter bile acid metabolism and insulin sensitivity in both humans and mice [79]. The specific composition of the microbiota alters the types and levels of SCFA that are formed, affecting numerous physiologic processes that are differentially modulated by acetate, propionate, and butyrate [80]. Besides the direct effect of antibiotics on the composition of the gut microbiota, antibiotics affect the manner in which this community interacts with the host and regulates basic physiological processes [81]. Bacterial byproducts can skew the dendritic cells of the gut-associated lymphoid tissue either towards a Treg-state, creating a local anti-inflammatory cytokine environment, or toward a Th17-state and creating an inflammatory phenotype [82]. The term dysbiosis refers to a persistent alteration of the gut microbiota, both in its composition and function [83], with a less diverse and less stable microbiota. Dysbiosis is commonly characterized by a reduced diversity of the phyla Firmicutes and Bacteroidetes and is often accompanied by an overgrowth of the family Enterobacteriaceae. Bacterial metabolites like the SCFA produced by Bifidobacteriaceae are involved in the host defense of colonic epithelium, affecting the mucus layer structure and consequently the host’s gastrointestinal barrier function [84]. As the microbiota composition changes, the altered microbial community will present a different repertoire of microbial-associated molecular patterns (MAMPS) to the receptors located in immune and epithelial cells, which results in an altered stimulation of receptors such as NOD1 and the Toll-like receptors (TLRs), which are involved in lymphoid tissue development, T cell differentiation, neutrophil priming and cytokine release [85]. HSC and progenitor cells express TLRs, which are able to recognize microbial components and initiate innate immune responses. Bacterial lipopolysaccharide (LPS) binds the TLR4/Myd88 receptor complex on hematopoietic stem cells and stimulates myeloid differentiation pathways, providing a means for rapid replenishment of the innate immune system during infection [86]. Nevertheless, repeated exposure to small amounts of LPS is harmful to long-term repopulating stem cells and induces the loss of the ability to differentiate towards lymphocytes with an enduring myeloid bias and lymphoid deficiency in HSC originating from marrow chronically exposed to LPS [87]. In Vicente-Dueñas et al.’s study [71] neither a single microbe nor the overall bacterial burden were connected with the development of pB-ALL, but in particular the depletion and subsequent reconstitution of the microbiome were significantly associated with pB-ALL. The intestinal microbiome of mice genetically predisposed to ALL through Pax5 heterozygosity or ETV6-RUNX1 fusion had a specific shape when compared to wild-type mice, and the alteration of this already distinct microbiome of Pax5 heterozygous mice through antibiotic treatment was able to trigger pB-ALL in the absence of infectious stimuli [71]. Depleting the gut microbiome by antibiotic treatment did not prevent infection-driven pB-ALL development in Pax5 1/2 mice in core facilities, but remarkably, this intervention promoted pB-ALL development in a specific pathogen-free environment, even in the absence of infectious stimuli [44]. Zhang et al. showed that microbiome regulates neutrophil ageing [88] and furthermore, that the microbiome is required to balance hematopoiesis. Microbiome depletion with antibiotics reduced the aged neutrophils, which are highly inflammatory. Neutrophil ageing is driven by the microbiome via TLR and myeloid differentiation factor88 (MyD88)-mediated signaling pathways, which transduce microbiome-derived signals. Microbiome depletion in specific-pathogen-free mice with prolonged broad-spectrum antibiotics resulted in significant and selective reductions of circulating aged neutrophils which were completely restored when TLR4 ligand LPS was added. In contrast, germ-free animals showed broad alterations in both innate and adaptive immune cells, with significantly reduced numbers of total and aged neutrophils. Treatment of germ-free mice with antibiotics did not further reduce the aged neutrophils but were partially restored when germ-free mice were reconstituted by fecal transplantation, suggesting that the microbiome is involved in a fine tuned and balanced regulation of immunity, influencing the proportion of highly active aged neutrophils and balancing hematopoiesis and the risk of tissue injury. These results are in line with Vicente-Dueñas et al.’s study [71], where the depletion and subsequent reconstitution of the microbiome by antibiotics in mice genetically predisposed to ALL through Pax5 heterozygosity was significantly associated with pB-ALL in specific pathogen-free environment but not in core facilities. One may hypothesize that whereas a healthy, species-rich microbiome balances normal hematopoiesis, the depletion of the microbiome by antibiotics followed by a state of dysbiosis, with persistently low plasmatic LPS levels may contribute to a lymphoid-myeloid shift and the generation of a myeloid inflammatory microenvironment, predisposing pre-leukemic cells to progression to overt B-ALL (Fig. 1).
Innate immune cells undergo epigenetic modifications following infections or vaccination, which provide them with a memory, which subsequently modulates their response to a second infection exposure later in life (“trained immunity”). Delayed recurrent infections cause a decline in antitumor immuno-surveillance and the maturation arrest of B-cells secondary to glucocorticoid release in response to infection. The release of pro-inflammatory (Th1) cytokines promotes cell survival and a hypermutable state, whereas the release of Th2 cytokines and IL-7 stimulates immature B-cell proliferation, including preleukemic cells. The putative pediatric pB-ALL initiating niche shows gene expression signatures of bone marrow mesenchymal stem cell (MSC) with pro-inflammatory and suppressor niches. The differentiation of human classical monocytes towards the non-classical monocyte lineage is enhanced by the presence of human B-ALL. Acute leukemia interacts with the vascular endothelium leading to increased vascular permeability and non-classical monocytes emerge in response to leukemia-induced inflammation. Hematopoietic stem cells (HSC) express Toll-like receptors (TLRs), which are able to recognize microbial components and initiate innate immune responses. Bacterial lipopolysaccharide (LPS) binds the TLR4/Myd88 receptor complex on HSC and stimulates myeloid differentiation pathways, providing a means for rapid replenishment of the innate immune system during infection. The microbiome regulates neutrophil ageing and is required to balance hematopoiesis. A state of dysbiosis with persistent low plasmatic LPS leves can induce loss of lymphoid differentiation in HSC and a myeloid shift. One may hypothesize that whereas a healthy, species-rich microbiome balances normal hematopoiesis, the depletion of the microbiome by antibiotics followed by a state of dysbiosis, with persistently low plasmatic LPS levels may contribute to a lymphoid-myeloid shift and the generation of a myeloid inflammatory microenvironment, predisposing pre-leukemic cells to progression to overt pB-ALL
3 Antibiotics in childhood, microbiome changes and data in childhood ALL patients
It has been shown in children, adults and in animal models that antibiotics dramatically alter the gut microbial composition [81, 89, 90]. Since antibiotics are among the most common medications prescribed to children in high-income countries [91], the potential for dysbiosis in the microbiome should be taken into account.
The table below (Table 1) summarizes standardized principles of antibiotic prescription for the most frequent infectious diseases in children obtaining care in an outpatient setting for the following diagnosis: acute rhinosinusitis, acute otitis media, bronchiolitis, pharyngitis, non-specific upper respiratory tract infections, and urinary tract infection.
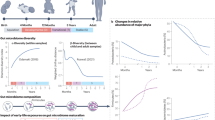
Yassour et al. performed an elegant longitudinal study on the infant gut microbiome and the impact of antibiotic treatment on bacterial strain diversity and stability. Based on whole-genome shotgun sequencing of monthly stool samples over 36 months in 39 children, half of whom received multiple courses of antibiotics during the first 3 years of life, it was shown that the microbiota of antibiotic-treated children had reduced bacterial and strain diversity and a less stable gut microbiome following antibiotic treatment. The most current evidence on gut microbiome dysbiosis in children after antibiotic treatment has been systematically reviewed by McDonnel et al. [101]. Only twelve studies met quality eligibility criteria for this systematic review and included five randomized controlled trials, five cohort studies and two cross-sectional studies that analyzed the relationship between antibiotics and gut microbiome dysbiosis, age 0–18 years, molecular techniques of assessment and outcomes of microbiome richness, diversity or composition. Five studies found a significant reduction in diversity and three of them a significant reduction in richness. Antibiotic exposure was associated with reduced microbiome diversity and richness, and with alterations in bacterial abundance, with reductions of Bifidobacteria and Lactobacillus and increases in Proteobacteria like E. coli. Macrolide exposure (azithromycin) was associated with reduced richness for twice as long as penicillin and a significant reduction in alpha-diversity. Quality of evidence in this rigorous meta-analysis was defined as good or fair [101], highlighting the need for further standardized research in this field [102].
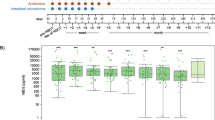
The microbiome of pediatric ALL patients has a specific shape starting from the moment of diagnosis when compared to healthy controls. Several studies have shown that ALL patients differ in their gastrointestinal microflora at diagnosis when compared to healthy controls [103,104,105,106,107,108]. Additional evidence supporting this fact is that also the supragingival plaque is less rich and less diverse in pediatric ALL patients at diagnosis comparted with healthy controls [109]. As described by Oldenburg et al. at the onset of ALL, reduced diversity in the oral and gut microbiomes in ALL patients is already observed [110]. Further reduction of diversity occurs during treatment due to the administration of chemotherapeutics and antibiotics, with dominance of Enteroccocaceae being predictive of infections. Changes to the microbiome can be detected up to several years after completion of treatment, with possible implications for long-term health [110].
4 Summary
The reviewed insights on the role of the innate immune responses in childhood pB-ALL models highlight the trained immunity concept as a possible target for future interventions towards prevention of pB-ALL in children. The role of the microbiome in normal hematopoiesis and the effect of dysbiosis on HSC is gaining increasing attention particularly since the observation that antibiotic treatment is able to trigger pB-ALL in the absence of infectious stimuli in Pax5 heterozygous mice genetically predisposed to ALL [71]. Since antibiotics, which represent one of the most widely used medical interventions, alter the gut microbial composition and can cause a state of dysbiosis, this raises exciting epidemiological questions regarding the implications for antibiotic use in early life especially in infants with a a preleukemic “first hit”. Sheading light through a rigorous epidemiologic study on this piece of the puzzle may have broad implications for clinical practice.
References
Bloom M, Maciaszek JL, Clark ME, Pui CH, Nichols KE. Recent advances in genetic predisposition to pediatric acute lymphoblastic leukemia. Expert Rev Hematol. 2020;13(1):55–70. https://doi.org/10.1080/17474086.2020.1685866.
Vijayakrishnan J, Qian M, Studd JB, Yang W, Kinnersley B, Law PJ, Broderick P, Raetz EA, Allan J, Pui CH, Vora A, Evans WE, Moorman A, Yeoh A, Yang W, Li C, Bartram CR, Mullighan CG, Zimmerman M, Hunger SP, Schrappe M, Relling MV, Stanulla M, Loh ML, Houlston RS, Yang JJ. Identification of four novel associations for B-cell acute lymphoblastic leukaemia risk. Nat Commun. 2019;10(1):5348. https://doi.org/10.1038/s41467-019-13069-6.
Pui CH, Relling MV, Downing JR. Acute lymphoblastic leukemia. N Engl J Med. 2004;350(15):1535–48. https://doi.org/10.1056/NEJMra023001.
Ford AM, Bennett CA, Price CM, Bruin MC, Van Wering ER, Greaves M. Fetal origins of the TEL-AML1 fusion gene in identical twins with leukemia. Proc Natl Acad Sci USA. 1998;95(8):4584–8. https://doi.org/10.1073/pnas.95.8.4584.
Maia AT, van der Velden VH, Harrison CJ, Szczepanski T, Williams MD, Griffiths MJ, van Dongen JJ, Greaves MF. Prenatal origin of hyperdiploid acute lymphoblastic leukemia in identical twins. Leukemia. 2003;17(11):2202–6. https://doi.org/10.1038/sj.leu.2403101.
Wiemels JL, Leonard BC, Wang Y, Segal MR, Hunger SP, Smith MT, Crouse V, Ma X, Buffler PA, Pine SR. Site-specific translocation and evidence of postnatal origin of the t(1;19) E2A-PBX1 fusion in childhood acute lymphoblastic leukemia. Proc Natl Acad Sci USA. 2002;99(23):15101–6. https://doi.org/10.1073/pnas.222481199.
Hein D, Borkhardt A, Fischer U. Insights into the prenatal origin of childhood acute lymphoblastic leukemia. Cancer Metastasis Rev. 2020;39(1):161–71. https://doi.org/10.1007/s10555-019-09841-1.
Cazzaniga G, van Delft FW, Lo Nigro L, Ford AM, Score J, Iacobucci I, Mirabile E, Taj M, Colman SM, Biondi A, Greaves M. Developmental origins and impact of BCR-ABL1 fusion and IKZF1 deletions in monozygotic twins with Ph+ acute lymphoblastic leukemia. Blood. 2011;118(20):5559–64. https://doi.org/10.1182/blood-2011-07-366542.
Vijayakrishnan J, Studd J, Broderick P, Kinnersley B, Holroyd A, Law PJ, Kumar R, Allan JM, Harrison CJ, Moorman AV, Vora A, Roman E, Rachakonda S, Kinsey SE, Sheridan E, Thompson PD, Irving JA, Koehler R, Hoffmann P, Nöthen MM, Heilmann-Heimbach S, Jöckel K-H, Easton DF, Pharaoh PDP, Dunning AM, Peto J, Canzian F, Swerdlow A, Eeles RA, Kote-Jarai Z, Muir K, Pashayan N, Greaves M, Zimmerman M, Bartram CR, Schrappe M, Stanulla M, Hemminki K, Houlston RS. Genome-wide association study identifies susceptibility loci for B-cell childhood acute lymphoblastic leukemia. Nat Commun. 2018. https://doi.org/10.1038/s41467-018-03178-z.
Inaba H, Mullighan CG. Pediatric acute lymphoblastic leukemia. Haematologica. 2020;105(11):2524–39. https://doi.org/10.3324/haematol.2020.247031.
Izraeli S. The acute lymphoblastic leukemia of Down Syndrome—genetics and pathogenesis. Eur J Med Genet. 2016;59(3):158–61. https://doi.org/10.1016/j.ejmg.2015.11.010.
Cave H, Caye A, Strullu M, Aladjidi N, Vignal C, Ferster A, Mechinaud F, Domenech C, Pierri F, Contet A, Cacheux V, Irving J, Kratz C, Clavel J, Verloes A. Acute lymphoblastic leukemia in the context of RASopathies. Eur J Med Genet. 2016;59(3):173–8. https://doi.org/10.1016/j.ejmg.2016.01.003.
Qian M, Cao X, Devidas M, Yang W, Cheng C, Dai Y, Carroll A, Heerema NA, Zhang H, Moriyama T, Gastier-Foster JM, Xu H, Raetz E, Larsen E, Winick N, Bowman WP, Martin PL, Mardis ER, Fulton R, Zambetti G, Borowitz M, Wood B, Nichols KE, Carroll WL, Pui CH, Mullighan CG, Evans WE, Hunger SP, Relling MV, Loh ML, Yang JJ. TP53 germline variations influence the predisposition and prognosis of B-cell acute lymphoblastic leukemia in children. J Clin Oncol. 2018;36(6):591–9. https://doi.org/10.1200/JCO.2017.75.5215.
Ripperger T, Schlegelberger B. Acute lymphoblastic leukemia and lymphoma in the context of constitutional mismatch repair deficiency syndrome. Eur J Med Genet. 2016;59(3):133–42. https://doi.org/10.1016/j.ejmg.2015.12.014.
Oshrine B, Grana N, Moore C, Nguyen J, Crenshaw M, Edwards M, Sudhaman S, Forster VJ, Tabori U. B-cell acute lymphoblastic leukemia with high mutation burden presenting in a child with constitutional mismatch repair deficiency. Blood Adv. 2019;3(12):1795–8. https://doi.org/10.1182/bloodadvances.2019000358.
Duployez N, Jamrog LA, Fregona V, Hamelle C, Fenwarth L, Lejeune S, Helevaut N, Geffroy S, Caillault A, Marceau-Renaut A, Poulain S, Roche-Lestienne C, Largeaud L, Prade N, Dufrechou S, Hebrard S, Berthon C, Nelken B, Fernandes J, Villenet C, Figeac M, Gerby B, Delabesse E, Preudhomme C, Broccardo C. Germline PAX5 mutation predisposes to familial B-cell precursor acute lymphoblastic leukemia. Blood. 2021;137(10):1424–8. https://doi.org/10.1182/blood.2020005756.
Shah S, Schrader KA, Waanders E, Timms AE, Vijai J, Miething C, Wechsler J, Yang J, Hayes J, Klein RJ, Zhang J, Wei L, Wu G, Rusch M, Nagahawatte P, Ma J, Chen SC, Song G, Cheng J, Meyers P, Bhojwani D, Jhanwar S, Maslak P, Fleisher M, Littman J, Offit L, Rau-Murthy R, Fleischut MH, Corines M, Murali R, Gao X, Manschreck C, Kitzing T, Murty VV, Raimondi S, Kuiper RP, Simons A, Schiffman JD, Onel K, Plon SE, Wheeler D, Ritter D, Ziegler DS, Tucker K, Sutton R, Chenevix-Trench G, Li J, Huntsman DG, Hansford S, Senz J, Walsh T, Lee M, Hahn CN, Roberts K, King MC, Lo SM, Levine RL, Viale A, Socci ND, Nathanson KL, Scott HS, Daly M, Lipkin SM, Lowe SW, Downing JR, Altshuler D, Sandlund JT, Horwitz MS, Mullighan CG, Offit K. A recurrent germline PAX5 mutation confers susceptibility to pre-B cell acute lymphoblastic leukemia. Nat Genet. 2013;45(10):1226–31. https://doi.org/10.1038/ng.2754.
Moriyama T, Metzger ML, Wu G, Nishii R, Qian M, Devidas M, Yang W, Cheng C, Cao X, Quinn E, Raimondi S, Gastier-Foster JM, Raetz E, Larsen E, Martin PL, Bowman WP, Winick N, Komada Y, Wang S, Edmonson M, Xu H, Mardis E, Fulton R, Pui CH, Mullighan C, Evans WE, Zhang J, Hunger SP, Relling MV, Nichols KE, Loh ML, Yang JJ. Germline genetic variation in ETV6 and risk of childhood acute lymphoblastic leukaemia: a systematic genetic study. Lancet Oncol. 2015;16(16):1659–66. https://doi.org/10.1016/S1470-2045(15)00369-1.
Topka S, Vijai J, Walsh MF, Jacobs L, Maria A, Villano D, Gaddam P, Wu G, McGee RB, Quinn E, Inaba H, Hartford C, Pui CH, Pappo A, Edmonson M, Zhang MY, Stepensky P, Steinherz P, Schrader K, Lincoln A, Bussel J, Lipkin SM, Goldgur Y, Harit M, Stadler ZK, Mullighan C, Weintraub M, Shimamura A, Zhang J, Downing JR, Nichols KE, Offit K. Germline ETV6 mutations confer susceptibility to acute lymphoblastic leukemia and thrombocytopenia. PLoS Genet. 2015;11(6): e1005262. https://doi.org/10.1371/journal.pgen.1005262.
Greaves M. A causal mechanism for childhood acute lymphoblastic leukaemia. Nat Rev Cancer. 2018;18(8):471–84. https://doi.org/10.1038/s41568-018-0015-6.
Schafer D, Olsen M, Lahnemann D, Stanulla M, Slany R, Schmiegelow K, Borkhardt A, Fischer U. Five percent of healthy newborns have an ETV6-RUNX1 fusion as revealed by DNA-based GIPFEL screening. Blood. 2018;131(7):821–6. https://doi.org/10.1182/blood-2017-09-808402.
Mori H, Colman SM, Xiao Z, Ford AM, Healy LE, Donaldson C, Hows JM, Navarrete C, Greaves M. Chromosome translocations and covert leukemic clones are generated during normal fetal development. Proc Natl Acad Sci USA. 2002;99(12):8242–7. https://doi.org/10.1073/pnas.112218799.
Lausten-Thomsen U, Hjalgrim H, Marquart H, Lutterodt M, Petersen BL, Schmiegelow K. ETV6-RUNX1 transcript is not frequent in early human haematopoiesis. Eur J Haematol. 2008;81(2):161–2. https://doi.org/10.1111/j.1600-0609.2008.01091.x.
Lausten-Thomsen U, Madsen HO, Vestergaard TR, Hjalgrim H, Nersting J, Schmiegelow K. Prevalence of t(12;21)[ETV6-RUNX1]-positive cells in healthy neonates. Blood. 2011;117(1):186–9. https://doi.org/10.1182/blood-2010-05-282764.
Hein D, Dreisig K, Metzler M, Izraeli S, Schmiegelow K, Borkhardt A, Fischer U. The preleukemic TCF3-PBX1 gene fusion can be generated in utero and is present in approximately 0.6% of healthy newborns. Blood. 2019;134(16):1355–8. https://doi.org/10.1182/blood.2019002215.
Kroll ME, Stiller CA, Murphy MF, Carpenter LM. Childhood leukaemia and socioeconomic status in England and Wales 1976–2005: evidence of higher incidence in relatively affluent communities persists over time. Br J Cancer. 2011;105(11):1783–7. https://doi.org/10.1038/bjc.2011.415.
Francis SS, Selvin S, Yang W, Buffler PA, Wiemels JL. Unusual space-time patterning of the Fallon, Nevada leukemia cluster: evidence of an infectious etiology. Chem Biol Interact. 2012;196(3):102–9. https://doi.org/10.1016/j.cbi.2011.02.019.
Cazzaniga G, Bisanti L, Randi G, Deandrea S, Bungaro S, Pregliasco F, Perotti D, Spreafico F, Masera G, Valsecchi MG, Biondi A, Greaves M. Possible role of pandemic AH1N1 swine flu virus in a childhood leukemia cluster. Leukemia. 2017;31(8):1819–21. https://doi.org/10.1038/leu.2017.127.
Dockerty JD, Draper G, Vincent T, Rowan SD, Bunch KJ. Case-control study of parental age, parity and socioeconomic level in relation to childhood cancers. Int J Epidemiol. 2001;30(6):1428–37. https://doi.org/10.1093/ije/30.6.1428.
Greenbaum S, Sheiner E, Wainstock T, Segal I, Ben-Harush M, Sergienko R, Walfisch A. Cesarean delivery and childhood malignancies: a single-center, population-based cohort study. J Pediatr. 2018;197(292–296): e293. https://doi.org/10.1016/j.jpeds.2017.12.049.
Wang R, Wiemels JL, Metayer C, Morimoto L, Francis SS, Kadan-Lottick N, DeWan AT, Zhang Y, Ma X. Cesarean section and risk of childhood acute lymphoblastic leukemia in a population-based, record-linkage study in California. Am J Epidemiol. 2017;185(2):96–105. https://doi.org/10.1093/aje/kww153.
Marcotte EL, Thomopoulos TP, Infante-Rivard C, Clavel J, Petridou ET, Schuz J, Ezzat S, Dockerty JD, Metayer C, Magnani C, Scheurer ME, Mueller BA, Mora AM, Wesseling C, Skalkidou A, Rashed WM, Francis SS, Ajrouche R, Erdmann F, Orsi L, Spector LG. Caesarean delivery and risk of childhood leukaemia: a pooled analysis from the Childhood Leukemia International Consortium (CLIC). Lancet Haematol. 2016;3(4):e176-185. https://doi.org/10.1016/S2352-3026(16)00002-8.
Sevelsted A, Stokholm J, Bonnelykke K, Bisgaard H. Cesarean section and chronic immune disorders. Pediatrics. 2015;135(1):e92-98. https://doi.org/10.1542/peds.2014-0596.
Amitay EL, Keinan-Boker L. Breastfeeding and childhood leukemia incidence: a meta-analysis and systematic review. JAMA Pediatr. 2015;169(6): e151025. https://doi.org/10.1001/jamapediatrics.2015.1025.
Urayama KY, Buffler PA, Gallagher ER, Ayoob JM, Ma X. A meta-analysis of the association between day-care attendance and childhood acute lymphoblastic leukaemia. Int J Epidemiol. 2010;39(3):718–32. https://doi.org/10.1093/ije/dyp378.
Gilham C, Peto J, Simpson J, Roman E, Eden TO, Greaves MF, Alexander FE, Investigators U. Day care in infancy and risk of childhood acute lymphoblastic leukaemia: findings from UK case-control study. BMJ. 2005;330(7503):1294. https://doi.org/10.1136/bmj.38428.521042.8F.
Ma X, Buffler PA, Selvin S, Matthay KK, Wiencke JK, Wiemels JL, Reynolds P. Daycare attendance and risk of childhood acute lymphoblastic leukaemia. Br J Cancer. 2002;86(9):1419–24. https://doi.org/10.1038/sj.bjc.6600274.
Kamper-Jorgensen M, Woodward A, Wohlfahrt J, Benn CS, Simonsen J, Hjalgrim H, Schmiegelow K. Childcare in the first 2 years of life reduces the risk of childhood acute lymphoblastic leukemia. Leukemia. 2008;22(1):189–93. https://doi.org/10.1038/sj.leu.2404884.
Neglia JP, Linet MS, Shu XO, Severson RK, Potter JD, Mertens AC, Wen W, Kersey JH, Robison LL. Patterns of infection and day care utilization and risk of childhood acute lymphoblastic leukaemia. Br J Cancer. 2000;82(1):234–40. https://doi.org/10.1054/bjoc.1999.0905.
Rudant J, Lightfoot T, Urayama KY, Petridou E, Dockerty JD, Magnani C, Milne E, Spector LG, Ashton LJ, Dessypris N, Kang AY, Miller M, Rondelli R, Simpson J, Stiakaki E, Orsi L, Roman E, Metayer C, Infante-Rivard C, Clavel J. Childhood acute lymphoblastic leukemia and indicators of early immune stimulation: a Childhood Leukemia International Consortium study. Am J Epidemiol. 2015;181(8):549–62. https://doi.org/10.1093/aje/kwu298.
Orsi L, Magnani C, Petridou ET, Dockerty JD, Metayer C, Milne E, Bailey HD, Dessypris N, Kang AY, Wesseling C, Infante-Rivard C, Wunsch-Filho V, Mora AM, Spector LG, Clavel J. Living on a farm, contact with farm animals and pets, and childhood acute lymphoblastic leukemia: pooled and meta-analyses from the Childhood Leukemia International Consortium. Cancer Med. 2018;7(6):2665–81. https://doi.org/10.1002/cam4.1466.
Ajrouche R, Rudant J, Orsi L, Petit A, Baruchel A, Lambilliotte A, Gambart M, Michel G, Bertrand Y, Ducassou S, Gandemer V, Paillard C, Saumet L, Blin N, Hemon D, Clavel J. Childhood acute lymphoblastic leukaemia and indicators of early immune stimulation: the Estelle study (SFCE). Br J Cancer. 2015;112(6):1017–26. https://doi.org/10.1038/bjc.2015.53.
Hauer J, Martin-Lorenzo A, Sanchez-Garcia I. Infection causes childhood leukemia. Aging. 2015;7(9):607–8. https://doi.org/10.18632/aging.100815.
Linet MS, Brown LM, Mbulaiteye SM, Check D, Ostroumova E, Landgren A, Devesa SS. International long-term trends and recent patterns in the incidence of leukemias and lymphomas among children and adolescents ages 0–19 years. Int J Cancer. 2016;138(8):1862–74. https://doi.org/10.1002/ijc.29924.
Steliarova-Foucher E, Colombet M, Ries LAG, Moreno F, Dolya A, Bray F, Hesseling P, Shin HY, Stiller CA, IICC-3 Contributors. International incidence of childhood cancer, 2001–10: a population-based registry study. Lancet Oncol. 2017;18(6):719–31. https://doi.org/10.1016/S1470-2045(17)30186-9.
Steliarova-Foucher E, Fidler MM, Colombet M, Lacour B, Kaatsch P, Pineros M, Soerjomataram I, Bray F, Coebergh JW, Peris-Bonet R, Stiller CA, IICC-3 Contributors. Changing geographical patterns and trends in cancer incidence in children and adolescents in Europe, 1991–2010 (Automated Childhood Cancer Information System): a population-based study. Lancet Oncol. 2018;19(9):1159–69. https://doi.org/10.1016/S1470-2045(18)30423-6.
Morra ME, Kien ND, Elmaraezy A, Abdelaziz OAM, Elsayed AL, Halhouli O, Montasr AM, Vu TL, Ho C, Foly AS, Phi AP, Abdullah WM, Mikhail M, Milne E, Hirayama K, Huy NT. Early vaccination protects against childhood leukemia: a systematic review and meta-analysis. Sci Rep. 2017;7(1):15986. https://doi.org/10.1038/s41598-017-16067-0.
Spix C, Eletr D, Blettner M, Kaatsch P. Temporal trends in the incidence rate of childhood cancer in Germany 1987–2004. Int J Cancer. 2008;122(8):1859–67. https://doi.org/10.1002/ijc.23281.
Hauer J, Fischer U, Borkhardt A. Toward prevention of childhood ALL by early-life immune training. Blood. 2021;138(16):1412–28. https://doi.org/10.1182/blood.2020009895.
Netea MG, Joosten LA, Latz E, Mills KH, Natoli G, Stunnenberg HG, O’Neill LA, Xavier RJ. Trained immunity: a program of innate immune memory in health and disease. Science. 2016;352(6284):aaf1098. https://doi.org/10.1126/science.aaf1098.
de Laval B, Maurizio J, Kandalla PK, Brisou G, Simonnet L, Huber C, Gimenez G, Matcovitch-Natan O, Reinhardt S, David E, Mildner A, Leutz A, Nadel B, Bordi C, Amit I, Sarrazin S, Sieweke MH. C/EBPbeta-dependent epigenetic memory induces trained immunity in hematopoietic stem cells. Cell Stem Cell. 2020;26(5):793. https://doi.org/10.1016/j.stem.2020.03.014.
Kaufmann E, Sanz J, Dunn JL, Khan N, Mendonca LE, Pacis A, Tzelepis F, Pernet E, Dumaine A, Grenier JC, Mailhot-Leonard F, Ahmed E, Belle J, Besla R, Mazer B, King IL, Nijnik A, Robbins CS, Barreiro LB, Divangahi M. BCG educates hematopoietic stem cells to generate protective innate immunity against tuberculosis. Cell. 2018;172(1–2):176-190 e119. https://doi.org/10.1016/j.cell.2017.12.031.
Katzmarski N, Dominguez-Andres J, Cirovic B, Renieris G, Ciarlo E, Le Roy D, Lepikhov K, Kattler K, Gasparoni G, Handler K, Theis H, Beyer M, van der Meer JWM, Joosten LAB, Walter J, Schultze JL, Roger T, Giamarellos-Bourboulis EJ, Schlitzer A, Netea MG. Transmission of trained immunity and heterologous resistance to infections across generations. Nat Immunol. 2021;22(11):1382–90. https://doi.org/10.1038/s41590-021-01052-7.
Fidanza M, Seif AE, DeMicco A, Rolf N, Jo S, Yin B, Li Y, Barrett DM, Duque-Afonso J, Cleary ML, Bassing CH, Grupp SA, Reid GS. Inhibition of precursor B-cell malignancy progression by toll-like receptor ligand-induced immune responses. Leukemia. 2016;30(10):2116–9. https://doi.org/10.1038/leu.2016.152.
Seif AE, Barrett DM, Milone M, Brown VI, Grupp SA, Reid GS. Long-term protection from syngeneic acute lymphoblastic leukemia by CpG ODN-mediated stimulation of innate and adaptive immune responses. Blood. 2009;114(12):2459–66. https://doi.org/10.1182/blood-2009-02-203984.
Richardson RB. Promotional etiology for common childhood acute lymphoblastic leukemia: the infective lymphoid recovery hypothesis. Leuk Res. 2011;35(11):1425–31. https://doi.org/10.1016/j.leukres.2011.07.023.
Vilchis-Ordoñez A, Ramírez-Ramírez D, Pelayo R. The triad inflammation-microenvironment-tumor initiating cells in leukemia progression. Curr Opin Physio. 2021;19:211–8. https://doi.org/10.1016/j.cophys.2020.10.010.
Balandran JC, Davila-Velderrain J, Sandoval-Cabrera A, Zamora-Herrera G, Teran-Cerqueda V, Garcia-Stivalet LA, Limon-Flores JA, Armenta-Castro E, Rodriguez-Martinez A, Leon-Chavez BA, Vallejo-Ruiz V, Hassane DC, Perez-Tapia SM, Ortiz-Navarrete V, Guzman ML, Pelayo R. Patient-derived bone marrow spheroids reveal leukemia-initiating cells supported by mesenchymal hypoxic niches in pediatric B-ALL. Front Immunol. 2021;12: 746492. https://doi.org/10.3389/fimmu.2021.746492.
Beneforti L, Dander E, Bresolin S, Bueno C, Acunzo D, Bertagna M, Ford A, Gentner B, Kronnie GT, Vergani P, Menendez P, Biondi A, D’Amico G, Palmi C, Cazzaniga G. Pro-inflammatory cytokines favor the emergence of ETV6-RUNX1-positive pre-leukemic cells in a model of mesenchymal niche. Br J Haematol. 2020;190(2):262–73. https://doi.org/10.1111/bjh.16523.
Pastorczak A, Domka K, Fidyt K, Poprzeczko M, Firczuk M. Mechanisms of immune evasion in acute lymphoblastic leukemia. Cancers. 2021. https://doi.org/10.3390/cancers13071536.
Ribechini E, Greifenberg V, Sandwick S, Lutz MB. Subsets, expansion and activation of myeloid-derived suppressor cells. Med Microbiol Immunol. 2010;199(3):273–81. https://doi.org/10.1007/s00430-010-0151-4.
Bronte V, Brandau S, Chen SH, Colombo MP, Frey AB, Greten TF, Mandruzzato S, Murray PJ, Ochoa A, Ostrand-Rosenberg S, Rodriguez PC, Sica A, Umansky V, Vonderheide RH, Gabrilovich DI. Recommendations for myeloid-derived suppressor cell nomenclature and characterization standards. Nat Commun. 2016;7:12150. https://doi.org/10.1038/ncomms12150.
Romanski A, Bug G, Becker S, Kampfmann M, Seifried E, Hoelzer D, Ottmann OG, Tonn T. Mechanisms of resistance to natural killer cell-mediated cytotoxicity in acute lymphoblastic leukemia. Exp Hematol. 2005;33(3):344–52. https://doi.org/10.1016/j.exphem.2004.11.006.
Moretta L, Montaldo E, Vacca P, Del Zotto G, Moretta F, Merli P, Locatelli F, Mingari MC. Human natural killer cells: origin, receptors, function, and clinical applications. Int Arch Allergy Immunol. 2014;164(4):253–64. https://doi.org/10.1159/000365632.
Blaeschke F, Willier S, Stenger D, Lepenies M, Horstmann MA, Escherich G, Zimmermann M, Rojas Ringeling F, Canzar S, Kaeuferle T, Rohlfs M, Binder V, Klein C, Feuchtinger T. Leukemia-induced dysfunctional TIM-3(+)CD4(+) bone marrow T cells increase risk of relapse in pediatric B-precursor ALL patients. Leukemia. 2020;34(10):2607–20. https://doi.org/10.1038/s41375-020-0793-1.
Duell J, Dittrich M, Bedke T, Mueller T, Eisele F, Rosenwald A, Rasche L, Hartmann E, Dandekar T, Einsele H, Topp MS. Frequency of regulatory T cells determines the outcome of the T-cell-engaging antibody blinatumomab in patients with B-precursor ALL. Leukemia. 2017;31(10):2181–90. https://doi.org/10.1038/leu.2017.41.
An F, Wang H, Liu Z, Wu F, Zhang J, Tao Q, Li Y, Shen Y, Ruan Y, Zhang Q, Pan Y, Zhu W, Qin H, Wang Y, Fu Y, Feng Z, Zhai Z. Influence of patient characteristics on chimeric antigen receptor T cell therapy in B-cell acute lymphoblastic leukemia. Nat Commun. 2020;11(1):5928. https://doi.org/10.1038/s41467-020-19774-x.
Gomez AM, Martinez C, Gonzalez M, Luque A, Melen GJ, Martinez J, Hortelano S, Lassaletta A, Madero L, Ramirez M. Chemokines and relapses in childhood acute lymphoblastic leukemia: a role in migration and in resistance to antileukemic drugs. Blood Cells Mol Dis. 2015;55(3):220–7. https://doi.org/10.1016/j.bcmd.2015.07.001.
Witkowski MT, Dolgalev I, Evensen NA, Ma C, Chambers T, Roberts KG, Sreeram S, Dai Y, Tikhonova AN, Lasry A, Qu C, Pei D, Cheng C, Robbins GA, Pierro J, Selvaraj S, Mezzano V, Daves M, Lupo PJ, Scheurer ME, Loomis CA, Mullighan CG, Chen W, Rabin KR, Tsirigos A, Carroll WL, Aifantis I. Extensive remodeling of the immune microenvironment in B cell acute lymphoblastic leukemia. Cancer Cell. 2020;37(6):867-882 e812. https://doi.org/10.1016/j.ccell.2020.04.015.
Roy A. A “gut feeling” about precursor B-ALL. Blood. 2020;136(18):1995–6. https://doi.org/10.1182/blood.2020008907.
Vicente-Duenas C, Janssen S, Oldenburg M, Auer F, Gonzalez-Herrero I, Casado-Garcia A, Isidro-Hernandez M, Raboso-Gallego J, Westhoff P, Pandyra AA, Hein D, Gossling KL, Alonso-Lopez D, De Las RJ, Bhatia S, Garcia-Criado FJ, Garcia-Cenador MB, Weber APM, Kohrer K, Hauer J, Fischer U, Sanchez-Garcia I, Borkhardt A. An intact gut microbiome protects genetically predisposed mice against leukemia. Blood. 2020;136(18):2003–17. https://doi.org/10.1182/blood.2019004381.
Koenig JE, Spor A, Scalfone N, Fricker AD, Stombaugh J, Knight R, Angenent LT, Ley RE. Succession of microbial consortia in the developing infant gut microbiome. Proc Natl Acad Sci USA. 2011;108(Suppl 1):4578–85. https://doi.org/10.1073/pnas.1000081107.
Palmer C, Bik EM, DiGiulio DB, Relman DA, Brown PO. Development of the human infant intestinal microbiota. PLoS Biol. 2007;5(7): e177. https://doi.org/10.1371/journal.pbio.0050177.
Yatsunenko T, Rey FE, Manary MJ, Trehan I, Dominguez-Bello MG, Contreras M, Magris M, Hidalgo G, Baldassano RN, Anokhin AP, Heath AC, Warner B, Reeder J, Kuczynski J, Caporaso JG, Lozupone CA, Lauber C, Clemente JC, Knights D, Knight R, Gordon JI. Human gut microbiome viewed across age and geography. Nature. 2012;486(7402):222–7. https://doi.org/10.1038/nature11053.
Bindels LB, Porporato P, Dewulf EM, Verrax J, Neyrinck AM, Martin JC, Scott KP, Buc Calderon P, Feron O, Muccioli GG, Sonveaux P, Cani PD, Delzenne NM. Gut microbiota-derived propionate reduces cancer cell proliferation in the liver. Br J Cancer. 2012;107(8):1337–44. https://doi.org/10.1038/bjc.2012.409.
Macia L, Tan J, Vieira AT, Leach K, Stanley D, Luong S, Maruya M, Ian McKenzie C, Hijikata A, Wong C, Binge L, Thorburn AN, Chevalier N, Ang C, Marino E, Robert R, Offermanns S, Teixeira MM, Moore RJ, Flavell RA, Fagarasan S, Mackay CR. Metabolite-sensing receptors GPR43 and GPR109A facilitate dietary fibre-induced gut homeostasis through regulation of the inflammasome. Nat Commun. 2015;6:6734. https://doi.org/10.1038/ncomms7734.
Kim CH, Park J, Kim M. Gut microbiota-derived short-chain fatty acids, T cells, and inflammation. Immune Netw. 2014;14(6):277–88. https://doi.org/10.4110/in.2014.14.6.277.
Trompette A, Gollwitzer ES, Yadava K, Sichelstiel AK, Sprenger N, Ngom-Bru C, Blanchard C, Junt T, Nicod LP, Harris NL, Marsland BJ. Gut microbiota metabolism of dietary fiber influences allergic airway disease and hematopoiesis. Nat Med. 2014;20(2):159–66. https://doi.org/10.1038/nm.3444.
Vrieze A, Out C, Fuentes S, Jonker L, Reuling I, Kootte RS, van Nood E, Holleman F, Knaapen M, Romijn JA, Soeters MR, Blaak EE, Dallinga-Thie GM, Reijnders D, Ackermans MT, Serlie MJ, Knop FK, Holst JJ, van der Ley C, Kema IP, Zoetendal EG, de Vos WM, Hoekstra JB, Stroes ES, Groen AK, Nieuwdorp M. Impact of oral vancomycin on gut microbiota, bile acid metabolism, and insulin sensitivity. J Hepatol. 2014;60(4):824–31. https://doi.org/10.1016/j.jhep.2013.11.034.
Macfarlane GT, Macfarlane S. Fermentation in the human large intestine: its physiologic consequences and the potential contribution of prebiotics. J Clin Gastroenterol. 2011;45(Suppl):S120-127. https://doi.org/10.1097/MCG.0b013e31822fecfe.
Francino MP. Antibiotics and the human gut microbiome: dysbioses and accumulation of resistances. Front Microbiol. 2015;6:1543. https://doi.org/10.3389/fmicb.2015.01543.
Honda K, Littman DR. The microbiota in adaptive immune homeostasis and disease. Nature. 2016;535(7610):75–84. https://doi.org/10.1038/nature18848.
Levy M, Kolodziejczyk AA, Thaiss CA, Elinav E. Dysbiosis and the immune system. Nat Rev Immunol. 2017;17(4):219–32. https://doi.org/10.1038/nri.2017.7.
Lange K, Buerger M, Stallmach A, Bruns T. Effects of antibiotics on gut microbiota. Dig Dis. 2016;34(3):260–8. https://doi.org/10.1159/000443360.
Ubeda C, Pamer EG. Antibiotics, microbiota, and immune defense. Trends Immunol. 2012;33(9):459–66. https://doi.org/10.1016/j.it.2012.05.003.
Nagai Y, Garrett KP, Ohta S, Bahrun U, Kouro T, Akira S, Takatsu K, Kincade PW. Toll-like receptors on hematopoietic progenitor cells stimulate innate immune system replenishment. Immunity. 2006;24(6):801–12. https://doi.org/10.1016/j.immuni.2006.04.008.
Esplin BL, Shimazu T, Welner RS, Garrett KP, Nie L, Zhang Q, Humphrey MB, Yang Q, Borghesi LA, Kincade PW. Chronic exposure to a TLR ligand injures hematopoietic stem cells. J Immunol. 2011;186(9):5367–75. https://doi.org/10.4049/jimmunol.1003438.
Zhang D, Chen G, Manwani D, Mortha A, Xu C, Faith JJ, Burk RD, Kunisaki Y, Jang JE, Scheiermann C, Merad M, Frenette PS. Neutrophil ageing is regulated by the microbiome. Nature. 2015;525(7570):528–32. https://doi.org/10.1038/nature15367.
Azad MB, Konya T, Persaud RR, Guttman DS, Chari RS, Field CJ, Sears MR, Mandhane PJ, Turvey SE, Subbarao P, Becker AB, Scott JA, Kozyrskyj AL, Investigators CS. Impact of maternal intrapartum antibiotics, method of birth and breastfeeding on gut microbiota during the first year of life: a prospective cohort study. BJOG. 2016;123(6):983–93. https://doi.org/10.1111/1471-0528.13601.
Yassour M, Vatanen T, Siljander H, Hamalainen AM, Harkonen T, Ryhanen SJ, Franzosa EA, Vlamakis H, Huttenhower C, Gevers D, Lander ES, Knip M, Group DS, Xavier RJ. Natural history of the infant gut microbiome and impact of antibiotic treatment on bacterial strain diversity and stability. Sci Transl Med. 2016;8(343):343ra381. https://doi.org/10.1126/scitranslmed.aad0917.
Chai G, Governale L, McMahon AW, Trinidad JP, Staffa J, Murphy D. Trends of outpatient prescription drug utilization in US children, 2002–2010. Pediatrics. 2012;130(1):23–31. https://doi.org/10.1542/peds.2011-2879.
Wald ER, Applegate KE, Bordley C, Darrow DH, Glode MP, Marcy SM, Nelson CE, Rosenfeld RM, Shaikh N, Smith MJ, Williams PV, Weinberg ST, American Academy of P. Clinical practice guideline for the diagnosis and management of acute bacterial sinusitis in children aged 1–18 years. Pediatrics. 2013;132(1):e262-280. https://doi.org/10.1542/peds.2013-1071.
Lieberthal AS, Carroll AE, Chonmaitree T, Ganiats TG, Hoberman A, Jackson MA, Joffe MD, Miller DT, Rosenfeld RM, Sevilla XD, Schwartz RH, Thomas PA, Tunkel DE. The diagnosis and management of acute otitis media. Pediatrics. 2013;131(3):e964-999. https://doi.org/10.1542/peds.2012-3488.
Coker TR, Chan LS, Newberry SJ, Limbos MA, Suttorp MJ, Shekelle PG, Takata GS. Diagnosis, microbial epidemiology, and antibiotic treatment of acute otitis media in children: a systematic review. JAMA. 2010;304(19):2161–9. https://doi.org/10.1001/jama.2010.1651.
Ralston SL, Lieberthal AS, Meissner HC, Alverson BK, Baley JE, Gadomski AM, Johnson DW, Light MJ, Maraqa NF, Mendonca EA, Phelan KJ, Zorc JJ, Stanko-Lopp D, Brown MA, Nathanson I, Rosenblum E, Sayles S 3rd, Hernandez-Cancio S, American Academy of P. Clinical practice guideline: the diagnosis, management, and prevention of bronchiolitis. Pediatrics. 2014;134(5):e1474-1502. https://doi.org/10.1542/peds.2014-2742.
Shulman ST, Bisno AL, Clegg HW, Gerber MA, Kaplan EL, Lee G, Martin JM, Van Beneden C, Infectious Diseases Society of A. Clinical practice guideline for the diagnosis and management of group A streptococcal pharyngitis: 2012 update by the Infectious Diseases Society of America. Clin Infect Dis. 2012;55(10):e86-102. https://doi.org/10.1093/cid/cis629.
Fashner J, Ericson K, Werner S. Treatment of the common cold in children and adults. Am Fam Physician. 2012;86(2):153–9.
Hersh AL, Jackson MA, Hicks LA, American Academy of Pediatrics Committee on Infectious D. Principles of judicious antibiotic prescribing for upper respiratory tract infections in pediatrics. Pediatrics. 2013;132(6):1146–54. https://doi.org/10.1542/peds.2013-3260.
Subcommittee on Urinary Tract Infection SCoQI, Management, Roberts KB. Urinary tract infection: clinical practice guideline for the diagnosis and management of the initial UTI in febrile infants and children 2–24 months. Pediatrics. 2011;128(3):595–610. https://doi.org/10.1542/peds.2011-1330.
White B. Diagnosis and treatment of urinary tract infections in children. Am Fam Physician. 2011;83(4):409–15.
McDonnell L, Gilkes A, Ashworth M, Rowland V, Harries TH, Armstrong D, White P. Association between antibiotics and gut microbiome dysbiosis in children: systematic review and meta-analysis. Gut Microbes. 2021;13(1):1–18. https://doi.org/10.1080/19490976.2020.1870402.
Fredricks DN. Evidence in microbiome science: standards for the field (and laboratory). Clin Infect Dis. 2021;72(9):1514–6. https://doi.org/10.1093/cid/ciaa766.
Gao X, Miao R, Zhu Y, Lin C, Yang X, Jia R, Linghan K, Wan C, Deng J. A new insight into acute lymphoblastic leukemia in children: influences of changed intestinal microfloras. BMC Pediatr. 2020;20(1):290. https://doi.org/10.1186/s12887-020-02192-9.
Liu X, Zou Y, Ruan M, Chang L, Chen X, Wang S, Yang W, Zhang L, Guo Y, Chen Y, Zhang Y, He H, Gan Y, Wang K, Zhu X. Pediatric acute lymphoblastic leukemia patients exhibit distinctive alterations in the gut microbiota. Front Cell Infect Microbiol. 2020;10: 558799. https://doi.org/10.3389/fcimb.2020.558799.
De Pietri S, Ingham AC, Frandsen TL, Rathe M, Krych L, Castro-Mejia JL, Nielsen DS, Nersting J, Wehner PS, Schmiegelow K, Hasle H, Pamp SJ, Muller K. Gastrointestinal toxicity during induction treatment for childhood acute lymphoblastic leukemia: the impact of the gut microbiota. Int J Cancer. 2020;147(7):1953–62. https://doi.org/10.1002/ijc.32942.
Rajagopala SV, Singh H, Yu Y, Zabokrtsky KB, Torralba MG, Moncera KJ, Frank B, Pieper R, Sender L, Nelson KE. Persistent gut microbial dysbiosis in children with acute lymphoblastic leukemia (ALL) during chemotherapy. Microb Ecol. 2020;79(4):1034–43. https://doi.org/10.1007/s00248-019-01448-x.
Rajagopala SV, Yooseph S, Harkins DM, Moncera KJ, Zabokrtsky KB, Torralba MG, Tovchigrechko A, Highlander SK, Pieper R, Sender L, Nelson KE. Gastrointestinal microbial populations can distinguish pediatric and adolescent acute lymphoblastic leukemia (ALL) at the time of disease diagnosis. BMC Genomics. 2016;17(1):635. https://doi.org/10.1186/s12864-016-2965-y.
Chua LL, Rajasuriar R, Lim YAL, Woo YL, Loke P, Ariffin H. Temporal changes in gut microbiota profile in children with acute lymphoblastic leukemia prior to commencement-, during-, and post-cessation of chemotherapy. BMC Cancer. 2020;20(1):151. https://doi.org/10.1186/s12885-020-6654-5.
Wang Y, Xue J, Zhou X, You M, Du Q, Yang X, He J, Zou J, Cheng L, Li M, Li Y, Zhu Y, Li J, Shi W, Xu X. Oral microbiota distinguishes acute lymphoblastic leukemia pediatric hosts from healthy populations. PLoS ONE. 2014;9(7): e102116. https://doi.org/10.1371/journal.pone.0102116.
Oldenburg M, Ruchel N, Janssen S, Borkhardt A, Gossling KL. The microbiome in childhood acute lymphoblastic leukemia. Cancers. 2021. https://doi.org/10.3390/cancers13194947.
Funding
Open Access funding enabled and organized by Projekt DEAL. This review was not funded.
Author information
Authors and Affiliations
Contributions
TMC-S wrote the article, AS and NG made substantial contributions to the conception and design of the work. All auhtors reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This article does not contain any studies with human participants or animals performed by any of the authors.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cardesa-Salzmann, T.M., Simon, A. & Graf, N. Antibiotics in early life and childhood pre-B-ALL. Reasons to analyze a possible new piece in the puzzle. Discov Onc 13, 5 (2022). https://doi.org/10.1007/s12672-022-00465-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-022-00465-6