Abstract
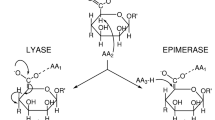
Alginates are heteropolysaccharides comprising β-D-mannuronate (M) and α-L-guluronate (G) units. Natural sources of alginate include brown algae and certain bacteria. Brown algal alginates possess a greater degree of sequence diversity than do those produced by bacteria, and this may be due to the presence of multiple mannuronan C5-epimerase (MC5E) isozymes in brown algae, although only three have been enzymatically studied to date. In this study, a novel MC5E candidate protein, CoC5-1, from the brown alga Cladosiphon okamuranus, for which no enzymatic function has been previously determined, was expressed as a glycosylated protein in insect cells. The action of recombinant CoC5-1 on polymannuronate (polyM) increased the amount of gel formed in the presence of Ca2+, indicating that CoC5-1 possesses MC5E activity. The optimum temperature, pH, and NaCl concentration were 40 °C, pH 8.0, and 100 mM, respectively. The thermal stability of CoC5-1 was higher than that of the Saccharina japonica MC5E SjC5-VI. CoC5-1 remained active after incubation at 50 °C for 1 h, whereas SjC5-VI was completely inactivated. 1H-nuclear magnetic resonance (NMR) analysis revealed that the M:G ratio of the polyM substrate was altered from 9.0 to 2.1 by CoC5-1. This is the first report of MC5E enzymatic activity in brown algae of the family Chordariaceae.







Similar content being viewed by others
References
Astbury WT (1945) Structure of alginic acid. Nature 155:667–668. https://doi.org/10.1038/155667a0
Black WAP (1950) The seasonal variation in weight and chemical composition of the common British Laminariaceae. J Mar Biol Ass 29:45–72. https://doi.org/10.1017/S0025315400056186
Blatny JM, Ertesvåg H, Nes IF, Valla S (2003) Heterologous gene expression in Lactococcus lactis; expression of the Azotobacter vinelandii algE6 gene product displaying mannuronan C-5 epimerase activity. FEMS Microbiol Lett 227:229–235. https://doi.org/10.1016/S0378-1097(03)00685-2
Costerton JW, Lewandowski Z (1995) Microbial Biofilms. Annu Rev Microbial 49:711–745. https://doi.org/10.1146/annurev.mi.49.100195.003431
Craigie JS, Morris ER, Rees DA, Thom D (1984) Alginate block structure in phaeophyceae from Nova Scotia: Variation with species, environment and tissue-type. Carbohydr Polym 4:237–252. https://doi.org/10.1016/0144-8617(84)90001-8
Donati I, Paoletti S (2009) Material Properties of Alginates. In: Rehm BHA (ed) Alginates: Biology and Applications. Springer, Berlin Heidelberg, Berlin, Heidelberg, pp 1–53
Ertesvåg H, Doseth B, Larsen B, Skjåk-Bræk G, Valla S (1994) Cloning and expression of an Azotobacter vinelandii mannuronan C-5-epimerase gene. J Bacteriol 176:2846–2853. https://doi.org/10.1128/jb.176.10.2846-2853.1994
Ertesvåg H, Hoidal HK, Skjåk-Bræk G, Valla S (1998) The Azotobacter vinelandii mannuronan C-5-epimerase AlgE1 consists of two separate catalytic domains. J Biol Chem 273:30927–30932. https://doi.org/10.1074/jbc.273.47.30927
Fischer FG, Dörfel H (1955) Die polyuronsäuren der braunalgen (Kohlenhydrate der Algen I). Biol Chem 302:186–203. https://doi.org/10.1515/bchm2.1955.302.1-2.186
Fischl R, Bertelsen K, Gaillard F, Coelho S, Michel G, Klinger M, Boyen C, Czjzek M, Hervé C (2016) The cell-wall active mannuronan C5-epimerases in the model brown alga Ectocarpus: From gene context to recombinant protein. Glycobiology 26:973–983. https://doi.org/10.1093/glycob/cww040
Franklin MJ, Chitnis CE, Gacesa P, Sonesson A, White DC, Ohman DE (1994) Pseudomonas aeruginosa AlgG is a polymer level alginate C5-mannuronan epimerase. J Bacteriol 176:1821–1830. https://doi.org/10.1128/jb.176.7.1821-1830.1994
Fukumoto R, Borlongan IA, Nishihara GN, Endo H, Terada R (2018) Photosynthetic responses to photosynthetically active radiation and temperature including chilling-light stress on the heteromorphic life history stages of a brown alga, Cladosiphon okamuranus (Chordariaceae) from Ryukyu Islands, Japan. Phycol Res 66:209–217. https://doi.org/10.1111/pre.12220
Gaardløs M, Heggeset TMB, Tøndervik A, Teze D, Svensson B, Ertesvåg H, Sletta H, Aachmann FL (2022) Mechanistic basis for understanding the dual activities of the bifunctional Azotobacter vinelandii mannuronan C-5-Epimerase and alginate lyase AlgE7. Appl Environ Microbiol 88:e0183621. https://doi.org/10.1128/AEM.01836-21
Gacesa P, Wusteman FS (1990) Plate assay for simultaneous detection of alginate lyases and determination of substrate specificity. Appl Environ Microbiol 56:2265–2267. https://doi.org/10.1128/aem.56.7.2265-2267.1990
Gawin A, Tietze L, Aarstad OA, Aachmann FL, Brautaset T, Ertesvåg H (2020) Functional characterization of three Azotobacter chroococcum alginate-modifying enzymes related to the Azotobacter vinelandii AlgE mannuronan C-5-epimerase family. Sci Rep 10:12470–12514. https://doi.org/10.1038/s41598-020-68789-3
Gimmestad M, Sletta H, Ertesvåg H, Bakkevig K, Jain S, Suh S-J, Skjåk-Bræk G, Ellingsen TE, Ohman DE, Valla S (2003) The Pseudomonas fluorescens AlgG protein, but not its mannuronan C-5-epimerase activity, is needed for alginate polymer formation. J Bacteriol 185:3515–3523. https://doi.org/10.1128/JB.185.12.3515-3523.2003
Grasdalen H (1983) High-field, 1H-n.m.r. spectroscopy of alginate: sequential structure and linkage conformations. Carbohydr Res 118:255–260. https://doi.org/10.1016/0008-6215(83)88053-7
Grasdalen H, Larsen B, Smidsrød O (1977) 13C-N.m.r. studies of alginate. Carbohydr Res 56:C11–C15. https://doi.org/10.1016/S0008-6215(00)83369-8
Grasdalen H, Larsen B, Smidsrød O (1979) A p.m.r. study of the composition and sequence of uronate residues in alginates. Carbohydr Res 68:23–31. https://doi.org/10.1016/S0008-6215(00)84051-3
Grasdalen H, Larsen B, Smisrød O (1981) 13C-n.m.r. studies of monomeric composition and sequence in alginate. Carbohydr Res 89:179–191. https://doi.org/10.1016/S0008-6215(00)85243-X
Harrison RL, Jarvis DL (2006) Protein N-glycosylation in the baculovirus-insect cell expression system and engineering of insect cells to produce “mammalianized” recombinant glycoproteins. Adv Virus Res 68:159–191. https://doi.org/10.1016/S0065-3527(06)68005-6
Haug A, Smidsrød O (1967) Strontium-calcium selectivity of alginates. Nature 215:757–757. https://doi.org/10.1038/215757a0
Haug A, Smidsrød O, Högdahl B, Øye HA, Rasmussen SE, Sunde E, Sørensen NA (1970) Selectivity of some anionic polymers for divalent metal ions. Acta Chem Scand 24:843–854. https://doi.org/10.3891/acta.chem.scand.24-0843
Hay ID, Rehman ZU, Moradali MF, Wang Y, Rehm BHA (2013) Microbial alginate production, modification and its applications. Microb Biotechnol 6:637–650. https://doi.org/10.1111/1751-7915.12076
Hirokawa T, Boon-Chieng S, Mitaku S (1998) SOSUI: classification and secondary structure prediction system for membrane proteins. Bioinformatics 14:378–379. https://doi.org/10.1093/bioinformatics/14.4.378
Hirst EL, Jones JKN, Jones WO (1939) Structure of alginic acid. Nature 143:857. https://doi.org/10.1038/143857a0
Hoidal HK, Ertesvåg H, Skjåk-Bræk G, Stokke BT, Valla S (1999) The recombinant Azotobacter vinelandii mannuronan C-5-epimerase AlgE4 epimerizes alginate by a nonrandom attack mechanism. J Biol Chem 274:12316–12322. https://doi.org/10.1074/jbc.274.18.12316
Inoue A (2018) Characterization of PL-7 family alginate lyases from marine organisms and their applications. Methods Enzymol 605:499–524. https://doi.org/10.1016/bs.mie.2018.01.030
Inoue A, Ojima T (2019) Functional identification of alginate lyase from the brown alga Saccharina japonica. Sci Rep 9:4937. https://doi.org/10.1038/s41598-019-41351-6
Inoue A, Ojima T (2021) Functional identification of the 4-deoxy-L-erythro-5-hexoseulose uronate reductase from a brown alga, Saccharina japonica. Biochem Biophys Res Commun 545:112–118. https://doi.org/10.1016/j.bbrc.2021.01.090
Inoue A, Satoh A, Morishita M, Tokunaga Y, Miyakawa T, Tanokura M, Ojima T (2016) Functional heterologous expression and characterization of mannuronan C5-epimerase from the brown alga Saccharina japonica. Algal Res 16:282–291. https://doi.org/10.1016/j.algal.2016.03.030
Inoue A, Iwayama T, Ojima T (2019) Complementary DNA cloning and functional analysis of lycopene β-cyclase in the brown alga Undaria pinnatifida. Fisheries Sci 85:717–729. https://doi.org/10.1007/s12562-019-01314-2
Kelly BJ, Brown MT (2000) Variations in the alginate content and composition of Durvillaea antarctica and D. willana from southern New Zealand. pp 317–324
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. https://doi.org/10.1038/227680a0
Liu F, Sun X, Wang W, Liang Z, Wang F (2013) De novo transcriptome analysis-gained insights into physiological and metabolic characteristics of Sargassum thunbergii (Fucales, Phaeophyceae). J Appl Phycol 26:1519–1526. https://doi.org/10.1007/s10811-013-0140-2
Michel G, Tonon T, Scornet D, Cock JM, Kloareg B (2010) The cell wall polysaccharide metabolism of the brown alga Ectocarpus siliculosus. Insights into the evolution of extracellular matrix polysaccharides in eukaryotes. New Phytol 188:82–97. https://doi.org/10.1111/j.1469-8137.2010.03374.x
Mitaku S, Hirokawa T (1999) Physicochemical factors for discriminating between soluble and membrane proteins: hydrophobicity of helical segments and protein length. Protein Eng 12:953–957. https://doi.org/10.1093/protein/12.11.953
Mitaku S, Hirokawa T, Tsuji T (2002) Amphiphilicity index of polar amino acids as an aid in the characterization of amino acid preference at membrane-water interfaces. Bioinformatics 18:608–616. https://doi.org/10.1093/bioinformatics/18.4.608
Moradali MF, Donati I, Sims IM, Ghods S, Rehm BHA (2015) Alginate polymerization and modification are linked in Pseudomonas aeruginosa. Mbio 6:e00453-e515. https://doi.org/10.1128/mBio.00453-15
Moradali MF, Ghods S, Rehm BHA (2018) Alginate Biosynthesis and Biotechnological Production. In: Rehm BHA, Moradali MF (eds) Alginates and their biomedical applications. Springer Singapore, Singapore
Mørch YA, Donati I, Strand BL, Skjåk-Bræk G (2006) Effect of Ca2+, Ba2+, and Sr2+ on alginate microbeads. Biomacromol 7:1471–1480. https://doi.org/10.1021/bm060010d
Nishitsuji K, Arimoto A, Iwai K, Sudo Y, Hisata K, Fujie M, Arakaki N, Kushiro T, Konishi T, Shinzato C, Satoh N, Shoguchi E (2016) A draft genome of the brown alga, Cladosiphon okamuranus, S-strain: a platform for future studies of “mozuku” biology. DNA Res 23:561–570. https://doi.org/10.1093/dnares/dsw039
Nyvall P, Corre E, Boisset C, Barbeyron T, Rousvoal S, Scornet D, Kloareg B, Boyen C (2003) Characterization of mannuronan C-5-epimerase genes from the brown alga Laminaria digitata. Plant Physiol 133:726–735. https://doi.org/10.1104/pp.103.025981
Pengyan Z, Chang L, Zhanru S, Fuli L, Jianting Y, Delin D (2021) Genome-wide transcriptome profiling and characterization of mannuronan C5-epimerases in Saccharina japonica. Algal Res. https://doi.org/10.1016/j.algal.2021.102491
Ramstad MV, Ellingsen TE, Josefsen KD, Hoidal HK, Valla S, Skjåk-Bræk G, Levine DW (1999) Properties and action pattern of the recombinant mannuronan C-5-epimerase AlgE2. Enzyme Microb Technol 24:636–646. https://doi.org/10.1016/S0141-0229(98)00148-3
Shan TF, Pang SJ, Li J, Li X (2014) De novo transcriptome analysis of the gametophyte of Undaria pinnatifida (Phaeophyceae). J Appl Phycol 27:1011–1019. https://doi.org/10.1007/s10811-014-0393-4
Skjåk-Bræk G, Paoletti S, Gianferrara T (1989) Selective acetylation of mannuronic acid residues in calcium alginate gels. Carbohydr Res 185:119–129. https://doi.org/10.1016/0008-6215(89)84027-3
Smidsrød O (1974) Molecular basis for some physical properties of alginates in the gel state. Farad Discuss 57:263–274. https://doi.org/10.1039/DC9745700263
Sun M, Sun C, Li T, Li K, Yan S, Yin H (2020) Characterization of a novel bifunctional mannuronan C-5 epimerase and alginate lyase from Pseudomonas mendocina. sp. DICP-70. Int J Biol Macromol 150:662–670. https://doi.org/10.1016/j.ijbiomac.2020.02.126
Svanem BI, Skjåk-Bræk G, Ertesvåg H, Valla S (1999) Cloning and expression of three new Azotobacter vinelandii genes closely related to a previously described gene family encoding mannuronan C-5-epimerases. J Bacteriol 181:68–77. https://doi.org/10.1128/JB.181.1.68-77.1999
Svanem BI, Strand WI, Ertesvåg H, Skjåk-Bræk G, Hartmann M, Barbeyron T, Valla S (2001) The catalytic activities of the bifunctional Azotobacter vinelandii mannuronan C-5-epimerase and alginate lyase AlgE7 probably originate from the same active site in the enzyme. J Biol Chem 276:31542–31550. https://doi.org/10.1074/jbc.M102562200
Tako M, Yoza E, Tohma S (2000) Chemical characterization of acetyl fucoidan and alginate from commercially cultured Cladosiphon okamuranus. Bot Mar 43:393–398
Teufel F, Armenteros JJA, Johansen AR, Gíslason MH, Pihl SI, Tsirigos KD, Winther O, Brunak S, von Heijne G, Nielsen H (2022) SignalP 6.0 predicts all five types of signal peptides using protein language models. Nat Biotechnol 40:1023–1025. https://doi.org/10.1038/s41587-021-01156-3
Tomoda M (1963) Colorimetric Determination of Pentoses. IV. Determination with Orcinol Reagent. Chem Pharm Bull 11:809–812. https://doi.org/10.1248/cpb.11.809
Tøndervik A, Aune R, Degelmann A, Piontek M, Ertesvåg H, Skjåk-Bræk G, Sletta H (2022) Strain construction and process development for efficient recombinant production of mannuronan C-5 epimerases in Hansenula polymorpha. Front Plant Sci 13:837891. https://doi.org/10.3389/fpls.2022.837891
Tsujino I, Saito T (1961) A new unsaturated uronide isolated from alginase hydrolysate. Nature 192:970–971. https://doi.org/10.1038/192970a0
Windhues T, Borchard W (2003) Effect of acetylation on physico-chemical properties of bacterial and algal alginates in physiological sodium chloride solutions investigated with light scattering techniques. Carbohydr Polym 52:47–52. https://doi.org/10.1016/S0144-8617(02)00265-5
Ye N, Zhang X, Miao M, Fan X, Zheng Y, Xu D, Wang J, Zhou L, Wang D, Gao Y, Wang Y, Shi W, Ji P, Li D, Guan Z, Shao C, Zhuang Z, Gao Z, Qi J, Zhao F (2015) Saccharina genomes provide novel insight into kelp biology. Nat Commun 6:6986. https://doi.org/10.1038/ncomms7986
Acknowledgements
This study was supported by the Japan Society for the Promotion of Science KAKENHI program (grant numbers 16H04977 and 19H03039), the Center of Innovation Program of the Japan Science and Technology Agency (grant number JPMJCE1301), the Center of Innovation NEXT Program of the Japan Science and Technology Agency (grant number JPMJPF2108), and the Hokkaido University NMR Facility as a program of the “NMR Platform” of the Ministry of Education, Culture, Sports, Science and Technology (MEXT).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no potential conflicts of interest regarding the research, authorship, or publication of this article.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sakagami, M., Ohnishi, Y., Kumaki, Y. et al. Enzymatic characterization of a mannuronan C5-epimerase from the subtropical brown alga Cladosiphon okamuranus. Fish Sci 89, 823–835 (2023). https://doi.org/10.1007/s12562-023-01720-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12562-023-01720-7