Abstract
Hepatitis E virus (HEV) is an emerging zoonotic pathogen that exhibits great host diversity. The primary means of transmission of the virus in low- and middle-income countries is contaminated water, often due to a lack of access to proper sanitation, which leads to faecal contamination of water sources. Environmental surveillance is an important tool that can be used to monitor virus circulation and as an early warning system for outbreaks. This study was conducted to determine the prevalence and genetic diversity of HEV in wastewater, surface water (rivers and standpipe/ablution water), and effluent from a piggery in South Africa. A total of 536 water samples were screened for HEV using real-time reverse transcription-polymerase chain reaction. Overall, 21.8% (117/536) of the wastewater, river, and ablution water samples tested positive for HEV, whereas 74.4% (29/39) of the samples from the piggery tested positive. Genotyping revealed sequences belonging to HEV genotypes 3 (98%, 53/54) and 4 (2%, 1/54), with subtypes 3c, 3f, and 4b being identified.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatitis E virus (HEV) is a pathogen of global importance as it is the major cause of acute viral hepatitis (Hoofnagle et al., 2012; Sridhar et al., 2017), surpassing hepatitis A virus (HAV) (Rein et al., 2012; Wong et al., 2021). The World Health Organization (WHO) estimates that HEV causes approximately 20 million infections, more than 44,000 deaths, and 3,000 stillbirths each year (Baez et al., 2017; Rein et al., 2012; Wong et al., 2021). Hepatitis E virus infection is typically self-limiting and presents with comparable clinical manifestations to HAV, such as fever, discomfort, body aches, vomiting, jaundice, and nausea. However, its primary distinguishing characteristic is its increased morbidity and case fatality rate in young adults and pregnant women (Kmush et al., 2013).
Hepatitis E virus belongs to the Hepeviridae family and is classified within the genus Paslahepevirus, the species Paslahepevirus balayani (Purdy et al., 2022). Eight HEV genotypes (HEV-1 – HEV-8) have been described to date (Smith et al., 2020). Genotypes 1 and 2 are exclusively associated with human infections through faecal-oral transmission. Genotypes 3 and 4 are zoonotic in nature and can cause chronic infection in the immunocompromised (Aslan & Balaban, 2020). The predominant reservoirs for HEV-3 include swine, rabbits, deer, and mongoose while the main reservoirs for HEV-4 are humans and swine (Sridhar et al., 2017). Genotypes 5 and 6 have been identified only in animals, specifically wild boars. Genotypes 7 and 8 are novel genotypes which have been recently isolated from camelids (Woo et al., 2014, 2016). Genotype 7, which was identified in dromedaries, could have a significant impact on human health as it was detected in a liver transplant patient who developed chronic HEV infection after consuming camel meat and camel-derived products (Lee et al., 2016). To date, scientists have identified more than 30 HEV subtypes (Smith et al., 2020). Only a few of these subtypes, such as subtype HEV-3c, 3f, 4b, and 4c, are of major clinical importance as they have been identified as key drivers of zoonotic transmission or severe clinical manifestations such as acute liver failure (Abravanel et al., 2020; Hakze-van der Honing et al., 2011; Nakano et al., 2018; Sato et al., 2020).
Hepatitis E virus can be transmitted through two main routes: the faecal-oral route, which involves the consumption of faecally contaminated water, and zoonotic transmission, which occurs when humans consume raw or undercooked meat from HEV-infected animals such as swine, deer and wild boar (van der Poel & Rzezutka, 2017). Person-to-person transmission is relatively infrequent but there have been reported cases of mother-to-infant transmission and transmission through solid organ transplants and blood transfusions (Arankalle & Chobe, 2000; Kumar & Sarin, 2013; Kumar et al., 2001; Murkey et al., 2017).
The presence of HEV in water is a growing concern as both drinking and irrigation water can be contaminated, causing a further spread of HEV through fresh produce and potable water consumption (Kokkinos et al., 2017; Salvador et al., 2020; Tripathy et al., 2019). Modes of water contamination may differ depending on the region but typical contamination scenarios include conditions such as heavy rains, water flowing through contaminated soil or leaking sewage pipes, and living in population-dense areas with no access to safe water supply (Khuroo, 2011). Studies in America, Asia and Europe have demonstrated the presence of HEV in groundwater, rivers, and wastewater (Baez et al., 2017; Takuissu et al., 2022; van der Poel & Rzezutka, 2017). Molecular analysis of wastewater runoff and faeces from pig farms in North Carolina, United States of America (USA), presented evidence of HEV RNA in these samples. Waste from pig farms is often disposed of through land application such as fertilisation and irrigation of crops, which means it could inadvertently seep into groundwater that is used for drinking, resulting in HEV exposure of nearby communities and other pigs on the farms (Kase et al., 2009; Meester et al., 2021). Employment at pig farms and wastewater treatment works (WWTWs) may also be considered a risk factor for HEV transmission (Beyer et al., 2020).
Presently, there is a paucity of data regarding the prevalence and diversity of HEV in South Africa (SA). Early studies in SA indicated a modest seroprevalence of 2.4% in 1994 and then 5.8–14.3% by 1996, suggesting persistent virus circulation and endemicity (Grabow et al., 1994, 1996). More recent studies have revealed a significant increase in the seroprevalence of HEV among blood donors as well as an increase in the incidence and genetic diversity of HEV in pigs and pig-derived products (Adelabu et al., 2017; Chauhan & Gordon, 2022; Lopes et al., 2017; Maponga et al., 2020). A linear relationship has also been reported between seroprevalence, age, and geographical area (Tucker et al., 1996). Despite progress in surveillance, characterisation, and risk factor identification of HEV, limited attention has been paid to asymptomatic individuals or those with mild symptoms who may still be shedding the virus but do not seek medical care. A comprehensive understanding of the prevalence and distribution of HEV in a region cannot be obtained based on clinical case data alone. Therefore, this exploratory study investigated the prevalence and diversity of HEV in SA in various water matrices. Water samples from WWTWs, rivers, runoff from selected ablution facilities, and a piggery were screened using real-time reverse transcription-polymerase chain reaction (RT-PCR), and HEV strains were further characterised by genotyping using Sanger sequencing.
Materials and Methods
Sample Collection
Systematic water sampling was conducted from February to September 2021; samples (1 L) were collected from selected rivers, influent from WWTWs, and surface runoff at communal standpipe/ablution sites in seven provinces of SA, namely: Free State (FS), Gauteng (GP), Kwa-Zulu Natal (KZN), Limpopo (L), Mpumalanga (MP), North West (NW), and Western Cape (WC). The sampling frequency varied from weekly to biweekly for WWTWs and surface water (river and standpipe/ablution) sites, respectively. Grab sampling was used for all sites except one WWTW site, which used composite sampling. From May to September 2022, 2-L samples of raw wastewater were collected from pig pens, effluent processed through the solid–liquid separation and autothermal thermophilic aerobic digestion (ATAD) system, and water from the adjacent river at a piggery in Tshwane. All the samples were transported in cooler boxes and refrigerated at 4 °C until further processing.
Virus Recovery
The skimmed milk flocculation protocol as outlined by Falman et al. (2019) was used to recover viruses from the water samples. Briefly, 5% weight/volume (w/v) pre-flocculated skimmed milk solution (2 mL) (Oxoid, Basingstoke, Hampshire, UK) was added to a 200 mL sample, followed by pH adjustment to 3.0–4.0 using 1 M hydrochloric acid (HCl) (Merck KGaA, Darmstadt, Germany) and gentle shaking at 200 revolutions per minute (rpm) at room temperature (20–25 °C) for 2 hours (h). The samples were then clarified by centrifugation (Sorvall® Super T20, du Pont, Wilmington, DE, USA) at 4500 × g for 30 min (min) at 4 °C. The supernatant was decanted, and the pellet was resuspended in 2 mL phosphate-buffered saline (PBS; pH 7.4; Sigma-Aldrich, St. Louis, MO, USA). Aliquots (1 mL) of each recovered virus concentrate were stored at − 20 °C for analysis, while the remaining volumes were stored at − 70 °C.
Nucleic Acid Extraction
The virus concentrates were subjected to chloroform treatment before extraction to reduce potential PCR inhibitors. Briefly, chloroform (Merck KGaA) (250 μL) was added to 1 mL of the virus concentrate. The solution was vortexed three times for 15 s (sec) and incubated at room temperature for 5 min, followed by centrifugation at 5000× g for 3 min. Following phase separation, the aqueous phase (1 mL) was transferred to a new 2 mL tube. This process was performed once for wastewater and surface water samples and twice for pig farm samples.
Total nucleic acids were extracted from the wastewater and surface water samples using a QIAamp® UltraSens® Virus Kit (QIAGEN, Hilden, Germany) following the manufacturer’s instructions. Nucleic acids were eluted in AVE buffer (100 μL) and stored at − 80 °C. Nucleic acids from recovered piggery and adjacent river samples were extracted using the EMAG® Nucleic Acid Extraction System (bioMérieux, Marcy-l’Étoile, France) following the manufacturer’s instructions, eluted in 100 μL of elution buffer, and then stored at − 80 °C.
Positive Control Preparation
An HEV-positive stool specimen, kindly provided by Prof. Wolfgang Preiser from Stellenbosch University, SA, was used to generate a positive control plasmid for the detection assay. Total nucleic acids were extracted from a 10% suspension of the stool specimen using the QIAamp® UltraSens® Virus Kit (QIAGEN) according to the manufacturer’s instructions. To construct the plasmid, complementary DNA (cDNA) was synthesised using a Protoscript® II Reverse Transcriptase Kit (New England Biolabs Inc., Ipswich, MA, USA), 0.5 mM deoxynucleotide triphosphates (dNTPs) (New England Biolabs Inc.), and random hexamer primers (Roche, Basel, Switzerland) according to the manufacturer’s instructions. A PCR was then performed using the cDNA (5 μL), EmeraldAmp MAX HS PCR Master Mix (25 μL) (Takara Bio Inc., Kusatsu, Shiga, Japan), nuclease-free water (18 μL; Promega Corporation, Madison, WI, USA), and published primers: JVHEVF and JVHEVR, at a final concentration of 0.4 μM for each primer (Table 1). The thermal cycling parameters were 95 °C for 3 min, followed by 40 cycles at 94 °C for 1 min, 45 °C for 1 min, and 72 °C for 1 min, with a final extension at 72 °C for 7 min.
The 69 bp product was then purified and cloned into a pJET1.2/blunt vector using the CloneJET™ PCR Cloning Kit (Thermo Fisher Scientific, Waltham, MA, USA) and 10-beta Escherichia coli (E. coli) chemically competent cells (New England Biolabs Inc.). Following colony PCR, the plasmid was purified using the Zyppy™ Plasmid Miniprep Kit (Zymo Research, Irvine, CA, USA) and stored at − 20 °C until further use.
Real-Time RT-PCR Assay for HEV Detection
A one-step real-time RT-PCR assay (QuantiFast Pathogen® RT-PCR + IC kit, QIAGEN) was used to amplify the 69 bp region of the highly conserved ORF3. The nucleic acid template (5 μL), nuclease-free water (7.55 μL; Promega Corp.), 5× Reaction Mix (5 μL), and published primers and probe (Table 1) at a final concentration of 0.4 μM and 0.08 μM respectively, were added to a final reaction volume of 25 μL. The kit’s internal control (IC) was used to monitor PCR inhibition.
The assay was performed using the QuantStudio™ 5 platform (Applied Biosystems, Waltham, MA, USA). The cycling conditions were as follows: reverse transcription at 50 °C for 30 min, enzyme activation at 95 °C for 5 min, and 45 cycles at 95 °C for 15 s and 60 °C for 1 min. Fluorescence was measured during the extension step. The data was analysed with QuantStudio™ software (Applied Biosystems).
A standard curve was constructed using triplicate tenfold serial dilutions of the positive control plasmid (r2 = 0.995, efficiency = 94.57%, error = 0.063). The limit of detection (LOD) of the assay, defined as the 95% confidence minimum detectable template concentration, and the limit of quantification (LOQ), defined as the lowest concentration of HEV RNA that could be reliably quantified, were determined based on the standard curve. A cycle threshold (Ct) value of 40 was used as the cut-off value for HEV-positive samples. In instances where substantial IC inhibition was observed, that is when no amplification signals were detected from both the target sequence and the IC or the Ct value of the IC was ≥ 40 with no amplification from the target sequence, the RT-PCR assay was repeated with diluted nucleic acid (1:10 dilution). If the IC tested positive after dilution, inhibition was considered as resolved. Hepatitis E virus RNA in selected positive samples (Ct < 33.9) was quantified using the real-time RT-qPCR assay described above.
Molecular Characterisation
Amplification of the Partial Capsid Region
Samples with a Ct value ≤ 35.5 were selected for molecular characterisation using a two-step RT-PCR. The extracted nucleic acid template (5 μL) was used to synthesise cDNA (20 μL) using random hexamer primers (30 μM) (Thermo Fisher Scientific), 0.5 mM dNTPs (New England Biolabs Inc.), and the Protoscript® II Reverse Transcriptase Kit (New England Biolabs Inc.), according to the manufacturer’s instructions. The 348 bp partial capsid region (ORF2) was amplified by nested PCR using a 50 μL reaction containing cDNA (5 μL), EmeraldAmp MAX HS PCR Master Mix (25 μL) (Takara Bio Inc.), nuclease-free water (Promega Corp.), and published primer sets (Table 1) at a final concentration of 0.2 μM. The first-round PCR product (1 μL) was used as the template for the second round. The cycling conditions for both the first and second rounds were as follows: 95 °C for 3 min, followed by 40 cycles of 94 °C for 1 min, 45 °C for 1 min, and 72 °C for 1 min, with a final extension at 72 °C for 7 min.
Visualisation of PCR Products and Purification
Amplicons were analysed by 1.5% agarose gel electrophoresis using Agarose LE (Cleaver Scientific, Rugby, Warwickshire, UK) and purified using the DNA Clean and Concentrator® − 25 Kit (Zymo Research) according to the manufacturer’s instructions. Amplicons with low yields were cloned using the CloneJET™ PCR Cloning Kit (Thermo Fisher Scientific) and 10-beta E. coli chemically competent cells (New England Biolabs Inc.) as per the manufacturer’s instructions.
Sanger Sequencing
The PRISM BigDye® Terminator v3.1 Cycle Sequencing Kit (Thermo Fisher Scientific) was used for cycle sequencing of amplicons of the correct size. The cycling conditions were as follows: initial denaturation (94 °C for 3 min), followed by 25 cycles of denaturation (94 °C for 30 s), annealing (50 °C for 10 s), and extension (60 °C for 4 min). Sequencing reactions were referred to Inqaba Biotec, Pretoria, SA, for purification and analysis.
Phylogenetic Analysis
The BioEdit Sequence Alignment Editor Software was used for base calling, while Sequencher™ DNA Sequence Analysis Software version 4.9 (Gene Codes Corporation, Ann Arbor, MI, USA) was utilised for sequence assembly and translation (Hall, 1999). To identify the genotypes, the Nucleotide Basic Local Alignment Search Tool (BLAST) was applied to compare them to known sequences in the GenBank database. The HEV Genotyping Tool version 1.0 (https://www.rivm.nl/mpf/typingtool/hev/) was used to assign the subtypes. All sequences were deposited in GenBank under accession numbers OR604636 – OR604689.
Sequence alignment was performed with the Multiple Alignment using Fast Fourier Transform (MAFFT) version 7 software (https://mafft.cbrc.jp/alignment/server/) (Katoh et al., 2019). The derived HEV sequences were aligned to GenBank top hits and HEV reference subtypes as recommended by Smith et al. (2020). A maximum-likelihood phylogenetic tree was constructed by using the Molecular Evolutionary Genetics Analysis Program Version X (MEGA X) software; the evolutionary distances were determined using the Jukes-Cantor method and validated by replication with 1000 bootstraps.
Statistical Analysis
Descriptive statistics were used to summarise HEV detection rates. The significance of differences in HEV detection rate according to sample type, province, and time period were assessed using the chi square test on OpenEpi (Sullivan et al., 2009) (https://www.openepi.com/TwobyTwo/TwobyTwo.htm, accessed 21/02/2024).
Results
Hepatitis E Virus Detection
A total of 536 water samples were collected from February to September 2021. Of these, 328 were influent samples from WWTWs, 188 were river samples, and only 20 were from standpipe/ablution sites. Hepatitis E virus was detected in 21.8% (117/536) of the samples, with detection rates of 22% (72/328) in wastewater influent, 22% (42/188) in river water, and 10% (2/20) in ablution runoff/standpipe site samples. The average Ct value for the positive samples was 34.5 (WWTWs), 34.6 (rivers), and 36.8 (standpipe/ablution). No statistically significant difference was observed in the detection rates in wastewater influent and river water (p = 0.3382). Inhibition was observed in 18.9% (98/536) of samples. After dilution, 14.3% (14/98) tested positive (average Ct value: 36.28) while 72% (71/98) were negative and 13 samples (2.4%) remained inconclusive. In total, 406 samples were negative. The LOD of the real-time RT-qPCR assay was determined as 100 genome copies (gc)/reaction and the LOQ as 1000 gc/reaction. The HEV RNA concentrations for the positive samples ranged between 1.89 × 103 gc/reaction and 2.35 × 104 gc/reaction.
Hepatitis E virus was detected in all seven provinces with the WC having the highest percentage of positive samples (38.1%, 32/84), followed by NW (37.5%, 9/24) and then GP (23.3%, 57/245) (Fig. 1A). The HEV detection rate in the WC was significantly higher than all other provinces (p < 0.006) except for NW (p = 0.4627). A higher frequency of positive samples was observed in the winter months of June and July, with a progressive decrease in positivity from August (Fig. 1B). No significant difference was observed from the samples in the WC (p > 0.05) between February-April (30% positivity), May–July (45% positivity), and August–September (40% positivity). However, GP demonstrated a statistically significant increase in positive samples (p < 0.02) during May–July (31% positivity) compared to both the February-April (14% positivity) and August–September (15% positivity) time periods.
Between May and September 2022, an additional 39 samples were collected from the Tshwane piggery and the adjacent river. These included 10 samples of raw effluent, 10 samples of effluent after settling, 9 samples of effluent after separation, and 10 samples from the adjacent river. All effluent samples from the piggery (29) tested positive for HEV (average Ct value: 27.5) with RNA concentrations ranging between 1.13 × 104 gc/reaction and 1.79 × 106 gc/reaction, while all samples from the river adjacent to the piggery (10) tested negative.
Molecular Characterisation
Amplification of the Partial Capsid Region
Of the 74 WWTW influent and surface water samples selected for genotyping (average Ct value: 33.18), the HEV partial capsid region was successfully amplified in 33.8% (25/74) of the samples. Additionally, 100% (29/29) of the HEV-positive piggery effluent samples were successfully amplified (average Ct value: 27.54). A total of 54 HEV strains were sequenced, genotyped, and phylogenetically analysed.
Phylogenetic Analysis
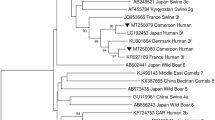
A maximum likelihood phylogenetic tree was constructed to determine the relatedness of the sequences obtained in this study to those in GenBank (Fig. 2). Most of the strains clustered with strains from SA and were classified as HEV-3 (53/54), with only one strain being identified as HEV-4 (1/54). Further analysis using the HEV typing tool revealed that three HEV-3 strains belonged to subtypes HEV-3f and HEV-3c, and the HEV-4 strain belonged to subtype HEV-4b. The remaining strains (n = 51) were categorically of genotype HEV-3, but the online HEV typing tool was unable to classify them into a subtype.
Maximum-likelihood phylogenetic tree of partial capsid gene sequences (348 bp, ORF2) representing the 54 HEV strains detected in the study. Evolutionary distances were determined using the Jukes-Cantor method, conducted in MEGA X. Numbers next to the branches indicate nodes where bootstrap support was > 70% out of the 1000 replicates. Reference sequences are based on Smith et al. (2020). Closely related strains from GenBank are indicated by accession numbers and study strains are indicated by triangles ▲(river), solid circles ● (WWTWs), and open circles ○ (piggery)
Discussion
Wastewater-based epidemiology (WBE) was applied to investigate the prevalence and diversity of HEV in wastewater and other water matrices across both rural and urban settings in SA. The WBE field is a relatively new and dynamic area of research that can complement conventional surveillance systems and serve as an early warning system for infectious disease outbreaks in specific regions (Sims & Kasprzyk-Hordern, 2020).
Real-time RT-PCR was used to detect HEV. This detection method has been used in previous studies from other countries, including Pakistan (Ahmad et al., 2010), Sweden (Hellmér et al., 2014), Portugal (Salvador et al., 2020), and Argentina (Lo Castro et al., 2023). Because of its widespread availability, dependability, and affordability, this method is regarded as the gold standard for virus detection in water matrices. Other assays, such as the reverse transcription droplet digital PCR (RT-ddPCR) assay, have been developed and have demonstrated improved sensitivity, specificity, and reproducibility compared to real-time RT-PCR (Nicot et al., 2016). However, the costs and need for specialised equipment make such assays impractical for most researchers, especially in low- and middle-income countries (LMICs).
Real-time RT-PCR inhibition could not be resolved for 2.4% (13/536) of the wastewater and surface water samples collected from February-September 2021. Compounds present in samples, such as metal ions, lipids, proteins, and polysaccharides, can inhibit nucleic acid amplification (Schrader et al., 2012). Diluting the RNA can reduce the concentration of such compounds, therefore assisting with the amplification of the target. Inhibition was predominantly detected in the wastewater samples, as well as in a few surface water samples extracted manually using the QIAamp® UltraSens® Virus Kit (QIAGEN). No inhibition was observed in the piggery samples collected between May and September 2022, which could be due to the use of a different extraction method (EMAG® Nucleic Acid Extraction System) or the less complex matrix of piggery wastewater with fewer potential PCR inhibitors. Inhibition leads to a possible underestimation of the positivity rate within a sample pool.
A 21.8% HEV positivity rate was found in wastewater and surface water samples collected between February and September 2021. The detection rate in wastewater influent was high (22%), this is likely because significant concentrations of HEV are shed in stool excreted by infected individuals, making it more detectable in wastewater influent, which is a pooled sample from thousands of people in each area (Hellmér et al., 2014; Iaconelli et al., 2020; Takuissu et al., 2022). A recent meta-analysis of HEV in environmental waters showed that its prevalence is higher in untreated wastewater (15.1%) compared to surface waters (7.4%) (Takuissu et al., 2022). Interestingly, the prevalence of HEV in wastewater is typically lower in most industrialised countries, such as Italy (5.4%) (Iaconelli et al., 2020), Norway (8.0%) (Myrmel et al., 2015), and Portugal (3.3%) (Matos et al., 2018) when compared to LMICs.
For the surface water samples collected between February and September 2021, the detection rate of HEV in the rivers and standpipe/ablution sites was 22% and 10%, respectively. These results are comparable to the detection rate in WWTW influent. This study shows frequent HEV contamination of SA river systems at low levels. According to the Green Drop Watch Report for 2023, 70.1% of wastewater treatment plants in SA are dysfunctional or have exceeded their design capacity (Department of Water and Sanitation, Republic of South Africa, 2023). There are also unsewered informal settlements with rapidly changing population numbers close to rivers leading to inadequate disposal of sewage. Many of these informal settlements have backyard pigs and other free-ranging animals which could shed HEV. The lower detection rates of HEV, or lack thereof, in surface water in industrialised countries can be attributed to access to proper sanitation, hygiene, and adequate disposal and treatment of sewage (Ahmad et al., 2022).
All piggery samples (29) tested positive for HEV. Pigs are one of the main reservoirs of HEV (Ahmed & Nasheri, 2023). Infected pigs can shed the virus in their stool, therefore, exposure to manure and faecally contaminated pig pens is a potential source of infection for both pigs and farm workers (Beyer et al., 2020; Meester et al., 2021). The presence of HEV RNA in the environment is not conclusive evidence of infectious virions, but it does highlight the potential risks. Studies have shown that working at WWTWs and piggeries/slaughterhouses as an HEV risk factor (Bagulo et al., 2020; Pavio et al., 2017; Vaidya et al., 2003). In Antioquia, Columbia, pig farm employees had an HEV seroprevalence of 11.25% (Baez et al., 2017). To mitigate the risk of HEV infection in farm workers, it is imperative to provide them with adequate personal protective equipment and to handle all waste carefully. Vaccination of these cohorts for disease prevention would be optimal. However, the only available vaccine that has demonstrated good safety and high efficacy is solely licensed in China and Pakistan since 2011 and 2020, respectively (Zhong et al., 2023).
None of the samples from the river adjacent to the piggery tested positive for HEV. These results align with a study conducted by Kasorndorkbua et al. (2005) in the USA, which suggested that contamination of the adjacent river may not be evident because the virions in the river being present at a low concentration and thus undetectable by the applied detection methods. Alternatively, the effective implementation of aerobic digestion similar to ATAD described by Wi et al. (2019), coupled with the initial release of the effluent into a wetland, likely ensured sufficient treatment of the waste before it reached the river.
Although the seasonality of HEV is unclear (Fares, 2015), an increased incidence of HEV in the summer months of May–July in Europe suggests a summer seasonality of HEV in that region (Healy et al., 2022). Lu and colleagues (2013) reported that the seasonality of HEV differed according to the geographical area in China (Lu et al., 2013), emphasising the complexity of describing HEV seasonality. In this study, HEV was detected throughout the sampling period at varying frequencies across the months and provinces. Analysis of the data revealed a gradual increase in the incidence of HEV from March, with a peak observed in June and July. As the winter season progressed, there was a gradual decrease in the incidence of HEV. Conclusions on the seasonality of HEV in SA could not be drawn from this study because the sample collection duration was too short, however it is clear the virus circulates in the population for most of the year. This suggests that regular environmental and clinical surveillance is required to predict potential outbreaks, especially in densely populated rural areas with limited access to clean water and proper sanitation.
The South African National Health Act (Act No. 61 of 2003) classifies hepatitis E as a category 2 medical condition, requiring health care professionals to report clinical/laboratory confirmed cases to the Department of Health through written or electronic communication within 7 days. However, between June 2018 and May 2023, only 247 cases were reported in the notifiable medical condition system for the entire country (NICD, 2023). Considering the HEV detection rate in the current study, HEV infections are likely underreported or misdiagnosed in the country. Research has shown that most cases of hepatitis E are misdiagnosed as drug-induced liver injury (DILI), especially in LMICs (El-Mokhtar et al., 2021; Kamar et al., 2014). In SA, over five million people were receiving antiretroviral therapy in 2022 (UNAIDS, 2022). Antiretroviral therapy causes DILI in patients with HIV infection (Pillaye et al., 2020), often leading to healthcare workers misdiagnosing HEV-induced liver inflammation as DILI. This is caused by limited HEV awareness in clinical settings. Environmental surveillance is therefore necessary to improve our understanding of the prevalence of HEV in the country. The low prevalence of the virus in the clinical setting could also be due to its low concentrations, causing sporadic, asymptomatic infections as it spreads (Grabow et al., 1996).
Among the 74 positive samples collected between February and September 2021 and selected for genotyping, only 33.8% (25/74) were positive by nested PCR. This low positivity rate may be due to low concentrations of the virus and inhibitors within the sample. All piggery samples collected between May and September 2022 were successfully genotyped. These samples had a 100-fold higher viral concentration than the 2021 samples, which likely facilitated efficient amplification using nested PCR. Phylogenetic analysis revealed the predominance of HEV-3 in the country (98.1% [53/54]), with a single detection of genotype 4 (1.85% [1/54]). Previous investigations in various countries, including Italy (La Rosa et al., 2015, 2017), Germany (Beyer et al., 2020), and Australia (Miura et al., 2016), have reported the presence of these genotypes in similar environmental sources. Nucleotide BLAST analysis indicated that most sequences from this study (74%) were closely related to an HEV-3 sequence identified in a renal transplant patient (KU178916.1) from the WC province in SA. Interestingly, the phylogenetic tree showed that strains from the piggery formed a distinct cluster that was more closely related to strains detected in a WWTW and river in this study than to their top hit from GenBank (renal transplant patient) (Andersson et al., 2015). The samples in this cluster could not be definitively subtyped due to insufficient bootstrap support, as the bootstrap cut-off was 70%. Future studies should characterise a larger region of the genome to determine whether or not these samples belong to a new, possibly unclassified subtype.
One study strain was identified as subtype 3f and clustered with a human strain from France (Lhomme et al., 2015) with a significant bootstrap value (80%). Infection with this subtype correlates with elevated HEV viral titres, a higher fever risk, and increased hospitalisation rates (Abravanel et al., 2020). Three strains (5.6%) were most closely related to an HEV-3 strain detected in pork liver spread (MF503296.1) from SA (Korsman et al., 2019). The remaining sequences (18.5%) matched those from swine in Nigeria (KJ451631.1), Italy (MK532915.1), SA (OM104034.1), Japan (AB094271.1), and China (KJ001830.1). Overall, the sequences from this study showed nucleotide identities between 93.0% and 97.0% with their top hits in GenBank. Within the SA context, there is limited sequence data available for HEV, with only 21 sequences from SA available in GenBank to date.
The subtype 3c strains, related to HEV-3 detected in backyard pigs in KZN (OM104034.1) (Chauhan & Gordon, 2022), were detected in GP, MP and NW. This subtype is widespread in pigs and wild boars (Fenaux et al., 2018). Although 3c infections are associated with a low viral load (Abravanel et al., 2020), a study spanning four European countries, comparing HEV infections in symptomatic and asymptomatic individuals found that infection with 3c leads to more asymptomatic infections (Smith et al., 2015). This could further perpetuate the spread of HEV. HEV strains found in human populations have been detected in water matrices in both industrialised and LMICs (Li et al., 2017). The association of the HEV strains in this study with strains from clinical cases suggests a wider community spread than previously thought. This poses a health risk, especially to immunocompromised people, who may suffer from chronic infection after exposure to the virus in rivers and other water sources.
Our findings are consistent with those of Beyer et al. (2020), who similarly identified both subtypes 3c and 3f in environmental samples in Germany. Subtypes 3c and 3f are predominant in Europe; however, subtype 3f has also been found in Thailand and Japan (Nakano et al., 2018). Infection with subtype 3f is associated with an increase in hospitalisation and a higher viral load, whereas subtype 3c has mostly been implicated in increased HEV incidence. (Abravanel et al., 2020; Nakano et al., 2018). The single HEV 4 strain was classified as subtype 4b and clustered with a strain from a pig in China and a reference strain, as described by Smith et al. (2020) (81% bootstrap support). Infections with genotype 4 have been noted to cause more severe clinical manifestations in humans (Hakze-van der Honing et al., 2011). The phylogenetic findings in this study are in line with most studies that have investigated the genetic diversity of HEV in water matrices, as HEV-3 has been recognised as the most predominant genotype, with HEV-4 being present in low quantities (Iaconelli et al., 2020; Takuissu et al., 2022). A limitation of our study is that we could not confirm whether the viruses identified in this study were infectious or not. Future studies should incorporate viability PCR in their methods to detect viable viruses that could potentially cause infection.
Conclusion
From the available information, this is the first study of its kind to be conducted in SA. Our analyses present compelling evidence for the presence of HEV in pig slurry, wastewater, and surface waters within SA. Phylogenetic analysis established a clear link between the strains detected in the environment and those previously detected in human cases, indicating an ongoing circulation of HEV in the population that extends beyond reported clinical cases. This study not only expands our understanding of water-based epidemiology, but also significantly contributes to bridging the existing knowledge gap concerning the prevalence and distribution of HEV in our region. Furthermore, the valuable data obtained from this research can inform and support the development of effective vaccines and implementation of preventive measures to mitigate HEV outbreaks.
References
Abravanel, F., Dimeglio, C., Castanier, M., Péron, J. M., Kamar, N., Lhomme, S., & Izopet, J. (2020). Does HEV-3 subtype play a role in the severity of acute hepatitis E? Liver International, 40(2), 333–337. https://doi.org/10.1111/liv.14329
Adelabu, O. A., Chuks Iweriebor, B., Nwodo, U. U., Obi, L. C., & Okoh, A. I. (2017). Incidence and molecular characterization of hepatitis E virus from swine in Eastern Cape. South Africa. Advances in Virology, 2017, 1073253. https://doi.org/10.1155/2017/1073253
Ahmad, T., Jin, H., Dhama, K., Yatoo, D. M., Tiwari, R., Bilal, M., Dhawan, M., Emran, T., Alestad, J., Alhani, H., BinKhalaf, H., & Rabaan, A. (2022). Hepatitis E virus in pigs and the environment: An updated review of public health concerns. Narra Journal, 2(2):e78. https://doi.org/10.52225/narra.v2i2.78
Ahmed, R., & Nasheri, N. (2023). Animal reservoirs for hepatitis E virus within the paslahepevirus genus. Veterinary Microbiology, 278, 109618. https://doi.org/10.1016/j.vetmic.2022.109618
Andersson, M. I., Stead, P. A., Maponga, T., Van Der Plas, H., & Preiser, W. (2015). Hepatitis E virus infection: An underdiagnosed infection in transplant patients in Southern Africa? Journal of Clinical Virology, 70, 23–5. https://doi.org/10.1016/j.jcv.2015.06.081
Arankalle, V. A., & Chobe, L. P. (2000). Retrospective analysis of blood transfusion recipients: Evidence for post-transfusion hepatitis E. Vox Sanguinis, 79(2), 72–4. https://doi.org/10.1159/000031215
Aslan, A. T., & Balaban, H. Y. (2020). Hepatitis E virus: Epidemiology, diagnosis, clinical manifestations, and treatment. World Journal of Gastroenterology, 26(37), 5543–5560. https://doi.org/10.3748/wjg.v26.i37.5543
Baez, P. A., Lopez, M. C., Duque-Jaramillo, A., Pelaez, D., & Molina, F. (2017). First evidence of the hepatitis E virus in environmental waters in Colombia. PLoS ONE, 12(5), e0177525. https://doi.org/10.1371/journal.pone.0177525
Bagulo, H., Majekodunmi, A. O., & Welburn, S. C. (2020). Hepatitis E in Sub Saharan Africa – A significant emerging disease. One Health, 11, 100186. https://doi.org/10.1016/j.onehlt.2020.100186
Beyer, S., Szewzyk, R., Gnirss, R., Johne, R., & Selinka, H.-C. (2020). Detection and characterization of hepatitis E virus genotype 3 in wastewater and urban surface waters in Germany. Food and Environmental Virology, 12(2), 137–47. https://doi.org/10.1007/s12560-020-09424-2
Chauhan, R. P., & Gordon, M. L. (2022). Characterization of a near full-length hepatitis E virus genome of subtype 3c generated from naturally infected South African backyard pigs. Pathogens, 11(9), 1030. https://doi.org/10.3390/pathogens11091030
Department of Water and Sanitation, Republic of South Africa. (2023). Green Drop Watch Report. Retrieved February 4, 2024, from https://ws.dws.gov.za/iris/releases/GDWR.pdf
El-Mokhtar, M. A., Ramadan, H. K., Thabet, M. M., Abd-Elkader, A. S., Fouad, M., Sallam, M. M., Elgohary, E. A., Abd El-Hafeez, A. A., Mohamed, M. E., & Sayed, I. M. (2021). The unmet needs of hepatitis E virus diagnosis in suspected drug-induced liver injury in limited resource setting. Frontiers in Microbiology, 12, 737486. https://doi.org/10.3389/fmicb.2021.737486
Falman, J. C., Fagnant-Sperati, C. S., Kossik, A. L., Boyle, D. S., & Meschke, J. S. (2019). Evaluation of secondary concentration methods for poliovirus detection in wastewater. Food and Environmental Virology, 11(1), 20–31. https://doi.org/10.1007/s12560-018-09364-y
Fares, A. (2015). Seasonality of hepatitis: A review update. Journal of Family Medicine and Primary Care, 4(1), 96–100. https://doi.org/10.4103/2249-4863.152263
Fenaux, H., Chassaing, M., Berger, S., Jeulin, H., Gentilhomme, A., Bensenane, M., Bronowicki, J. P., Gantzer, C., Bertrand, I., & Schvoerer, E. (2018). Molecular features of hepatitis E virus circulation in environmental and human samples. Journal of Clinical Virology, 103, 63–70. https://doi.org/10.1016/j.jcv.2018.04.003
Garson, J. A., Ferns, R. B., Grant, P. R., Ijaz, S., Nastouli, E., Szypulska, R., & Tedder, R. S. (2012). Minor groove binder modification of widely used TaqMan probe for hepatitis E virus reduces risk of false negative real-time PCR results. Journal of Virological Methods, 186(1–2), 157–60. https://doi.org/10.1016/j.jviromet.2012.07.027
Grabow, W. O., Favorov, M. O., Khudyakova, N. S., Taylor, M. B., & Fields, H. A. (1994). Hepatitis E seroprevalence in selected individuals in South Africa. Journal of Medical Virology, 44(4), 384–8. https://doi.org/10.1002/jmv.1890440412
Grabow, W. O. K., Taylor, M. B., & Webber, L. M. (1996). Hepatitis E virus in South Africa. South African Journal of Science, 92(4), 178–180.
Hakze-Van Der Honing, R. W., Van Coillie, E., Antonis, A. F., & Van Der Poel, W. H. (2011). First isolation of hepatitis E virus genotype 4 in Europe through swine surveillance in the Netherlands and Belgium. PLoS ONE, 6(8), e22673. https://doi.org/10.1371/journal.pone.0022673
Hall, T. A. (1999). BioEdit: A user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series, 41(41), 95–98.
Healy, K., Freij, U., Ellerstad, M., Aulin, L. B. S., Brückle, L., Hillmering, H., Chen, M. S., & Gustafsson, R. (2022). Evaluating the prevalence of hepatitis E virus infection in a large cohort of European blood donors, 2015–2018. Journal of Viral Hepatitis, 29(9), 835–9. https://doi.org/10.1111/jvh.13682
Hellmér, M., Paxéus, N., Magnius, L., Enache, L., Arnholm, B., Johansson, A., Bergström, T., & Norder, H. (2014). Detection of pathogenic viruses in sewage provided early warnings of hepatitis A virus and norovirus outbreaks. Applied and Environmental Microbiology, 80(21), 6771–81. https://doi.org/10.1128/AEM.01981-14
Hoofnagle, J. H., Nelson, K. E., & Purcell, R. H. (2012). Hepatitis E. New England Journal of Medicine, 367(13), 1237–1244. https://doi.org/10.1056/NEJMra1204512
Huang, F. F., Haqshenas, G., Guenette, D. K., Halbur, P. G., Schommer, S. K., Pierson, F. W., Toth, T. E., & Meng, X. J. (2002). Detection by reverse transcription-PCR and genetic characterization of field isolates of swine hepatitis E virus from pigs in different geographic regions of the United States. Journal of Clinical Microbiology, 40(4), 1326–1332. https://doi.org/10.1128/JCM.40.4.1326-1332.2002
Iaconelli, M., Bonanno Ferraro, G., Mancini, P., Suffredini, E., Veneri, C., Ciccaglione, A. R., Bruni, R., Della Libera, S., Bignami, F., Brambilla, M., De Medici, D., Brandtner, D., Schembri, P., D’Amato, S., & La Rosa, G. (2020). Nine-year nationwide environmental surveillance of hepatitis E virus in urban wastewaters in Italy (2011–2019). International Journal of Environmental Research and Public Health, 17(6), 2059. https://doi.org/10.3390/ijerph17062059
Jothikumar, N., Cromeans, T. L., Robertson, B. H., Meng, X. J., & Hill, V. R. (2006). A broadly reactive one-step real-time RT-PCR assay for rapid and sensitive detection of hepatitis E virus. Journal of Virological Methods, 131(1), 65–71. https://doi.org/10.1016/j.jviromet.2005.07.004
Kamar, N., Dalton, H. R., Abravanel, F., & Izopet, J. (2014). Hepatitis E virus infection. Clinical Microbiology Reviews, 27(1), 116–138. https://doi.org/10.1128/CMR.00057-13
Kase, J. A., Correa, M. T., & Sobsey, M. D. (2009). Detection and molecular characterization of swine hepatitis E virus in North Carolina swine herds and their faecal wastes. Journal of Water Health, 7(2), 344–357. https://doi.org/10.2166/wh.2009.137
Kasorndorkbua, C., Opriessnig, T., Huang, F. F., Guenette, D. K., Thomas, P. J., Meng, X. J., & Halbur, P. G. (2005). Infectious swine hepatitis E virus is present in pig manure storage facilities on United States farms, but evidence of water contamination is lacking. Applied and Environmental Microbiology, 71(12), 7831–7837. https://doi.org/10.1128/AEM.71.12.7831-7837.2005
Katoh, K., Rozewicki, J., & Yamada, K. D. (2019). MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Briefings in Bioinformatics, 20(4), 1160–1166. https://doi.org/10.1093/bib/bbx108
Khuroo, M. S. (2011). Discovery of hepatitis E: The epidemic non-A, non-B hepatitis 30 years down the memory lane. Virus Research, 161(1), 3–14. https://doi.org/10.1016/j.virusres.2011.02.007
Kmush, B., Wierzba, T., Krain, L., Nelson, K., & Labrique, A. B. (2013). Epidemiology of hepatitis E in low- and middle-income countries of Asia and Africa. Seminars in Liver Disease, 33(1), 15–29. https://doi.org/10.1055/s-0033-1338111
Kokkinos, P., Kozyra, I., Lazic, S., Söderberg, K., Vasickova, P., Bouwknegt, M., Rutjes, S., Willems, K., Moloney, R., De Roda Husman, A. M., Kaupke, A., Legaki, E., D’Agostino, M., Cook, N., Von Bonsdorff, C. H., Rzeżutka, A., Petrovic, T., Maunula, L., Pavlik, I., & Vantarakis, A. (2017). Virological quality of irrigation water in leafy green vegetables and berry fruits production chains. Food and Environmental Virology, 9(1), 72–78. https://doi.org/10.1007/s12560-016-9264-2
Korsman, S., Bloemberg, J., Brombacher, M., Giuricich, A., Halley-Stott, R., & Kaba, M. (2019). Hepatitis E virus in patients with acute hepatitis in Cape Town, South Africa, 2011. South African Medical Journal, 109(8), 582–583. https://doi.org/10.7196/SAMJ.2019.v109i8.13867
Kumar, N., & Sarin, S. K. (2013). Hepatitis E – Is it a risk to transfusion safety? Asian Journal of Transfusion Science, 7(1), 1–3. https://doi.org/10.4103/0973-6247.106708
Kumar, R. M., Uduman, S., Rana, S., Kochiyil, J. K., Usmani, A., & Thomas, L. (2001). Sero-prevalence and mother-to-infant transmission of hepatitis E virus among pregnant women in the United Arab Emirates. European Journal of Obstetrics & Gynecology and Reproductive Biology, 100(1), 9–15. https://doi.org/10.1016/s0301-2115(01)00448-1
La Rosa, G., Della Libera, S., Brambilla, M., Bisaglia, C., Pisani, G., Ciccaglione, A. R., Bruni, R., Taffon, S., Equestre, M., & Iaconelli, M. (2017). Hepatitis E virus (genotype 3) in slurry samples from swine farming activities in Italy. Food Environmental Virology, 9(2), 219–229. https://doi.org/10.1007/s12560-016-9270-4
La Rosa, M. I., Purpari, G., Libera, S. D., Petricca, S., Guercio, A., Ciccaglione, A. R., Bruni, R., Taffon, S., Equestre, M., Fratini, M., Muscillo, M., & Giuseppina, M. (2015). Hepatitis A and E viruses in wastewaters, in river waters, and in bivalve molluscs in Italy. Food and Environmental Virology, 7(4), 316–324. https://doi.org/10.1007/s12560-015-9207-3
Lee, G.-H., Tan, B.-H., Chi-Yuan Teo, E., Lim, S.-G., Dan, Y.-Y., Wee, A., Kim Aw, P. P., Zhu, Y., Hibberd, M. L., Tan, C.-K., Purdy, M. A., & Teo, C.-G. (2016). Chronic infection with camelid hepatitis E virus in a liver transplant recipient who regularly consumes camel meat and milk. Gastroenterology, 150(2), 355–7.e3. https://doi.org/10.1053/j.gastro.2015.10.048
Lhomme, S., Abravanel, F., Dubois, M., Chapuy-Regaud, S., Sandres-Saune, K., Mansuy, J. M., Rostaing, L., Kamar, N., & Izopet, J. (2015). Temporal evolution of the distribution of hepatitis E virus genotypes in Southwestern France. Infection, Genetics and Evolution, 35, 50–55. https://doi.org/10.1016/j.meegid.2015.07.028
Li, H., Li, W., She, R., Yu, L., Wu, Q., Yang, J., Hu, F., Soomro, M. H., Shi, R., Hao, W., Zhao, Y., & Mao, J. (2017). Hepatitis E virus genotype 4 sequences detected in sewage from treatment plants of China. Food and Environmental Virology, 9(2), 230–233. https://doi.org/10.1007/s12560-016-9276-y
Lo Castro, I., Espul, C., de Paula, V. S., Altabert, N. R., Gonzalez, J. E., Lago, B. V., & Villar, L. M. (2023). High prevalence of hepatitis A and E viruses in environmental and clinical samples from West Argentina. Brazilian Journal of Infectious Diseases, 27(2), 102738. https://doi.org/10.1016/j.bjid.2022.102738
Lopes, T., Cable, R., Pistorius, C., Maponga, T., Ijaz, S., Preiser, W., Tedder, R., & Andersson, M. I. (2017). Racial differences in seroprevalence of HAV and HEV in blood donors in the Western Cape, South Africa: A clue to the predominant hev genotype? Epidemiology and Infection, 145(9), 1910–1912. https://doi.org/10.1017/S0950268817000565
Lu, Y. H., Qian, H. Z., Hu, A. Q., Qin, X., Jiang, Q. W., & Zheng, Y. J. (2013). Seasonal pattern of hepatitis E virus prevalence in swine in two different geographical areas of China. Epidemiology & Infection, 141(11), 2403–9. https://doi.org/10.1017/S0950268813000113
Maponga, T. G., Lopes, T., Cable, R., Pistorius, C., Preiser, W., & Andersson, M. I. (2020). Prevalence and risks of hepatitis E virus infection in blood donors from the Western Cape. South Africa. Vox Sanguinis, 115(8), 695–702. https://doi.org/10.1111/vox.12966
Matos, A., Mesquita, J. R., Gonçalves, D., Abreu-Silva, J., Luxo, C., & Nascimento, M. S. (2018). First detection and molecular characterization of hepatitis E virus in water from wastewater treatment plants in Portugal. Annals of Agricultural and Environmental Medicine, 25(2), 364–367. https://doi.org/10.26444/aaem/90497
Meester, M., Tobias, T. J., Bouwknegt, M., Kusters, N. E., Stegeman, J. A., & Van Der Poel, W. H. M. (2021). Infection dynamics and persistence of hepatitis E virus on pig farms – A review. Porcine Health Management, 7(1), 16. https://doi.org/10.1186/s40813-021-00189-z
Miura, T., Lhomme, S., Le Saux, J. C., Le Mehaute, P., Guillois, Y., Couturier, E., Izopet, J., Abranavel, F., & Le Guyader, F. S. (2016). Detection of hepatitis E virus in sewage after an outbreak on a French island. Food and Environmental Virology, 8(3), 194–9. https://doi.org/10.1007/s12560-016-9241-9
Murkey, J. A., Chew, K. W., Carlson, M., Shannon, C. L., Sirohi, D., Sample, H. A., Wilson, M. R., Vespa, P., Humphries, R. M., Miller, S., Klausner, J. D., & Chiu, C. Y. (2017). Hepatitis E virus-associated meningoencephalitis in a lung transplant recipient diagnosed by clinical metagenomic sequencing. Open Forum Infectious Diseases, 4(3), ofx121. https://doi.org/10.1093/ofid/ofx121
Myrmel, M., Lange, H., & Rimstad, E. (2015). A 1-year quantitative survey of Noro-, Adeno-, Human Boca-, and Hepatitis E viruses in Raw and Secondarily Treated Sewage from Two Plants in Norway. Food and Environmental Virology, 7(3), 213–23. https://doi.org/10.1007/s12560-015-9200-x
Nakano, T., Takahashi, M., Takahashi, K., Nagashima, S., Suzuki, Y., Nishigaki, Y., Tomita, E., Okano, H., Oya, Y., Shiraki, K., Takase, K., Sugimoto, K., Koyama, J., Mizuo, H., Ikezawa, K., Aikawa, T., Arai, M., & Okamoto, H. (2018). Hepatitis E virus subtype 3f strains isolated from Japanese hepatitis patients with no history of travel to endemic areas – The origin analyzed by molecular evolution. Virology, 513, 146–152. https://doi.org/10.1016/j.virol.2017.08.008
National Department of Health, South Africa. National Health ActNational Department of Health, South Africa. National Health Act, 2003 (Act No. 61 of 2003). Government Gazette No. 26595:869. Retrieved April 8, 2023, from https://www.gov.za/documents/national-health-act
National Institute of Communicable Diseases (NICD). (2023). Communicable Diseases Communique 22(6), 5. Retrieved July 9, 2023, from https://www.nicd.ac.za/publications/communicable-diseases-publications/communicable-diseases-communique/
Nicot, F., Cazabat, M., Lhomme, S., Marion, O., Sauné, K., Chiabrando, J., Dubois, M., Kamar, N., Abravanel, F., & Izopet, J. (2016). Quantification of HEV RNA by Droplet Digital PCR. Viruses, 8(8), 233. https://doi.org/10.3390/v8080233
Pavio, N., Doceul, V., Bagdassarian, E., & Johne, R. (2017). Recent knowledge on hepatitis E virus in suidae reservoirs and transmission routes to human. Veterinary Research, 48(1), 78. https://doi.org/10.1186/s13567-017-0483-9
Pillaye, J. N., Marakalala, M. J., Khumalo, N., Spearman, W., & Ndlovu, H. (2020). Mechanistic insights into antiretroviral drug-induced liver injury. Pharmacology Research & Perspectives, 8(4), e00598. https://doi.org/10.1002/prp2.598
Van Der Poel, W., & Rzezutka, A. (2017). Hepatitis E in Part 3: Specific excreted pathogens: Environmental and epidemiology aspects. Section 1: Viruses. Global Water Pathogen Project. https://doi.org/10.14321/waterpathogens.13
Purdy, M., Drexler, J., Meng, X.-J., Norder, H., Okamoto, H., Poel, W., Reuter, G., Souza, W., Ulrich, R., & Smith, D. (2022). ICTV virus taxonomy profile: Hepeviridae 2022. Journal of General Virology, 103(9), 001778. https://doi.org/10.1099/jgv.0.001778
Rein, D. B., Stevens, G. A., Theaker, J., Wittenborn, J. S., & Wiersma, S. T. (2012). The global burden of hepatitis E virus genotypes 1 and 2 in 2005. Hepatology, 55(4), 988–97. https://doi.org/10.1002/hep.25505
Salvador, D., Neto, C., Benoliel, M. J., & Caeiro, M. F. (2020). Assessment of the presence of hepatitis E virus in surface water and drinking water in Portugal. Microorganisms, 8(5), 761. https://doi.org/10.3390/microorganisms8050761
Sato, N., Watanabe, S., Miura, K., Morimoto, N., Takaoka, Y., Nomoto, H., Isoda, N., Nagashima, S., Takahashi, M., Okamoto, H., & Yamamoto, H. (2020). Clinical and virologic features of hepatitis E virus infection at a university hospital in Japan between 2000 and 2019. Journal of Medical Virology, 92, 3572–3583. https://doi.org/10.1002/jmv.26248
Schrader, C., Schielke, A., Ellerbroek, L., & Johne, R. (2012). PCR inhibitors – Occurrence, properties and removal. Journal of Applied Microbiology, 113(5), 1014–26. https://doi.org/10.1111/j.1365-2672.2012.05384.x
Sims, N., & Kasprzyk-Hordern, B. (2020). Future perspectives of wastewater-based epidemiology: Monitoring infectious disease spread and resistance to the community level. Environment International, 139, 105689. https://doi.org/10.1016/j.envint.2020.105689
Smith, D. B., Ijaz, S., Tedder, R. S., Hogema, B., Zaaijer, H. L., Izopet, J., Bradley-Stewart, A., Gunson, R., Harvala, H., Kokki, I., & Simmonds, P. (2015). Variability and pathogenicity of hepatitis E virus genotype 3 variants. Journal of General Virology, 96(11), 3255–3264. https://doi.org/10.1099/jgv.0.000264
Smith, D. B., Izopet, J., Nicot, F., Simmonds, P., Jameel, S., Meng, X.-J., Norder, H., Okamoto, H., Van Der Poel, W. H. M., Reuter, G., & Purdy, M. A. (2020). Update: Proposed reference sequences for subtypes of hepatitis E virus (species Orthohepevirus A). Journal of General Virology, 101(7), 692–698. https://doi.org/10.1099/jgv.0.001435
Sridhar, S., Teng, J. L. L., Chiu, T. H., Lau, S. K. P., & Woo, P. C. Y. (2017). Hepatitis E virus genotypes and evolution: Emergence of camel hepatitis E variants. International Journal of Molecular Sciences, 18(4), 869. https://doi.org/10.3390/ijms18040869
Sullivan, K. M., Dean, A., & Soe, M. M. (2009). Openepi: A web-based epidemiologic and statistical calculator for public health. Public Health Reports, 124(3), 471–474. https://doi.org/10.1177/003335490912400320
Takuissu, G. R., Kenmoe, S., Ndip, L., Ebogo-Belobo, J. T., Kengne-Ndé, C., Mbaga, D. S., Bowo-Ngandji, A., Oyono, M. G., Kenfack-Momo, R., Tchatchouang, S., Kenfack-Zanguim, J., Lontuo Fogang, R., Zeukoo Menkem, E., Kame-Ngasse, E., Magoudjou-Pekam, G. I., J. N., Nkie Esemu, Veneri, S., Mancini, C., et al. (2022). Hepatitis E virus in water environments: A systematic review and meta-analysis. Food and Environmental Virology, 14(3), 223–235. https://doi.org/10.1007/s12560-022-09530-3
The Joint United Nations Programme on HIV/AIDS (UNAIDS). (2022). Country factsheets, South Africa. Retrieved July 31, 2023, from https://www.unaids.org/en/regionscountries/countries/southafrica
Tripathy, A. S., Sharma, M., Deoshatwar, A. R., Babar, P., Bharadwaj, R., & Bharti, O. K. (2019). Study of a hepatitis E virus outbreak involving drinking water and sewage contamination in Shimla, India, 2015–2016. Transactions of the Royal Society of Tropical Medicine and Hygiene, 113(12), 789–96. https://doi.org/10.1093/trstmh/trz072
Tucker, T. J., Kirsch, R. E., Louw, S. J., Isaacs, S., Kannemeyer, J., & Robson, S. C. (1996). Hepatitis E in South Africa: Evidence for sporadic spread and increased seroprevalence in rural areas. Journal of Medical Virology, 50(2), 117–119. https://doi.org/10.1002/(SICI)1096-9071(199610)50:2<117::AID-JMV3>3.0.CO;2-D
Vaidya, S. R., Tilekar, B. N., Walimbe, A. M., & Arankalle, V. A. (2003). Increased risk of hepatitis E in sewage workers from India. Journal of Occupational and Environmental Medicine, 45(11), 1167–70. https://doi.org/10.1097/01.jom.0000088874.43855.2f
Wi, J., Lee, S., Kim, E., Lee, M., Koziel, J. A., & Ahn, H. (2019). Evaluation of semi-continuous pit manure recharge system performance on mitigation of ammonia and hydrogen sulfide emissions from a swine finishing barn. Atmosphere, 10(4), 170. https://doi.org/10.3390/atmos10040170
Wong, R. J., Cheung, R., Gish, R. G., & Chitnis, A. S. (2021). Prevalence of hepatitis e infection among adults with concurrent chronic liver disease. Journal of Viral Hepatitis, 28(11), 1643–55. https://doi.org/10.1111/jvh.13597
Woo, P. C. Y., Lau, S. K. P., Teng, J. L. L., Cao, K.-Y., Wernery, U., Schountz, T., Chiu, T. H., Tsang, A. K. L., Wong, P.-C., Wong, E. Y. M., & Yuen, K.-Y. (2016). New hepatitis E virus genotype in bactrian camels, Xinjiang, China, 2013. Emerging Infectious Diseases, 22(12), 2219–2221. https://doi.org/10.3201/eid2212.160979
Woo, P. C. Y., Lau, S. K. P., Teng, J. L. L., Tsang, A. K. L., Joseph, M., Wong, E. Y. M., Tang, Y., Sivakumar, S., Xie, J., Bai, R., Wernery, R., Wernery, U., & Yuen, K.-Y. (2014). New hepatitis E virus genotype in camels, the Middle East. Emerging Infectious Diseases, 20(6), 1044–1048. https://doi.org/10.3201/eid2006.140140
Zhong, G., Zhuang, C., Hu, X., Chen, Q., Bi, Z., Jia, X., et al. (2023). Safety of hepatitis E vaccination for pregnancy: A post-hoc analysis of a randomized, double-blind, controlled phase 3 clinical trial. Emerging Microbes & Infections, 12(1), 2185456. https://doi.org/10.1080/22221751.2023.2185456
Acknowledgements
This work is based on research supported wholly/in part by the Poliomyelitis Research Foundation (PRF), the Department of Science and Innovation (DSI) (EUREKA Network Call), and the Water Research Commission (WRC). We would also like to acknowledge the following entities for their bursaries awarded to K Salemane: National Research Foundation of South Africa (NRF), PRF, the DSI and the WRC. The research team would like to express their sincere gratitude to the management of the WWTWs and the Tshwane piggery for granting us permission to collect samples for the study at their facilities and to publish the data.
Funding
Open access funding provided by University of Pretoria. This work is based on research supported wholly/in part by the Poliomyelitis Research Foundation (J Mans grant number 21/74), the Department of Science and Innovation (DSI) (EUREKA Network Call, contract DSI/CON 0095/2020), and the Water Research Commission (WRC) (Project number C2020/2021-00686). The authors would also like to acknowledge the following entities for the bursaries awarded to K Salemane: NRF (grant number MND210714622684), PRF (grant number 21/43), the DSI (DSI/CON 0095/2020) and the WRC (C2020/2021-00686). Opinions expressed and conclusions arrived at, are those of the authors and are not necessarily to be attributed to any of the previously mentioned organisations.
Author information
Authors and Affiliations
Contributions
KS performed RNA extractions, sample screening and virus genotyping, data curation and formal analysis, and drafted and edited the manuscript. JM, LZC, GP, BG and MBT conceptualised the study and acquired funding. LZC, GP collected samples and recovered viruses. BG collected and processed samples. MBT co-supervised KS, collected samples, and critically reviewed the manuscript. JM supervised KS, extracted samples, and critically reviewed and edited the manuscript. All authors have approved the final version of the manuscript for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics Approval
All procedures performed in this study were by the ethical standards of the institutional research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. The study was approved by the Research Ethics Committee of the Faculty of Health Sciences at the University of Pretoria, Tswelopele Building, Level 4 in April 2021 and granted on 29 April 2021 (ethics approval no.:148/2021).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Salemane, K., Coetzee, L.Z., Pocock, G. et al. Water-Based Epidemiological Investigation of Hepatitis E Virus in South Africa. Food Environ Virol (2024). https://doi.org/10.1007/s12560-024-09596-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12560-024-09596-1