Abstract
Human noroviruses impose a considerable health burden globally. Here, a flow cytometry approach designed for their detection in biological waste and food samples was developed using antibody-coated magnetic beads. Antipeptide antibodies against murine norovirus and various human norovirus genotypes were generated for capture and coated onto magnetic beads. A flow cytometry assay was then implemented to detect bead-bound human norovirus GI.3 in patient stool samples and in norovirus-spiked mussel digestive tissues. The detection limit for stool samples was 105 gc/mL, thus bettering detection limits of commercially available norovirus diagnosis quick kits of 100-fold; the detection limit in spiked mussels however was ten-fold higher than in stool samples. Further assays showed a decrease in fluorescence intensity for heat- or UV-inactivated virus particles. Overall, we demonstrate the application of a flow cytometry approach for direct detection of small non-enveloped virus particles such as noroviruses. An adaptation of the technology to routine diagnostics has the potential to contribute a rapid and sensitive tool to norovirus outbreak investigations. Further improvements to the method, notably decreasing the detection limit of the approach, may allow the analysis of naturally contaminated food and environmental samples.
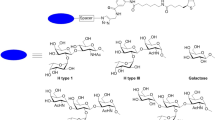
Graphic Abstract








Similar content being viewed by others
References
Arakelyan, A., Fitzgerald, W., Margolis, L., & Grivel, J.-C. (2013). Nanoparticle-based flow virometry for the analysis of individual virions. Journal of Clinical Investigation, 123(9), 3716–3727. https://doi.org/10.1172/JCI67042
Atmar, R. L., & Estes, M. K. (2006). The Epidemiologic and Clinical Importance of Norovirus Infection. Gastroenterology Clinics of North America, 35(2), 275–290. https://doi.org/10.1016/j.gtc.2006.03.001
Atmar, R. L., Opekun, A. R., Gilger, M. A., Estes, M. K., Crawford, S. E., Neill, F. H., & Graham, D. Y. (2008). Norwalk virus shedding after experimental human infection. Emerging Infectious Diseases, 14(10), 1553–1557. https://doi.org/10.3201/eid1410.080117
Brussaard, C. P. D., Marie, D., & Bratbak, G. (2000). Flow cytometric detection of viruses. Journal of Virological Methods, 85(1–2), 175–182. https://doi.org/10.1016/S0166-0934(99)00167-6
Chhabra, P., de Graaf, M., Parra, G. I., Chan, M. C. W., Green, K., Martella, V., et al. (2019). Updated classification of norovirus genogroups and genotypes. The Journal of General Virology, 100(10), 1393–1406. https://doi.org/10.1099/jgv.0.001318
Crawford, S. E., Ajami, N., Parker, T. D., Kitamoto, N., Natori, K., Takeda, N., Tanaka, T., Kou, B., Atmar, R. L., & Estes, M. K. (2015). Mapping broadly reactive norovirus genogroup i and II monoclonal antibodies. Clinical and Vaccine Immunology, 22(2), 168–177. https://doi.org/10.1128/CVI.00520-14
Devant, J., Hofhaus, G., & Hansman, G. S. (2019). Novel structural features of human norovirus capsid. bioRxiv. https://doi.org/10.1101/528240
Dolin, R., Blacklow, N. R., DuPont, H., Buscho, R. F., Wyatt, R. G., Kasel, J. A., Hornick, R., & Chanock, R. M. (1972). Biological properties of Norwalk agent of acute infectious nonbacterial gastroenteritis. Experimental Biology and Medicine, 140(2), 578–583. https://doi.org/10.3181/00379727-140-36508
El Bilali, N., Duron, J., Gingras, D., & Lippé, R. (2017). Quantitative Evaluation of Protein Heterogeneity within Herpes Simplex Virus 1 Particles. Journal of Virology. https://doi.org/10.1128/jvi.00320-17
Fuller, R. R., & Sweedler, J. V. (1996). Characterizing submicron vesicles with wavelength-resolved fluorescence in flow cytometry. Cytometry, 25(2), 144–155. https://doi.org/10.1002/(SICI)1097-0320(19961001)25:2%3c144::AID-CYTO3%3e3.0.CO;2-H
Glass, R. I., Parashar, U. D., & Estes, M. K. (2009). Norovirus gastroenteritis. New England Journal of Medicine, 361(18), 1776. https://doi.org/10.1056/NEJMra0804575
Gozalbo-Rovira, R., Rubio-Del-Campo, A., Santiso-Bellón, C., Vila-Vicent, S., Buesa, J., Delgado, S., Molinero, N., Margolles, A., Yebra, M. J., Collado, M. C., Monedero, V., & Rodríguez-Díaz, J. (2021). Interaction of Intestinal bacteria with human rotavirus during infection in children. International Journal of Molecular Sciences, 22(3), 1010. https://doi.org/10.3390/ijms22031010
Hansman, G. S., Taylor, D. W., McLellan, J. S., Smith, T. J., Georgiev, I., Tame, J. R. H., Park, S. Y., Yamazaki, M., Gondaira, F., Miki, M., Katayama, K., Murata, K., & Kwong, P. D. (2012). Structural basis for broad detection of genogroup II noroviruses by a monoclonal antibody that binds to a site occluded in the viral particle. Journal of Virology, 86(7), 3635–3646. https://doi.org/10.1128/jvi.06868-11
Hardy, M. E. (2005). Norovirus protein structure and function. FEMS Microbiology Letters, 253(1), 1–8. https://doi.org/10.1016/J.FEMSLE.2005.08.031
Hassard, F., Sharp, J. H., Taft, H., LeVay, L., Harris, J. P., McDonald, J. E., Tuson, K., Wilson, J., Jones, D. L., & Malham, S. K. (2017). Critical review on the public health impact of norovirus contamination in shellfish and the environment: A UK perspective. Food and Environmental Virology, 9(2), 123–141. https://doi.org/10.1007/s12560-017-9279-3
Huang, W., Samanta, M., Crawford, S. E., Estes, M. K., Neill, F. H., Atmar, R. L., & Palzkill, T. (2014). Identification of human single-chain antibodies with broad reactivity for noroviruses. Protein engineering design and selection (Vol. 27, pp. 339–349). Oxford University Press.
Iannelli, D. (1996). Cytofluorimetric method for the detection of the cucumber mosaic virus. Phytopathology, 86(9), 959. https://doi.org/10.1094/Phyto-86-959
Iannelli, D., D’Apice, L., Cottone, C., Viscardi, M., Scala, F., Zoina, A., Del Sorbo, G., Spigno, P., & Capparelli, R. (1997). Simultaneous detection of cucumber mosaic virus, tomato mosaic virus and potato virus Y by flow cytometry. Journal of Virological Methods, 69(1–2), 137–145. https://doi.org/10.1016/S0166-0934(97)00149-3
Jiang, X., Wang, M., Graham, D. Y., & Estes, M. K. (1992). Expression, self-assembly, and antigenicity of the Norwalk virus capsid protein. Journal of Virology, 66(11), 6527–6532. https://doi.org/10.1128/jvi.66.11.6527-6532.1992
Jung, J., Grant, T., Thomas, D. R., Diehnelt, C. W., Grigorieff, N., & Joshua-Tor, L. (2019). High-resolution cryo-EM structures of outbreak strain human norovirus shells reveal size variations. Proceedings of the National Academy of Sciences of the United States of America, 116(26), 12828–12832. https://doi.org/10.1073/pnas.1903562116
Kapikian, A. Z., Wyatt, R. G., Dolin, R., Thornhill, T. S., Kalica, A. R., & Chanock, R. M. (1972). Visualization by immune electron microscopy of a 27-nm particle associated with acute infectious nonbacterial gastroenteritis. Journal of virology, 10(5), 1075–81. http://www.ncbi.nlm.nih.gov/pubmed/4117963. Accessed 30 July 2018
Katpally, U., Wobus, C. E., Dryden, K., Virgin, H. W., & Smith, T. J. (2008). Structure of antibody-neutralized murine norovirus and unexpected differences from viruslike particles. Journal of Virology, 82(5), 2079–2088. https://doi.org/10.1128/jvi.02200-07
Khamrin, P., Thongprachum, A., Takanashi, S., Okitsu, S., Maneekarn, N., Hayakawa, S., & Ushijima, H. (2015). Evaluation of immunochromatography tests for detection of novel GII.17 norovirus in stool samples. Eurosurveillance. https://doi.org/10.2807/1560-7917.ES2015.20.28.21185
Kim, B. C., Ju, M. K., Dan-Chin-Yu, A., & Sommer, P. (2009). Quantitative detection of HIV-1 particles using HIV-1 neutralizing antibody-conjugated beads. Analytical Chemistry, 81(6), 2388–2393. https://doi.org/10.1021/ac802267u
Kolawole, A. O., Xia, C., Li, M., Gamez, M., Yu, C., Rippinger, C. M., Yucha, R. E., Smith, T. J., & Wobus, C. E. (2014). Newly isolated mAbs broaden the neutralizing epitope in murine norovirus. Journal of General Virology, 95(Pt 9), 1958–1968. https://doi.org/10.1099/vir.0.066753-0
Lambden, P. R., Caul, E. O., Ashley, C. R., & Clarke, I. N. (1993). Sequence and genome organization of a human small round-structured (Norwalk-like) virus. Science, 259(5094), 516–519. https://doi.org/10.1126/science.8380940
Le Guyader, F. S., Parnaudeau, S., Schaeffer, J., Bosch, A., Loisy, F., Pommepuy, M., & Atmar, R. L. (2009). Detection and quantification of noroviruses in shellfish. Applied and Environmental Microbiology, 75(3), 618–624. https://doi.org/10.1128/AEM.01507-08
Li, X., Zhou, R., Tian, X., Li, H., & Zhou, Z. (2010). Characterization of a cross-reactive monoclonal antibody against Norovirus genogroups I, II, III and V. Virus Research, 151(2), 142–147. https://doi.org/10.1016/J.VIRUSRES.2010.04.005
Li, X., Zhou, R., Wang, Y., Sheng, H., Tian, X., Li, H., & Qiu, H. (2009). Identification and characterization of a native epitope common to norovirus strains GII/4, GII/7 and GII/8. Virus Research, 140(1–2), 188–193. https://doi.org/10.1016/J.VIRUSRES.2008.12.004
Loisy, F., Atmar, R. L., Guillon, P., Le Cann, P., Pommepuy, M., & Le Guyader, F. S. (2005). Real-time RT-PCR for norovirus screening in shellfish. Journal of Virological Methods, 123(1), 1–7. https://doi.org/10.1016/j.jviromet.2004.08.023
Loret, S., El Bilali, N., & Lippé, R. (2012). Analysis of herpes simplex virus type I nuclear particles by flow cytometry. Cytometry Part A, 81A(11), 950–959. https://doi.org/10.1002/cyto.a.22107
Ludwig-Begall, L. F., Mauroy, A., & Thiry, E. (2018). Norovirus recombinants: Recurrent in the field, recalcitrant in the lab-a scoping review of recombination and recombinant types of noroviruses. The Journal of General Virology, 99(8), 970–988. https://doi.org/10.1099/jgv.0.001103
Madrigal, J. L., & Jones, M. K. (2020). Quantifying human norovirus virus-like particles binding to commensal bacteria using flow cytometry. Journal of Visualized Experiments, 2020(158), e61048. https://doi.org/10.3791/61048
Mallory, M. L., Lindesmith, L. C., Graham, R. L., & Baric, R. S. (2019). GII.4 human norovirus: Surveying the antigenic landscape. Viruses. https://doi.org/10.3390/v11020177
Mead, P. S., Slutsker, L., Dietz, V., McCaig, L. F., Bresee, J. S., Shapiro, C., Griffin, P. M., & Tauxe, R. V. (1999). Food-related illness and death in the United States. Emerging Infectious Diseases, 5(5), 607–625. https://doi.org/10.3201/eid0505.990502
Parker, T. D., Kitamoto, N., Tanaka, T., Hutson, A. M., & Estes, M. K. (2005). Identification of Genogroup I and Genogroup II broadly reactive epitopes on the norovirus capsid. Journal of Virology, 79(12), 7402–7409. https://doi.org/10.1128/jvi.79.12.7402-7409.2005
Parra, G. I., Azure, J., Fischer, R., Bok, K., Sandoval-Jaime, C., Sosnovtsev, S. V., Sander, P., & Green, K. Y. (2013). Identification of a broadly cross-reactive epitope in the inner shell of the norovirus capsid. PLoS ONE, 8(6), e67592. https://doi.org/10.1371/journal.pone.0067592
Prasad, B. V. V., Hardy, M. E., Dokland, T., Bella, J., Rossmann, M. G., & Estes, M. K. (1999). X-ray crystallographic structure of the Norwalk virus capsid. Science, 286(5438), 287–290. https://doi.org/10.1126/science.286.5438.287
Razafimahefa, R. M., Ludwig-Begall, L. F., Le Guyader, F. S., Farnir, F., Mauroy, A., & Thiry, E. (2021). Optimisation of a PMAxxTM-RT-qPCR assay and the preceding extraction method to selectively detect infectious murine norovirus particles in mussels. Food and Environmental Virology, 13, 93–106. https://doi.org/10.1007/s12560-020-09454-w
Reed, L. J., & Muench, H. (1938). A simple method of estimating fifty per cent endpoints12. American Journal of Epidemiology, 27(3), 493–497. https://doi.org/10.1093/oxfordjournals.aje.a118408
Ruis, C., Roy, S., Brown, J. R., Allen, D. J., Goldstein, R. A., & Breuer, J. (2017). The emerging GII.P16-GII.4 Sydney 2012 norovirus lineage is circulating worldwide, arose by late-2014 and contains polymerase changes that may increase virus transmission. PLoS ONE. https://doi.org/10.1371/journal.pone.0179572
Schindelin, J., Arganda-Carreras, I., Frise, E., Kaynig, V., Longair, M., Pietzsch, T., Preibisch, S., Rueden, C., Saalfeld, S., Schmid, B., Tinevez, J. Y., White, D. J., Hartenstein, V., Eliceiri, K., Tomancak, P., & Cardona, A. (2012). Fiji: An open-source platform for biological-image analysis. Nature Methods, 9(7), 676–682. https://doi.org/10.1038/nmeth.2019
Siebenga, J. J., Vennema, H., Zheng, D. P., Vinjé, J., Lee, B. E., Pang, X. L., Ho, E. C., Lim, W., Choudekar, A., Broor, S., Halperin, T., Rasool, N. B., Hewitt, J., Greening, G. E., Jin, M., Duan, Z. J., Lucero, Y., O’Ryan, M., Hoehne, M., & Koopmans, M. (2009). Norovirus illness is a global problem: Emergence and spread of norovirus GII.4 variants, 2001–2007. The Journal of Infectious Diseases, 200(5), 802–812. https://doi.org/10.1086/605127
Smith, H. Q., & Smith, T. J. (2019). The dynamic capsid structures of the noroviruses. Viruses. https://doi.org/10.3390/v11030235
Song, C., Takai-Todaka, R., Miki, M., Haga, K., Fujimoto, A., Ishiyama, R., Oikawa, K., Yokoyama, M., Miyazaki, N., Iwasaki, K., Murakami, K., Katayama, K., & Murata, K. (2020). Dynamic rotation of the protruding domain enhances the infectivity of norovirus. PLoS Pathogens. https://doi.org/10.1371/journal.ppat.1008619
Stals, A., Baert, L., Coillie, E. V., & Uyttendaele, M. (2012). Extraction of food-borne viruses from food samples: A review. International Journal of Food Microbiology, 153(1–2), 1–9. https://doi.org/10.1016/j.ijfoodmicro.2011.10.014
Steen, H. B. (2004). Flow cytometer for measurement of the light scattering of viral and other submicroscopic particles. Cytometry, 57A(2), 94–99. https://doi.org/10.1002/cyto.a.10115
Svraka, S., Duizer, E., Vennema, H., De Bruin, E., Van Der Veer, B., Dorresteijn, B., & Koopmans, M. (2007). Etiological role of viruses in outbreaks of acute gastroenteritis in The Netherlands from 1994 through 2005. Journal of Clinical Microbiology, 45(5), 1389–1394. https://doi.org/10.1128/JCM.02305-06
Tan, M., Fang, P., Chachiyo, T., Xia, M., Huang, P., Fang, Z., Jiang, W., & Jiang, X. (2008). Noroviral P particle: Structure, function and applications in virus-host interaction. Virology, 382(1), 115–123. https://doi.org/10.1016/j.virol.2008.08.047
Tang, V. A., Renner, T. M., Varette, O., Le Boeuf, F., Wang, J., Diallo, J. S., Bell, J. C., & Langlois, M. A. (2016). Single-particle characterization of oncolytic vaccinia virus by flow virometry. Vaccine, 34(42), 5082–5089. https://doi.org/10.1016/j.vaccine.2016.08.074
Thackray, L. B., Wobus, C. E., Chachu, K. A., Liu, B., Alegre, E. R., Henderson, K. S., Kelley, S. T., & Virgin, H. W. (2007). Murine noroviruses comprising a single genogroup exhibit biological diversity despite limited sequence divergence. Journal of virology, 81(19), 10460–73. https://doi.org/10.1128/JVI.00783-07
Tracy, B. P., Gaida, S. M., & Papoutsakis, E. T. (2010). Flow cytometry for bacteria: Enabling metabolic engineering, synthetic biology and the elucidation of complex phenotypes. Current Opinion in Biotechnology, 21(1), 85–99. https://doi.org/10.1016/j.copbio.2010.02.006
Ushijima, H., Thongprachum, A., Khamrin, P., Takanashi, S., Okitsu, S., Maneekarn, N., & Hayakawa, S. (2017). Evaluation of immunochromatographic tests and a new enzyme immunoassay for detection of a novel GII.17 norovirus in stool samples. Japanese Journal of Infectious Diseases, 70(3), 326–328. https://doi.org/10.7883/yoken.JJID.2016.413
Vongpunsawad, S., Venkataram Prasad, B. V., & Estes, M. K. (2013). Norwalk virus minor capsid protein VP2 associates within the VP1 shell domain. Journal of Virology, 87(9), 4818–4825. https://doi.org/10.1128/JVI.03508-12
Vorauer-Uhl, K., Wagner, A., Borth, N., & Katinger, H. (2000). Determination of liposome size distribution by flow cytometry. Cytometry, 39(2), 166–171. https://doi.org/10.1002/(SICI)1097-0320(20000201)39:2%3c166::AID-CYTO10%3e3.0.CO;2-M
Yan, X., Schielke, E. G., Grace, K. M., Hassell, C., Marrone, B. L., & Nolan, J. P. (2004). Microsphere-based duplexed immunoassay for influenza virus typing by flow cytometry. Journal of Immunological Methods, 284(1–2), 27–38. https://doi.org/10.1016/j.jim.2003.09.016
Yan, X., Zhong, W., Tang, A., Schielke, E. G., Hang, W., & Nolan, J. P. (2005). Multiplexed flow cytometric immunoassay for influenza virus detection and differentiation protein detection. A Four-Plexed Assay for Influenza Virus Was Developed to Demonstrate the Potential for Multi-, 77(23), 7673–7678.
Yang, S.-Y., Lien, K.-Y., Huang, K.-J., Lei, H.-Y., & Lee, G.-B. (2008). Micro flow cytometry utilizing a magnetic bead-based immunoassay for rapid virus detection. Biosensors and Bioelectronics, 24, 855–862. https://doi.org/10.1016/j.bios.2008.07.019
Yoda, T., Suzuki, Y., Terano, Y., Yamazaki, K., Sakon, N., Kuzuguchi, T., Oda, H., & Tsukamoto, T. (2003). Precise characterization of norovirus (Norwalk-like virus)-specific monoclonal antibodies with broad reactivity. Journal of Clinical Microbiology, 41(6), 2367–2371. https://doi.org/10.1128/JCM.41.6.2367-2371.2003
Zheng, L., Wang, W., Liu, J., Chen, X., Li, S., Wang, Q., Huo, Y., Qin, C., Shen, S., & Wang, M. (2018). Characterization of a Norovirus-specific monoclonal antibody that exhibits wide spectrum binding activities. Journal of Medical Virology, 90(4), 671–676. https://doi.org/10.1002/jmv.25001
Zicari, S., Arakelyan, A., Fitzgerald, W., Zaitseva, E., Chernomordik, L. V., Margolis, L., & Grivel, J. C. (2016). Evaluation of the maturation of individual Dengue virions with flow virometry. Virology, 488, 20–27. https://doi.org/10.1016/j.virol.2015.10.021
Acknowledgements
We thank Dr Pascale Huynen (Liège CHU, Belgium), Dr Bavo Verhaegen and Dr Katelijne Dierick (Sciensano, Brussels, Belgium) for providing clinical samples. This project was supported by a Grant from the Service Public Fédéral ‘Santé Publique, Sécurité de la Chaîne alimentaire et Environnement’ (RT15/8 IQUINOR2) (R.M.R, L. F. L.-B., E. T.).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12560_2021_9494_MOESM1_ESM.jpg
Supplementary file1 GI.3 human norovirus (HuNoV) detection in faecal suspension by flow cytometry. Gating strategy for flow cytometry analysis: forward versus side scatter (SSC vs FSC) gating (dot plots), the graphs on the right for each condition (histogram) show the fluorescence profiles. (A) Negative control. (B) GI.3 diluted 1:10, (C) 1:100, (D) 1:1000, (E) 1: 10000, (F) 1:100000, (G) 1:1000000 (JPG 10838 kb)
12560_2021_9494_MOESM2_ESM.jpg
Supplementary file2 Non-inactivated or inactivated GI.3 human norovirus (HuNoV) detection in faecal suspension by flow cytometry. Gating strategy for flow cytometry analysis: forward versus side scatter (SSC vs FSC) gating (dot plots), the graphs on the right for each condition (histogram) show the fluorescence profiles. (H) Negative control: PBS. (I) Non-inactivated GI.3. (J) Heat-inactivated GI.3. (K) UV-inactivated GI.3 (JPG 6619 kb)
12560_2021_9494_MOESM3_ESM.jpg
Supplementary file3 GI.3 human norovirus (HuNoV) detection in spiked mussel DTs by flow cytometry. Gating strategy for flow cytometry analysis: forward versus side scatter (SSC vs FSC) gating (dot plots), the graphs on the right for each condition (histogram) show the fluorescence profiles. (A) Negative control: PBS. (B) Negative control: negative mussel DTs supernatant. (C) GI.3 diluted 1:10, (D) 1:100, (E) 1:1000, (F) 1: 10000, (G) 1:100000, (H) 1:1000000 (JPG 12771 kb)
12560_2021_9494_MOESM4_ESM.jpg
Supplementary file4 Non-inactivated or inactivated GI.3 human norovirus (HuNoV) detection in spiked mussel DTs by flow cytometry. Gating strategy for flow cytometry analysis: forward versus side scatter (SSC vs FSC) gating (dot plots), the graphs on the right for each condition (histogram) show the fluorescence profiles. (I) Negative control: PBS. (J) Negative control: negative mussel DTs supernatant. (K) Non-inactivated GI.3. (L) Heat-inactivated GI.3. (M) UV-inactivated GI.3 (JPG 7899 kb)
Rights and permissions
About this article
Cite this article
Razafimahefa, R.M., Ludwig-Begall, L.F., Diallo, M.A. et al. Development of a Specific Anti-capsid Antibody- and Magnetic Bead-Based Immunoassay to Detect Human Norovirus Particles in Stool Samples and Spiked Mussels via Flow Cytometry. Food Environ Virol 13, 493–506 (2021). https://doi.org/10.1007/s12560-021-09494-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12560-021-09494-w