Abstract
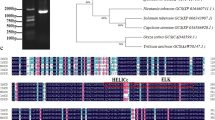
Myo-inositol oxygenase (MIOX), the only catabolic enzyme of the inositol pathway, catalyzes conversion of myo-inositol to D-GlcA (glucuronic acid). The present study encompasses bioinformatic analysis of MIOX gene across phylogenetically related plant lineages and representative animal groups. Comparative motif analysis of the MIOX gene(s) across various plant groups suggested existence of abiotic- stress related cis-acting elements such as, DRE, MYB, MYC, STRE, MeJa among others. A detailed analysis revealed a single isoform of MIOX gene, located in chromosome 6 of indica rice (Oryza sativa) with an open reading frame of 938 bp coding for 308 amino acids producing a protein of ~ 35 kD. Secondary structure prediction of the protein gave the predicted number of 144 alpha helices and 154 random coils. The three-dimensional structure suggested it to be a monomeric protein with a single domain. Bacterial overexpression of the protein, purification and enzyme assay showed optimal catalytic activity at pH 7.5–8 at an optimal temperature of 37 °C with Michaelis constant of 40.92 mM. The range of Km was determined as 22.74–28.7 mM and the range of Vmax was calculated as 3.51–3.6 µM/min, respectively. Four salt-tolerant and salt-sensitive rice cultivars displayed differential gene expression of OsMIOX at different time points in different tissues under salinity and drought stress as observed from qRT-PCR data, microarray results and protein expression profile in immunoblot analysis. Gel volumetric analysis confirmed a very high expression of MIOX in roots and leaves on 7th day following germination. Microarray data showed high expression of MIOX at all developmental stages including seedling growth and reproduction. These data suggest that OsMIOX might have a role to play in rice abiotic stress responses mediated through the myo-inositol oxidation pathway.










Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
Abbreviations
- CRE:
-
Cis Regulatory element
- DRE:
-
Dehydration responsive element
- MYB:
-
Myeloblastosis
- MYC:
-
Master regulator of cell cycle entry and proliferative metabolism
- STRE:
-
Stress responsive element
- MeJa:
-
Methyl jasmonate
References
Adak S, Roy A, Das P, Mukherjee A, Sengupta S, Majumder AL (2019) Soil salinity and mechanical obstruction differentially affects embryonic root architecture in different rice genotypes from West Bengal. Plant Physiol Rep 24:192–209
Alam MNU, Jewel GMNA, Azim T, Seraj ZI (2021) Novel QTLs for salinity tolerance revealed by genome-wide association studies of biomass, chlorophyll and tissue ion content in 176 rice landraces from Bangladesh. PLoS ONE 16:e0259456
Alford SR, Rangarajan P, Williams P, Gillaspy GE (2012) Myo-Inositol oxygenase is required for responses to low energy conditions in Arabidopsis thaliana. Front Plant Sci 3:69
Alok A, Kaur H, Bhati KK, Kumar J, Pandey P, Upadhay SK, Pandey A, Sharma NC, Pandey AK, Tiwari S (2015) Biochemical characterization and spatio-temporal expression of myo-inositol oxygenase (MIOX) from wheat (Triticum aestivum L.). Plant Gene 4:10–19
Ambawat S, Sharma P, Yadav NR, Yadav RC (2013) MYB transcription factor genes as regulators for plant responses: an overview. Physiol Mol Biol Plants 19:307–321
Antosch M, Mortensen SA, Grasser KD (2012) Plant proteins containing high mobility group box DNA-binding domains modulate different nuclear processes. Plant Physiol 159:875–883
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Charalampous FC (1958) Biochemical studies on inositol: V. Purification and properties of the enzyme that cleaves inositol to D-Glucuronic acid. J Biol Chem 234:220–227
Charalampous FC, Lyras C (1957) Biochemical studies on inositol: IV. Conversion of inositol to glucuronic acid by rat kidney extracts. J Biol Chem 228:1–13
Dietz KJ, Vogel MO, Viehhauser A (2010) AP2/EREBP transcription factors are part of gene regulatory networks and integrate metabolic, hormonal and environmental signals in stress acclimation and retrograde signalling. Protoplasma 245:3–14
Doherty CJ, Van Buskirk HA, Myers SJ, Thomashow MF (2009) Roles for Arabidopsis CAMTA transcription factors in cold-regulated gene expression and freezing tolerance. Plant Cell 21:972–984
Duan J, Zhang M, Zhang H, Xiong H, Liu P, Ali J, Li J, Li Z (2012) OsMIOX, a myo-inositol oxygenase gene, improves drought tolerance through scavenging of reactive oxygen species in rice (Oryza sativa L.). Plant Sci 196:143–151
Endres S, Tenhaken R (2009) Myo inositol oxygenase controls the level of myo inositol in Arabidopsis, but does not increase ascorbic acid. Plant Physiol 149:1042–1049
Fan M, Xu C, Xu K, Hu Y (2012) LATERAL ORGAN BOUNDARIES DOMAIN transcription factors direct callus formation in Arabidopsis regeneration. Cell Res 22:1169–1180
Guo J, Sun B, He H, Zhang Y, Tian H, Wang B (2021) Current understanding of bHLH transcription factors in plant abiotic stress tolerance. Int J Mol Sci 22:4921
İnal B, Büyük İ, İlhan E, Aras S (2017) Genome-wide analysis of Phaseolus vulgaris C2C2-YABBY transcription factors under salt stress conditions. 3 Biotech 7:302
Jakoby M, Weisshaar B, Dröge-Laser W, Vicente-Carbajosa J, Tiedemann J, Kroj T, Parcy F (2002) bZIP transcription factors in Arabidopsis. Trends Plant Sci 7:106–111
Kanter U, Becker M, Friauf E, Tenhaken R (2003) Purification, characterization and functional cloning of inositol oxygenase from Cryptococcus. Yeast 20:1317–1329
Kanter U, Usadel B, Guerineau F, Li Y, Pauly M, Tenhaken R (2005) The inositol oxygenase gene family of Arabidopsis is involved in the biosynthesis of nucleotide sugar precursors for cell-wall matrix polysaccharides. Planta 221:243–254
Kaur A, Pati PK, Pati AM, Nagpal AK (2017) In-silico analysis of cis-acting regulatory elements of pathogenesis-related proteins of Arabidopsis thaliana and Oryza sativa. PLoS ONE 12:e0184523
Koller E, Koller F, Hoffmann-Ostenhof O (1976) Myo-inositol oxygenase from oat seedlings. Mol Cell Biochem 10:33–39
Li S (2015) The Arabidopsis thaliana TCP transcription factors: a broadening horizon beyond development. Plant Signal Behav 10:e1044192
Li Y, He L, Li J, Chen J, Liu C (2018) Genome-wide identification, characterization, and expression profiling of the legume BZR transcription factor gene family. Front Plant Sci 9:1332
Li Z, Liu Z, Wei Y, Liu Y, Xing L, Liu M, Li P, Lu Q, Peng R (2021) Genome-wide identification of the MIOX gene family and their expression profile in cotton development and response to abiotic stress. PLoS ONE 16:e0254111. https://doi.org/10.1371/journal.pone.0254111
Loewus FA, Loewus MW (1983) Myo-Inositol: its biosynthesis and metabolism. Annu Rev Plant Physiol 34:137–161
Lorence A, Chevone BI, Mendes P, Nessler CL (2004) Myo-Inositol oxygenase offers a possible entry point into plant ascorbate biosynthesis. Plant Physiol 134:1200–1205
Manoli A, Trevisan S, Quaggiotti S, Varotto S (2018) Identification and characterization of the BZR transcription factor family and its expression in response to abiotic stresses in Zea mays L. Plant Growth Regul 84:423–436
Moon TS, Yoon SH, Lanza AM, Roy-Mayhew JD, Prather KL (2009) Production of glucaric acid from a synthetic pathway in recombinant Escherichia coli. Appl Environ Microbiol 75:589–595
Mukherjee R, Mukherjee A, Bandyopadhyay S, Mukherjee S, Sengupta S, Ray S, Majumder AL (2019) Selective manipulation of the inositol metabolic pathway for induction of salt-tolerance in indica rice variety. Sci Rep 9:5358
Munir S, Mumtaz MA, Ahiakpa JK et al (2020) Genome-wide analysis of Myo-inositol oxygenase gene family in tomato reveals their involvement in ascorbic acid accumulation. BMC Genomics 21:284
Nepal N, Yactayo-Chang JP, Medina-Jiménez K, Acosta-Gamboa LM, González-Romero ME, Arteaga-Vázquez MA, Lorence A (2019) Mechanisms underlying the enhanced biomass and abiotic stress tolerance phenotype of an Arabidopsis MIOX over-expresser. Plant Direct 3:e00165
Noctor G, Foyer CH (1998) Ascorbate and glutathione: keeping active oxygen under control. Annu Rev Plant Physiol Plant Mol Biol 49:249–279
Quesada V (2016) The roles of mitochondrial transcription termination factors (MTERFs) in plants. Physiol Plant 157:389–399
Reddy CC, Swan JS, Hamilton GA (1981) Myo-Inositol oxygenase from hog kidney. J Biol Chem 256:8510–8518
Shi F, Dong Y, Wang M, Qiu D (2020) Transcriptomics analyses reveal that OsMIOX improves rice drought tolerance by regulating the expression of plant hormone and sugar related genes. Plant Biotechnol Rep 14:339–349
Tamura K, Nei M (1993) Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol 10:512–526
Varland S, Osberg C, Arnesen T (2015) N-terminal modifications of cellular proteins: the enzymes involved, their substrate specificities and biological effects. Proteomics 15:2385–2401
Wang X, Niu Y, Zheng Y (2021) Multiple functions of MYB transcription factors in abiotic stress responses. Int J Mol Sci 22:6125
Xie Z, Nolan TM, Jiang H, Yin Y (2019) AP2/ERF transcription factor regulatory networks in hormone and abiotic stress responses in Arabidopsis. Front Plant Sci 10:228
Xing H, Pudake RN, Guo G, Xing G, Hu Z, Zhang Y, Sun Q, Ni Z (2011) Genome-wide identification and expression profiling of auxin response factor (ARF) gene family in maize. BMC Genomics 12:178
Yang J, Wang Y, Zhang Y (2015) ResQ: an approach to unified estimation of B-factor and residue-specific error in protein structure prediction. J Mol Biol 4:693–701
Ye W, Ren W, Kong L, Zhang W, Wang T (2016) Transcriptomic profiling analysis of Arabidopsis thaliana treated with exogenous Myo-inositol. PLoS ONE 11:e0161949
Zhang T, Lv W, Zhang H, Ma L, Li P, Ge L, Li G (2018a) Genome-wide analysis of the basic Helix-Loop-Helix (bHLH) transcription factor family in maize. BMC Plant Biol 18:235
Zhang T, Zhao Y, Wang Y, Liu Z, Gao C (2018b) Comprehensive analysis of MYB gene family and their expressions under abiotic stresses and hormone treatments in Tamarix hispida front. Plant Sci 9:1303
Acknowledgements
The work is supported by grants from Department of Biotechnology, GOI (BT/AB/05/02/2007-III dt.21/09/2010 and BT/IN/NWO/17/ALM dated 02.09.2015) awarded to ALM, while an INSA Senior Scientist. The research is also supported by grants from, Department of Biotechnology to SR [WBDBT Sanction No.237(Sanc.)/BT(Estt.)/RD-28/2016 dated 29-03-2017]. SA is funded by Govt. of India University Grants Commission-Rajiv Gandhi National Fellowship (2011-12/RGNF-SC-WES-13271). PD is funded by Department of Science and Technology-SERB research grant (YSS/2015/001872).
Author information
Authors and Affiliations
Contributions
ALM, SA and SR designed the research. SA, TA and PD conducted the experiments. SA, TA, PD and SR prepared the draft manuscript. ALM finalized the manuscript with input and approval from all the authors.
Corresponding author
Ethics declarations
Conflict of interest
All authors have approved and there is no conflict of interest in the submission of this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Adak, S., Agarwal, T., Das, P. et al. Characterization of myo-inositol oxygenase from rice (OsMIOX): influence of salinity stress in different indica rice cultivars. Physiol Mol Biol Plants 29, 927–945 (2023). https://doi.org/10.1007/s12298-023-01340-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-023-01340-6