Abstract
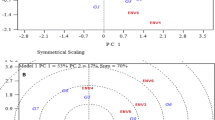
Among the tropical tuber crops, cassava (Manihot esculenta Crantz) deserves special attention as regards to its higher biological efficiency in terms of dry matter production which incidentally implies to the higher amount of nutrient extraction from the soil resulting in better response to the application of manures and fertilizers. Among the major nutrients, Potassium (K) is considered as the key nutrient for cassava owing to its influence both in tuber yield and tuber quality. The above facts as well as the availability of sufficient cassava genotypes in the germplasm collection of ICAR-CTCRI made us to initiate research work to screen cassava germplasm including the pre breeding lines. The objective being to identify K efficient genotypes which can yield well under limited availability of K so that the external application of K can be reduced. This paper describes the wide variation noticed during the pre evaluation of 83 elite genotypes which was done as a prelude in the screening and identification of K efficient genotypes. The characters studied were tuber yield, tuber characters, plant dry matter percentage, plant K content, tuber quality (starch, cyanogenic glucosides) attributes, physiological efficiency and plant biometric characters. The variation among the genotypes for the above traits was assessed by making some yardstick for classification which in turn helped in determining the percent distribution of genotypes in each category. The variation among the genotypes were further affirmed through principal component analysis, wherein the first five components explained more than 77% of variability and the cluster analysis performed grouped these genotypes into five clusters. The biplot showed the traits which are closely linked to the genotypes. The dendrogram constructed indicated similar genotypes to that of the clusters to the extent of more than 50% revealing the association of members with similar traits in clusters and dendrograms. The study helped in establishing the drastic variation among the genotypes along with identification of six genotypes viz., Aniyoor, 7 Sahya (2), 7 III E3-5, W-19, CR 43-8, 6-6 for further detailed experimentation to identify K efficient genotypes.








Similar content being viewed by others
References
Agong SG, Schittenhelm S, Friedt W (2001) Genotypic variation of Kenyan tomato (Lycopersiconesculentum L.) germplasm. J Food Tech Africa 6(1):13–17. https://doi.org/10.4314/jfta.v6i1.19277
AOAC (1984) Official methods of analysis, 14th edn. Association of Analytical Chemists, Washington DC
Baligar VC, Fageria NK (1997) Nutrient use efficiency in acid soils: nutrient management and plant use efficiency. In: Monitz AC, Furlani AMC, Fageria NK, Rosolem CA, Cantarells H (eds) Plant-soil interactions at low pH: sustainable agriculture and forestry production. Brazilian Soil Science Society Campinas, Brazil, pp 75–93
Ceballos H, Perez JC, Orlando JB, Lenis JI, Morante N, Calle F, Pino L, Hershey H (2016) Cassava breeding I: the value of breeding value. Front Plant Sci. https://doi.org/10.3389/fpls.2016.01227
Chowdhury J, Karim M, Khaliq Q, Solaiman A, Ahmed J (2016) Genotypic variations in growth, yield and yield components of soybean genotypes under drought stress conditions. Bangladesh J Agric Res 40(4):537–550. https://doi.org/10.3329/bjar.v40i4.26929
Clark RB, Duncan RR (1991) Improvement of plant mineral nutrition through breeding. Field Crops Res 27:219–240
Crossa J, Delacy IH, Taba S (1995) The use of multivariate methods in developing a core collection. In: Hodgkin T, Brown AHD, Van Hintum TJL, Morales EAV (eds) Core collections of plant genetic resources. Wiley, Chichester, pp 77–92
Daehler CC, Yorkston M, DudleyN SW (1999) Genetic variation in morphology and growth characters of Acacia koa in the Hawaiian Islands. Int J Plant Sci 160(4):767–773
Dar TH, Shakeel R, Verma S (2018) Comparative germplasm characterization of maize (Zea mays L.) in Rajouri Region of PirPanjal Himalaya J & K (India), based on morphological and ISSR markers. J Crop Sci Biotechnol 21:43–55. https://doi.org/10.1007/s12892-017-0128-0
Duncan RR, Carrow RN (1999) Turfgrass molecular genetic improvement for abiotic/edaphic stress environment. Adv Agron 67:233–306
Edison S, Sheela MN (2008) Diversity of tuber crops and wild relatives. In: Kannaiyan S (ed) Agrobiodiversity hotspots and access and benefit sharing. Narosa Publishing House, New Delhi, pp 241–248
Graham RD (1984) Breeding for nutritional characteristics in cereals. In: Tinker PB, Lauchi A (eds) Advances in plant nutrition. Praiger Publisher, New York, pp 57–102
Hahn SK, Terry ER, Leuschner K (1980) Breeding cassava for resistance to cassava mosaic disease. Euphytica 29:673–683
Heitjan DF, Roderick JAL (1991) Multiple imputation for the fatal accident reporting system. J R Stat Soc C Appl 40(1):13–29. https://doi.org/10.2307/2347902
Hughes AN, Louise A, Arsene IZB, Konan KC (2015) Phenotypic diversity of farmers’s traditional maize (Zea mays L) varieties in Cote d’ Ivoire. Maydica 60(2):112–118
Iqbal A, Gui H, Zhang H, Wang X, Pang N, Dong Q, Song M (2019) Genotypic variation in cotton genotypes for phosphorus use efficiency. Agron 9(11):689
Isfan D (1990) Nitrogen physiological efficiency index in some selected spring barley cultivars. J Plant Nutr 13(8):907–914
Jordan WR, Miller FR, Morris DE (1979) Genetic variation in root and shoot growth of sorghum in hydroponics. Crop Sci 19:468–472
Kouam EB, Dongmo JR, Djeugap JF (2018) Exploring morphological variation in tomato (Solanumlycopersicum): acombined study of disease resistance, genetic divergence and association of characters. Agricultura Tropica Et Subtropica 51(2):71–82
Lopez M, El-Dahan MAA, Leidi EO (2008) Genotypic variation in potassium uptake in dryland cotton. J Plant Nutr 31(11):1947–1962
Malik AI, Kongsil P, Nguyen VA, Sholihin WO, Srean P, Sheela MN, López-Lavalle LAB, Utsumi Y, Lu C, Kittipadakul P, Nguyen HH, Ceballos H, Nguyen TH, Selvaraj M, Aiemnaka GP, Labarta R, Chen S, Amawan S, Sok S, Youabee L, Seki M, Tokunaga H, Wang W, Li K, Nguyen HA, Van Dong N, Ham LH, Ishitani M (2020) Cassava breeding and agronomy in Asia: 50 years of history and future directions. Breed Sci 70(2):145–166
McAdam EL, Vaillancourt RE, Koutoulis A, Whittock SP (2014) Quantitative genetic parameters for yield, plant growth and cone chemical traits in hop (Humuluslupulus L.). BMC Genet 15:22. https://doi.org/10.1186/1471-2156-15-22
More SJ, Ravi V, Saravanan R (2019) Tropical tuber crops. In: de Freitas ST, Pareek S (eds) Postharvest physiological disorders in fruits and vegetables. CRC Press, Boca Raton, pp 719–757. https://doi.org/10.1201/b22001
Otayk SM (2019) Evaluation of agronomic traits and assessment of genetic variability in some popular wheat genotypes cultivated in Saudi Arabia. Aust J Crop Sci 13(6):847–856
Padmaja G, Moorthy SN, Bala N, Lila B, Sundaresan S, Sajeev MS, Nanda SK, Susan John K, Rajalekshmy L, Sudha Devi KS, Manikantan NM (2005) Analytical methodologies for tropical tuber crops, laboratory manual series-2. ICAR-Central Tuber Crops Research Institute, Thiruvananthapuram, p 119
Pahadi P, Sapkota M, Thapa DB, Pradhan S (2017) Cluster and principal component analysis for the selection of maize (Zea mays L.) genotypes. Int J Exp Res Rev 9:5–10
Rahim MA, Mia AA, Mahmud F, Zeba N, Afrin K (2010) Genetic variability, character association and genetic divergence in mungbean (Vignaradiata L. Wilczek). Plant Omics 3:1–6
Sadegh B (2017) Evaluation of agronomic, physiological, agro-physiological, recovery and utilization efficiencies in three cultivars of soyabean. Biosci Biotechnol Res Commun 10(1):230–235
Schenker N, Taylor JMG (1996) Partially parametric techniques for multiple imputation. Comput Stat Data Ann 22(4):425–446
Sheela MN, Radhika VS, Susan John K, Abraham K (2008) Variation in crude protein, dry matter and starch in inbred and backcross lines of cassava. J Root Crops 34(2):115–119
Silva SA, Souza DR, Araujo GM, Almeida VO, Neves CG (2019) Genetic parameters in agronomic characters for selection of superior lineages of Ricinuscommunis L. Cienc Rural 49(1):e20180547. https://doi.org/10.1590/0103-8478cr20180547
Singh D, Ckonkar PK, Dwivedi BS (2005) A manual on soil, plant and water analysis. Centre for Soil-Plant-Water Analysis, Division of Soil Science and Agricultural Chemistry, Indian Agricultural Research Institute, New Delhi, Westville Publishing House, New Delhi, p 200
Soon YK (1992) Differential response of wheat genotypes to phosphorus in acid soils. J Plant Nutr 15:513–526
Suryanarayana L, Reddi Sekhar D, RatnaBabu D, Ramana AV, Srinivasa Rao V (2017) Cluster and principal component analysis in maize. Int J Curr Microbiol Appl Sci 6(7):354–359
Susan John K, Ravindran CS, James G (2005) Long term fertilizer experiments: 3 decades experience in cassava, technical bulletin series No. 45, ICAR- Central Tuber Crops Research Institute, Thiruvananthapuram, Kerala, India, p 83
Susan John K, Suja G, Sheela MN, Ravindran CS (2010) Potassium: the key nutrient for cassava production, tuber quality and soil productivity: an overview. J Root Crops 36:44–52
Van Buuren S, GroothuisOudshoorn CG (2011) MICE: multivariate imputation by chained equations in R. J Stat Softw 45(3):1–67. https://doi.org/10.18637/jss.v045.i03
Van Hintum TJL (1995) Hierarchical approaches to the analysis of genetic diversity in crop plants. In: Hodgkin T, Brown AHD, Van Hintum TJL, Morales EAV (eds) Core collection of plant genetic resources. Wiley Sayce Publication, New York, pp 23–34
Wang L, Chen F (2012) Genotypic variation of potassium uptake and use efficiency in cotton (Gossypiumhirsutum). J Plant Nutr Soil Sci 175:303–308
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
John, K.S., Sreekumar, J., Sheela, M.N. et al. Pre evaluation of cassava (Manihot esculenta Crantz) germplasm for genotypic variation in the identification of K efficient genotypes through different statistical tools. Physiol Mol Biol Plants 26, 1911–1923 (2020). https://doi.org/10.1007/s12298-020-00867-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12298-020-00867-2