Abstract
Severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2), the etiological agent of coronavirus disease-2019 (COVID-19), is a highly contagious pathogenic coronavirus to emerge and spread in human populations. Although substantial exertions have been laid to avert spread of COVID-19 by therapeutic and preventive countermeasures, but emergence of SARS-CoV-2 variants as a result of mutations make the infection more ominous. New viral confers a higher nasopharyngeal viral load, increased viral transmissibility, higher infectiousness, immune escape, increased resistance to monoclonal/polyclonal antibodies from convalescence sera/vaccine, and an enhanced virulence. Thus, it is pertinent to monitor evolving mutations and genetic diversity of SARS-CoV-2 as it is decisive for understanding the viral variants. In this review we provide an overview of colloquial nomenclature and the genetic characteristics of different SARS-CoV-2 variants in the context of mutational changes of the circulating strains, transmissibility potential, virulence and infectivity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coronavirus disease-2019 (COVID-19), a highly contagious disease, caused by severe acute respiratory syndrome coronavirus-2 (SARS-CoV-2), is a global mounting public health problem, that often leads to a series of pathologies which are detrimental to human health condition with unprecedented consequences. The first case of this viral infection was testified in Wuhan, the capital city of the Hubei province of China in December 2019, then SARS-CoV-2 rapidly disseminated all over the world and circumstances became worse within a short period of time, compelling the World Health Organization (WHO) to declare it as pandemic on 11th March 2020; as most of the countries experienced massive spikes in cases of COVID-19 [1]. However, numbers of infected cases, acute events and mortality rates allied to COVID-19 displayed vast variation from country to country or in different populace. As per records of WHO by 16th of May 2021, 162,177,376 infection cases have been reported with a global death of 3,364,178 [2].
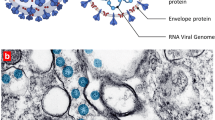
The reservoir of coronaviruses are bats, but it is believed to be transmitted to the human via an intermediate animal host [3]. SARS CoV-2 is an enveloped virion that measure approximately 120 nm in diameter, belonging to subfamily Orthocoronavirinae, in the family Coronaviridae, order Nidovirales, and realm Riboviria; containing positive-sense-single strand of RNA with a genome size of 29.99 kb encoding for multiple nonstructural and structural proteins [4, 5]. SARS-CoV-2 virus enters into type II pneumocytes of lung through Angiotensin Converting Enzyme 2 (ACE2) receptor [6], and then it starts replicating and migrating down to the airways and enters alveolar epithelial cells in the lungs. The swift replication of SARS-CoV-2 in the lungs might induces a robust immune response and pathogenesis manifests as mild viral pneumonia to severe acute respiratory distress syndrome [7, 8]. Initial cytokine storm is responsible for occurrence of macrophage activation syndrome and acute respiratory distress syndrome/respiratory failure. Simultaneously, SARS-CoV-2 spreads to the other organ(s) and infects cells which express ACE2 receptor, resulting in a in multi-organ damage [7]. Cytokine storm is followed by phase of immune dysregulation which is considered as a core culprit in the development of sepsis related complications [9].
Communal symptoms of SARS-CoV-2 infection are pyrexia, tussis, dyspnea, pharyngitis, myalgia, headache, olfactory and taste dysfunction (hyposmia/anosmia or ageusia). The widespread concern is the emergence of SARS-CoV-2 variants with concerning phenotypes as a result of mutation(s). SARS-CoV-2, despite being endowed with proofreading activity (a function attributed to nsp14) during viral replication [6], have a high mutation rate, and the absolute number of mutations increases with every round of infection either through genetic drift or through selection, and become fixed in different populations. Monitoring of these evolving mutations and genetic diversity of SARS-CoV-2 is decisive for understanding the viral variants and assuring the performance of new diagnostic tests and countermeasures (vaccines and therapies) against COVID-19. Thus, this study aims to perform a descriptive review of the basic virology of variants of SARS-CoV-2 i.e. we will highlight the current understanding; provide an overview of the nomenclature and genetic characteristics of viral variants in the context of mutational changes of the circulating strains, transmissibility potential, virulence and infectivity (Table 1).
Phylogeny and Genomics
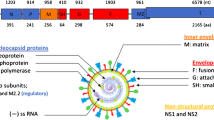
Phylogenetic analysis reveals SARS-CoV-2 genome shares high homology with other CoVs (79% with SARS-CoV; 50% with MERS-CoV; 88% with bat-derived CoVs) and accordingly is placed in the subgenus Sarbecovirus of the genus betacoronavirus [4, 5]. SARS‐CoV‐2 consists of positive-sense single-stranded RNA with distinctive 5′‐cap and 3′‐poly‐A tail structure and a low GC content compared to other CoVs (38%) [5]. The six functional open reading frames (ORFs) are organized in order from 5′ to 3′ as: replicase (ORF1a/ORF1b), spike (S), membrane (M), envelope (E), and nucleocapsid (N). ORF1a is the longest ORF, covers two-third portion of the 5′ genome and encodes a polyprotein (pp1ab), which is autoproteolytically processed into 16 non-structural proteins [4, 5]. The downstream regions encode for four main structural proteins (S, M, E, and N proteins) sequences along with seven accessory proteins encoding sequences, interspersed between the structural genes [4, 5].
The SARS‐CoV‐2 genome also contains leader and transcription‐regulatory sequence (TRS) [5]. The leader sequence (~ 70 bases) is present at 5′ end, out of which 7–10 bases are transcription‐regulatory sequences referred as TRS‐L. Similarly, adjacent to each ORF, TRS‐B motifs are present; TRS‐L and TRS‐B are responsible for the discontinuous synthesis of intermediate negative strands of sgRNA [6]. Of the four structural genes, S gene is significantly diverged from the corresponding sequence in SARS CoV, while other shares more than 90% homology [4,5,6]. The SARS-CoV-2 S is a trimeric glycoprotein, consists of 1273 amino acids. It contains two subunits viz S1 and S2 [6]. S1 is responsible for the viral entry by attaching to host cell’s ACE2 receptor through the S1 receptor-binding domain (RBD), while the S2 subunit allows virus-cell fusion of viral and cellular membranes. This process requires S protein priming by host proteases such as TMPRSS2 in cleavage sites S1/S2, a polybasic (furin) cleavage motif, at the S1/S2 boundary, and the S20 site [5, 6]. Additional distinctive genomic features of SARS-CoV-2 is, accessory gene orf8 and a presence of four amino acid residues (PRRA) at the junction of subunits S1 and S2. This insertion creates a polybasic cleavage site (RRAR), which allows effective cleavage by furin and other proteases [6].
Mutation: Variants and Strain
Mutation is a definite natural change that occurs in the genomic sequence of SARS-CoV-2 virus during process of viral replication [10, 11]. Natural selection processes will decide the fate of newly arising mutation. Mutations that are deleterious to the virus will be purged from the population, and few are essentially maintained in the population; they facilitate viral replication, transmissibility or immune escape i.e. if mutation favors the viral fitness, then it increases in frequency and evolute as a new variant [12]. However, frequency of mutation also depends on chance events. Hence, interplay between natural selection and chance events decide the evolution of new variants in host cells, populations and across the world [13]. If a genome differs from the reference sequence by one or more than one mutation then it is refereed as variant [14]. Specific variant which has different phenotypic characteristics like, virulence or transmissibility is called as a strain [15].
Nomenclature of SARS-CoV-2 Variants
Nomenclature helps to understand genomic epidemiology, surveillance and control of the infections [16]. Genetic diverseness of virus is primarily classified into specific large group called ‘clades’. On a phylogent tree, each specific ‘clade’ refers to a monophyletic group which is having a common ancestor. Thus, ‘clades’ classify genetic diverseness and pathogen phylogeny into mutually exclusive and equally divergent set of groups [17]. Wuhan/hCoV-19/ /WIV04/2019 sequence is considered as reference or zero sequence and it belongs to the A lineage of PANGO / 19B clade of Nextstrain/S clade of GISAID. No consistent nomenclature has been established for SARS-CoV-2, but colloquially four nomenclatures have been proposed for clades:
-
1.
According to Rambaut A, et al., 2020: It is also known as PANGO (Phylogenetic Assignment of Named Global Outbreak) Lineages. They recognized total 81 lineages of SARS-CoV-2 virus. These lineages were primary pertains to A, B, and B.1. Further six lineages, A.1 to A.6 were recognized from lineage A and two descendant sublineages, A.1.1 and A.3 were recognized from A.1. They also recognized 16 lineages directly originated from lineage B. So far, lineage B.1 is the major recognized lineage and it additionally subclassified into more than 70 sublineages [18].
-
2.
According to Nextstrain.org/ncov: Total 11 major clades were distinguished on the basis of analysis of clades.nextstrain.org. As per this system, clades are named by the year they are estimated to have emerged [19].
-
•
19A and 19B: Identified in early outbreak at Wuhan.
-
•
20A: Identified in outbreak of Europe in March 2020.
-
•
20C and 20D: They are genetically different sub-clades of 20A
-
•
20D to 20I: They emerged in early summer of 2020
-
•
-
3.
According to Guan Q, et al., 2020: They identified five main clades like, D392, I378, G614, S84, and V251 based on mutations profiles [20].
-
4.
According to Global Initiative on Sharing All Influenza Data (GISAID): Gisaid.org identified main nine clades such as GH, GRY, G, S, O, GV, L and V. [21].
Notable Variants of SARS-CoV-2
-
1.
Spike D614G variant: Substitution of aspartic acid to glycine at 614 amino acid position in spike protein of SARS-CoV-2 virus has been found in January and February 2020 and this mutation governs a vital conformational alteration in the spike protein between the S1 and S2 domains, that favors the binding to the ACE2 receptor and thus increases the probability of infection. Over a few months after the perception of ancestral strain, spike D614G variant became globally dominant, presumably because of enhanced ability to bind with human ACE2 receptor. D614G mutation also exhibit enhanced replication activity in both human (nasal and bronchial airway epithelial cell) and in animal models with a rapid transmissibility. Patients infected with the G variant shows a higher nasopharyngeal viral load than the wild-type strain, but there seems to be no difference in disease severity. Loss of smell is found to be the predominant symptom with G variant infectivity as this variant has boosted ability to bind with ACE2 receptor in olfactory epithelium. Sera of animal which were infected with this strain contains higher neutralizing antibody compared to original strain, suggestive of no additional effect of this variant on the efficacy of vaccine or of diagnostic and therapeutic measures [22,23,24,25].
-
2.
Lineage B.1.1.7: It is also called Variant of Concern 202,012/01 or 20B/501Y.V1 or UK COVID-19 variant and is characterized by 23 mutations (4 deletion, 6 synonymous mutation and 13 non-synonymous mutations) with 17 amino acid changes [26,27,28]. This variant was first announced in the South East of England on December 14th 2020 and it unfurled rapidly during late-December. Epidemiological studies and mathematical modeling revealed that transmission of this lineage is 56% faster than other lineage, as around 28% cases were infected with this strain by the end of December 2020. Rapid and widespread transmission of this variant is suggestive of natural selection of virus at a population level, though it does not have any significant effect on efficacy of vaccine or of diagnostic and therapeutic measures. Variants from this lineage are associated with multiple amino acid changes in the spike protein, including a deletion at 69/70, mutation in ORF8, P681H, N501Y and E484K mutation.
-
A.
N501Y: The N501Y (substitution of Asparagine to Tyrosine at 501 amino acid position) mutation is of major concern of B.1.1.7 lineage, as it is present in Receptor Binding Motif of Receptor Binding Domain (RBD) in Spike glycoprotein. Analysis through modelling methods reveled that N501Y mutation would allow a potential aromatic ring-ring interaction and an additional hydrogen bond between RBD and ACE2 and, hence, confers an increased binding affinity of S RBD for the ACE2 receptor, raising the viral transmission rate. This mutation confers an increased infectivity and virulence as it alters the antibody identification and ACE2 receptor binding specificity of spike glycoprotein [29].
-
B.
P681H: This mutation occurs spontaneously and is located adjacent to the amino acids 682–685, the furin cleavage site (FCS) identified at the S1/S2 in the spike protein. The function of the P681H mutation is not yet clear, but SARS-CoV-2 FCS promotes the promotes the entry of the virus into respiratory epithelial cells and enhances transmembrane serine protease (TMPRSS) induced cleavage ability. It is one the three mutations (along with H69-V70del, N501Y) of B.1.1.7 with the utmost potential to affect the biological behavior of the SARS-CoV-2 [30].
-
C.
69/70 deletion: It refers to 6 bp deletion at 69 and 70 amino acid positions in spike protein which causes a conformational modification in the spike protein. In immunocompromised patients, this deletion has been associated with immune escape mechanism and also deemed plausible a determinant for drop out in detection of S gene in some diagnostic method [31].
-
D.
Mutation in ORF8: It refers to mutation in stop codon (Q27stop) in ORF8 [32] and is responsible to truncates the ORF8 protein or makes it inactive, permitting the accumulation of further mutations in other regions.
-
E.
E484K mutation: Lineage B.1.1.7 with E484K mutation was identified in England in February 2021, which was also detected in Brazil and South Africa Variants. This mutation plays a pivotal role in immune evasion and in the binding affinity of the virus to the receptor by bringing a conformational alteration in the flexible loop region of S RBD. E484K mutation is also deemed to be responsible for the reduced efficacy of both vaccine and convalescent sera [33].
-
A.
-
3.
Lineage B.1.351 or 501.2 variant: It is also known as 20C/501Y.V2 or South African Covid Variant and was first detected in South Africa during mid-December 2020. This variant has multiple mutations (12 non-synonymous mutations and one deletion) equated to index Wuhan strain and about three-fourth of these mutations are in the spike glycoprotein viz: A701V, D80A, D215G, E484K, L18F, AL 242–244 del, R246I, K417N, D614G and N501Y, while the other are sited in ORF1a [K1655N], envelope [P71L], and N [T205I] viral proteins. This variant is found to be more prevalent in young individuals (without any comorbidity), confers increased binding to ACE2 receptor and is deemed responsible for the second wave. In view of multiple spike mutation(s), it spreads rapidly compared to other variants and vaccine might have reduced efficacy against this variant, presumably because of K417N and E484K mutations in the RBD domain [34].
-
4.
A701B variant: This variant was identified in December 2020 at Malaysia and is characterized by substitution of alanine by aspartic acid at the 701 amino acid position in spike protein. Rate of transmission and infectivity of this strain is uncertain [35].
-
5.
Spike N453Y variant: It is also known as cluster 5 variant or ΔFVI-spike and was discovered in mid-August to early part of September 2020 at North Jutland of Denmark and spreads from mink to humans through mink farms. Infection with this variant may cause decrease sensitivity for neutralization of virus, which may result in decrease in duration of immune protection by vaccination and natural course of infection. Adaptation of this variant in mink is an important concern in future because evolution of virus in mink reservoirs results in recurrent chance of infection to human from mink. Hence, some countries implemented extensive slaughtering of mink to reduce the further spread of infection [36].
-
6.
Lineage B.1.1.207: This variant was identified in Nigeria in late August 2020 with mutation of P681H, which is also found in Lineage B.1.1.7. Transmission and virulence of this variant is unclear [37].
-
7.
Lineage B.1.258∆: It has been identified within B.1.258 clade in Czech Republic and Slovakia during September-December 2020. It contains following imperative mutations:
-
A.
N439K mutation: It occurs in the receptor binding domain of spike protein and has been associated with high viral load as it increases the binding affinity of virus to ACE2 receptor. It has been reported that it escapes immune response developed by previous infection and from neutralizing monoclonal antibodies [38].
-
B.
∆H69/∆V70 deletion: It occurs in amino terminal domain of spike protein and has been associated with high infectivity substantiated by a two-fold increase in S protein-mediated infectivity in vitro along with the flair to escape from the immune response. This deletion has not been identified by RT-PCR of TaqPath and often get misdiagnosed as B.1.1.7 [39, 40]; H69/V70 deletion is considered as a permissive mutation i.e. it amends the immunodominant epitopes sited at amino terminal domain (variable loops), providing resistance to neutralization by both convalescent sera and vaccine.
-
A.
-
8.
P.1 Variant: It is also known as a 20 J/501Y.V3 and was identified in Brazil as dominating circulating virus. This variant belongs to B.1.1.28 lineage and contains 17 non-synonymous mutations, 11 in S protein viz L18F, T20N, P26S, D138Y, R190S, K417T, E484K, N501Y, D614G, H655Y, T1027I, and V1176F; 3 in ORF1ab viz S1188L, K1795Q, and E5665D; one in ORF8 namely E92K, and one P80K in N protein; 1 deletion namely SGF 3675-3677del in ORF1ab; and 4 synonymous mutations. This SARS-CoV-2 variant contains the highest number of mutations in spike protein and all these mutations collectively has important implications for transmissibility, reinfection rates and evasion of antibody-mediated immunity [41].
-
9.
Midwest variant or S Q677H: This variant contains Q677H mutation in spike protein and was reported to be highly prevalent in Ohio and Midwest states from December 2020 to January 2021. Functional effect of this variant regarding antigenicity and transmissibility of SARS-CoV-2 is uncertain [42].
-
10.
CAL.20C: It arises from lineage B.1.429 and was first identified in Europe and Los Angeles of United states in the year 2020, then it rapidly escalated in California during January 2021. This variant contains L452R mutation in receptor binding domain of spike protein, which has been found to resistant against therapeutic monoclonal antibodies [43]. Besides L452R, CAL.20C variant contains four mutations viz, I4205V in ORF1a; D1183Y in ORF1b; S13I, and W152C in spike protein. Functional effect of this variant on infectivity, antigenicity and disease severity of SARS-CoV-2 is ambiguous.
-
11.
Lineage B.1.525: This lineage was first identified in United Kingdom during December 2020 and contains four mutations viz Q677H, F888L, E484K, and Q52R in spike protein [44]
-
12.
Lineage B.1.617: This lineage was first identified at Maharashtra state of India in October 2020 during second wave surge. It possesses common novel mutations viz E484Q, L452R, P681R, D111D, D614G, and G142D in receptor binding domain of spike protein; amongst these E484Q, L452R, and P681R mutations are major circulatory variants and area of concern Each of the three mutations are in furin cleavage site and might accelerate S1-S2 cleavage, binding affinity of virus to ACE2 receptor, that leads to superior transmissibility [45]. Effect of this lineage is found to be neutralized with currently available vaccine and convalescent plasma of previously infected with SARS-CoV-2 infection. Hence, this lineage hasn’t any effect on immune escape and vaccine efficacy [46].
-
13.
Lineage B.1.168: This lineage was first acknowledged in October 2020 at West Bengal state of India and is found to be significantly increased lately. This lineage contains E484K, D618G and deletion of two amino acid (Y145 and H146) in region of spike protein. E484K variant has been considered as main immune escape variant as it escapes from convalescent plasma and multiple monoclonal antibodies. [47]
Consequences of Emerging Variants of SARS-CoV-2
-
1.
Escaping from specific diagnostic investigation: Mutation(s) in the spike glycoprotein affects the detection of spike protein by RT-PCR. However, most of the commercially available PCR protocols are using multiple targets and presumably no significant impact on diagnosis [48].
-
2.
Faster transmissibility in the population: D614G [49], B.1.1.7 [31], and B.1.617 [45] variant of SARS-CoV-2 has been associated with faster transmissibility compared to 614D variant (wild type). These variants propagate more quickly in human respiratory epithelial cells.
-
3.
Severity of infection: Mainly Lineage B.1.1.7 variant has been associated with increased severity of infection [31].
-
4.
Impact on vaccine effectiveness: Spike glycoprotein of SARS-CoV-2 attaches to ACE2 receptor in respiratory tract and it is a central target for neutralizing antibody, thus a potential target for vaccine efficacy. All these notable variants still have not spread at population level and are not unequivocally proficient to escape from host antibody response; hence should not have any substantial significant influence on the vaccine efficacy [50]. However, additional mutation(s) in the spike glycoprotein could lead to immune escape from neutralizing antibody and replacing other circulating variants by increasing transmissibility at a large populace level, then it could be impending cause of diminish effectiveness of the vaccine. mRNA vaccines stimulate virus specific cytotoxic and helper T cells and generate efficient neutralizing antibody response which is quiet enough to fight against newer variants. However, inactivated vaccines might be less effective against newer variants as they produce weak neutralizing antibody response. Genomic sequencing of virus should be done from fully vaccinated individuals admitted to hospital with SARS-CoV-2 infection to identify the new variants [50]. Genomic surveillance of these entire notable variant should be continued at large scale to know their impact on host immune response and on vaccine efficacy and to prevent global spread of new variants.
Major Clades/Variants in India
There was progressive decrease in cases of SARS-CoV-2 infection from 15th September 2020 to 9th of February 2021 in India. But there was abrupt spike in statistics of new cases from 10th February 2021, signifying evidence of second wave of SARS-CoV-2 infection. A2a, A3, AI/A3i and B4 are major clades found in India. Primarily, A/A3i clade emerged in March 2020 and was subsequently replaced by A2a clade. Clade AI/A3i was significantly reported in Delhi and clade B4 in West Bengal and Odisha [51]. Lineage B.1.617 and Lineage B.1.168 are responsible for recent quick surge of SARS-CoV-2 cases in India.
Role of Genome Sequencing in Surveillance
Genomics evaluate the particular virology in real time. First library of consistently sized fragments is generated by cleaving multiple genomes at randomly. Then unordered sequenced segments are generated by automated computation. Finally segments of overlapping sequence are constructed to know genome consensus [52]. Figure 1 depicts the steps involved in the genomic sequencing of SRAS-CoV-2 [53]. Genome sequencing helps to, identify the origin of infection; comprehend the evolution of pathogen, confirm the circulating variants and to develop the therapeutics and vaccines. Next generation sequencing (NGS) has sensitivity of 98.4% and specificity of 97.2% which is distantly improved equated to conventional RT-PCR [53]. Evaluation of samples in NGS will be imperative to guide the policy makers for the fight against SARS-CoV-2 in future. Figure 2 depicts proposed flow of work to identify and genomic counseling of new SARS-CoV-2 variant in future.
Conclusion
Emergent SARS-CoV-2 variants (evolving largely due to mutations in S protein, NTD and RBD), have in common an increased viral transmissibility, higher infectiousness, immune escape, increased resistance to monoclonal/polyclonal antibodies from convalescence sera/vaccine, and an enhanced virulence; altogether enabling a higher rate of severe disease. Thus, national authorities should continue the genomic surveillance, which might aid disease control, management and prevention efforts.
References
Mitra P, Misra S, Sharma P. One year of COVID-19: the “new normal”. Indian J Clin Biochem. 2021;36(1):1–2.
WHO Coronavirus Disease (COVID-19) Dashboard n.d. https://covid19.who.int. Accessed 16th May 2021.
Wang LF, Eaton BT. Bats, civets and the emergence of SARS. Curr Top Microbiol Immunol. 2007;315:325–44.
Lu R, Zhao X, Li J, Niu P, Yang B, Wu H, Wang W, et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet. 2020;395(10224):565–74.
Chan JF, Kok KH, Zhu Z, Chu H, To KK, Yuan S. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg Microbes Infect. 2020;9(1):221–36.
Ou X, Liu Y, Lei X, Li P, Mi D, Ren L, et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat Commun. 2020;11(1):1620.
Ni W, Yang X, Yang D, Bao J, Li R, Xiao Y, et al. Role of angiotensin-converting enzyme 2 (ACE2) in COVID-19. Crit Care. 2020;24:422.
Baj J, Karakuła-Juchnowicz H, Teresinski G, Buszewicz G, Ciesielka M, Sitarz E, et al. COVID-19: specific and non-specific clinical manifestations and symptoms: the current state of knowledge. J Clin Med. 2020;9:1753.
Ye Q, Wang B, Mao J. The pathogenesis and treatment of the `cytokine storm’ in COVID-19. J Infect. 2020;80:607–13.
Wang H, Li X, Li T, Zhang S, Wang L, Wu X, et al. The genetic sequence, origin, and diagnosis of SARS-CoV-2. Eur J Clin Microbiol Infect Dis. 2020;39:1629–35.
Sanjuan R, Domingo-Calap P. Mechanisms of viral mutation. Cell Mol Life Sci. 2016;73:4433–48.
Rodpothong P, Auewarakul P. Viral evolution and transmission effectiveness. World J Virol. 2012;1:131–4.
Livnat A. Interaction-based evolution: how natural selection and nonrandom mutation work together. Biol Direct. 2013;8:24.
Karki R, Pandya D, Elston RC, Ferlini C. Defining, “mutation” and “polymorphism” in the era of personal genomics. BMC Med Genomics. 2015;8:37.
Andino R, Domingo E. Viral quasispecies. Virology. 2015;479–480:46–51.
Sironi M, Hasnain SE, Rosenthal B, Phan T, Luciani F, Shaw MA, et al. Editors of infection, genetics and evolution. SARS-CoV-2 and COVID-19: a genetic, epidemiological, and evolutionary perspective. Infect Genet Evol. 2020;84:104384.
Zhao Z, Sokhansanj BA, Malhotra C, Zheng K, Rosen GL. Genetic grouping of SARS-CoV-2 coronavirus sequences using informative subtype markers for pandemic spread visualization. PLoS Comput Biol. 2020;16:e1008269.
Rambaut A, Holmes EC, O'Toole A, Hill V, McCrone JT, Ruis C, et al. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat Microbiol. 2020;5:1403–7.
https://clades.nextstrain.org/. Accessed 15 Feb 2021.
Guan Q, Sadykov M, Mfarrej S, Hala S, Naeem R, Nugmanova R, et al. A genetic barcode of SARS-CoV-2 for monitoring global distribution of different clades during the COVID-19 pandemic. Int J Infect Dis. 2020;100:216–23. https://www.gisaid.org/phylodynamics/global/nextstrain/ Accessed 15 Feb 2021.
https://www.gisaid.org/phylodynamics/global/nextstrain/. Accessed 15 Feb 2021.
Yurkovetskiy L, Wang X, Pascal KE, Tomkins-Tinch C, Nyalile TP, Wang Y, et al. Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant. Cell. 2020;183:739-751.e8.
Zhou B, Thao TTN, Hoffmann D, Taddeo A, Ebert N, Labroussaa F, et al. SARS-CoV-2 spike D614G variant confers enhanced replication and transmissibility. bioRxiv [Preprint]. 2020:2020.10.27.357558.
Groves DC, Rowland-Jones SL, Angyal A. The D614G mutations in the SARS-CoV-2 spike protein: implications for viral infectivity, disease severity and vaccine design. Biochem Biophys Res Commun. 2021;538:104–7.
Zhang L, Jackson CB, Mou H, Ojha A, Rangarajan ES, Izard T, et al. The D614G mutation in the SARS-CoV-2 spike protein reduces S1 shedding and increases infectivity. bioRxiv [Preprint]. 2020 Jun 12:2020.06.12.148726.
Galloway SE, Paul P, MacCannell DR, Johansson MA, Brooks JT, MacNeil A, et al. Emergence of SARS-CoV-2 B.1.1.7 lineage - United States, december 29, 2020-january 12, 2021. MMWR Morb Mortal Wkly Rep. 2021;70:95–9.
Raskin S. Genetics of COVID-19. J Pediatr (Rio J). 2020;S0021–7557(20):30211–4.
Su S, Wang Q, Jiang S. Facing the challenge of viral mutations in the age of pandemic: developing highly potent, broad-spectrum, and safe COVID-19 vaccines and therapeutics. Clin Transl Med. 2021;11:e284.
Leung K, Shum MH, Leung GM, Lam TT, Wu JT. Early transmissibility assessment of the N501Y mutant strains of SARS-CoV-2 in the United Kingdom, october to november 2020. Euro Surveill. 2021;26:2002106.
Maison DP, Ching LL, Shikuma CM, Nerurkar VR. Genetic characteristics and phylogeny of 969-bp S gene sequence of SARS-CoV-2 from hawaii reveals the worldwide emerging P681H mutation. bioRxiv [Preprint]. 2021:2021.01.06.425497.
Rahimi F, Talebi Bezmin Abadi A. Implications of the emergence of a new variant of SARS-CoV-2, VUI-202012/01. Arch Med Res. 2021. https://doi.org/10.1016/j.arcmed.2021.01.001
Pereira F. Evolutionary dynamics of the SARS-CoV-2 ORF8 accessory gene. Infect Genet Evol. 2020;85:104525.
Wang P, Nair MS, Liu L, Iketani S, Luo Y, Guo Y, et al. Increased resistance of SARS-CoV-2 variants B.1.351 and B.1.1.7 to antibody neutralization. bioRxiv [Preprint]. 2021:2021.
Tang JW, Toovey OTR, Harvey KN, Hui DDS. Introduction of the South African SARS-CoV-2 variant 501Y.V2 into the UK. J Infect. 2021;82:e8–10.
Chong YM, Sam IC, Chong J, Kahar Bador M, Ponnampalavanar S, Syed Omar S, et al. SARS-CoV-2 lineage B.6 was the major contributor to early pandemic transmission in Malaysia. PLoS Negl Trop Dis. 2020;14:e0008744.
Oude Munnink BB, Sikkema RS, Nieuwenhuijse DF, Molenaar RJ, Munger E, Molenkamp R, et al. Transmission of SARS-CoV-2 on mink farms between humans and mink and back to humans. Science. 2021;371:172–7.
CDC. Centers for Disease Control and Prevention. 18 January 2021. https://www.cdc.gov/mmwr/Novel_Coronaviurs_Reports.html. Retrieved 2 Feb 2021.
Thomson EC, Rosen LE, Shepherd JG, Spreafico R, Filipe ADS, Wojcechowskyj JA, et al. Circulating SARS-CoV-2 spike N439K variants maintain fitness while evading antibody-mediated immunity. Cell (in press). 2021. https://doi.org/10.1016/j.cell.2021.01.037.
Vogels CBF, Breban M, Alpert T, Petrone ME, Watkins AE, Hodcroft EB, et al. Tracking SARS-CoV-2 VOC 202012/01 (lineage B.1.1.7) dissemination in Portugal: insights from nationwide RT-PCR Spike gene drop out data. 2021. https://www.virological.org/ Accessed 14 Feb 2021.
Bal A, Destras G, Gaymard A, Stefic K, Marlet J, Eymieux S, et al. Two-step strategy for the identification of SARS-CoV-2 variant of concern 202012/01 and other variants with spike deletion H69–V70, France, august to december 2020. Euro Surveill. 2021;26:2100008.
Maggi F, Novazzi F, Genoni A, Baj A, Spezia PG, Focosi D, et al. Imported SARS-COV-2 variant p.1 detected in traveler returning from Brazil to Italy. Emerg Infect Dis. 2021;27:1249–51.
Kim JS, Jang JH, Kim JM, Chung YS, Yoo CK, Han MG. Genome-wide identification and characterization of point mutations in the SARS-CoV-2 genome. Osong Public Health Res Perspect. 2020;11:101–11.
Li Q, Wu J, Nie J, Zhang L, Hao H, Liu S, et al. The impact of mutations in SARS-CoV-2 spike on viral infectivity and antigenicity. Cell. 2020;182:1284-1294.e9.
Gomez CE, Perdiguero B, Esteban M. Emerging SARS-CoV-2 variants and impact in global vaccination programs against SARS-CoV-2/COVID-19. Vaccines (Basel). 2021;9:243.
Cherian S, Potdar V, JadhavS, Yadav P, Gupta N, Das M, et al. Convergent evolution of SARS-CoV-2 spike mutateons, L452R, E484Q and P681R, in the second wave of COVID-19 in Maharashtra, India. bioRxiv [Preprint]. 2021:2021.
Yadav PD, Sapkal GN, Abraham P, Ella R, Patil DY, Deshpande G. et al. Neutralization of variant under investigation B/1/617 with sera of BBV152 vaccines. bioRxiv [Preprint]. 2021:2021.
https://cov-lineages.org/lineages/lineage_B.1.618.html/. Accessed on 8 May 2021.
Korber B, Fischer WM, Gnanakaran S, Yoon H, Theiler J, Abfalterer W, et al. Tracking changes in SARS-CoV-2 spike: evidence that D614G increases infectivity of the COVID-19 virus. Cell. 2020;182:812-827.e19.
Conti P, Caraffa A, Gallenga CE, Kritas SK, Frydas I, Younes A, et al. The British variant of the new coronavirus-19 (Sars-Cov-2) should not create a vaccine problem. J Biol Regul Homeost Agents. 2020;35:1–4.
Koyama T, Weeraratne D, Snowdon JL, Parida L. Emergence of drift variants that may affect COVID-19 vaccine development and antibody treatment. Pathogens. 2020;9:324.
Banu S, Jolly B, Mukherjee P, et al. A distinct phylogenetic cluster of Indian severe acute respiratory syndrome coronavirus 2 isolates. Open Forum Infect Dis. 2020;7:ofaa434.
Shi W, Li J, Zhou H, Gao GF. Pathogen genomic surveillance elucidates the origins, transmission and evolution of emerging viral agents in China. Sci China Life Sci. 2017;60:1317–30.
Bhoyar R, Jain A, Sehgal P, Divakar MV, Sharma E, Imran M et al. High throughput detection and genetic epidemiology of SARS-CoV-2 using COVIDSeq next generation sequencing. PlOS ONE. 2021. https://doi.org/10.1371/journal.pone.0247115
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Dholariya, S., Parchwani, D.N., Singh, R. et al. Notable and Emerging Variants of SARS-CoV-2 Virus: A Quick Glance. Ind J Clin Biochem 36, 451–458 (2021). https://doi.org/10.1007/s12291-021-00991-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12291-021-00991-0