Abstract
Background
Breast cancer is a prevalent cancer in female. This study aims to investigate the therapeutic potential and mechanism of curcumin in breast cancer.
Methods
After cultivation, human breast cancer cells (MCF-7 cells) were treated with 0.1% (v/v) 15 µmol/ml curcumin-dimethylsulfoxide solution and 0.1% (v/v) dimethylsulfoxide, respectively, at 37 °C and 5% CO2 for 48 h. Total RNA was extracted, cDNA library was constructed, and cDNAs were amplified and sequenced. After data preprocessing, the Cufflinks software was utilized to identify differentially expressed genes (DEGs, |log2 fold change| > 0.5 and p value < 0.05). Then, functional and pathway enrichment analyses were performed through DAVID (p value < 0.05) and WebGestalt [false discovery rate (FDR) < 0.05], respectively. Furthermore, drug and disease association analyses (FDR < 0.05) were conducted through WebGestalt and DAVID, respectively. STRING was employed to construct protein–protein interaction (PPI) network (combined score > 0.4).
Results
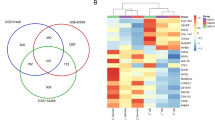
After DEGs screening, 347 DEGs were identified. Up-regulated DEGs were enriched in 14 functions and 3 pathways, and associated with 12 drugs. Down-regulated DEGs were enriched in 14 functions and 9 pathways, and associated with 14 drugs. Moreover, 5 DEGs were associated with breast cancer, including PGAP3, MAP3K1, SERPINE1, PON2, and GSTO2. PPI network was constructed, and the top DEGs were FOS, VIM, FGF2, MAPK1, SPARC, TOMM7, PSMB10, TCEB2, SOCS1, COL4A1, UQCR11, SERPINE1, and ISG15.
Conclusion
Curcumin might have therapeutic potential in breast cancer through regulating breast cancer-related genes, including SERPINE1, PGAP3, MAP3K1, MAPK1, GSTO2, VIM, SPARC, and FGF2. However, validations are required.


Similar content being viewed by others
References
Siegel R, Ma J, Zou Z, Jemal A. Cancer statistics, 2014. CA Cancer J Clin. 2014;64:9–29.
Zilli M, Grassadonia A, Tinari N, Di Giacobbe A, Gildetti S, Giampietro J, et al. Molecular mechanisms of endocrine resistance and their implication in the therapy of breast cancer. Biochim Biophys Acta. 2009;1795:62–81.
Baselga J, Cortés J, Kim S-B, Im S-A, Hegg R, Im Y-H, et al. Pertuzumab plus trastuzumab plus docetaxel for metastatic breast cancer. N Engl J Med. 2012;366:109–19.
Basnet P, Skalko-Basnet N. Curcumin: an anti-inflammatory molecule from a curry spice on the path to cancer treatment. Molecules. 2011;16:4567–98.
Ozawa H, Imaizumi A, Sumi Y, Hashimoto T, Kanai M, Makino Y, et al. Curcumin β-d-Glucuronide plays an important role to keep high levels of free-form curcumin in the blood. Biol Pharm Bull. 2017;40:1515.
Ammon HP, Wahl MA. Pharmacology of Curcuma longa. Planta Med. 1991;57:1–7.
Choudhuri T, Pal S, Agwarwal ML, Das T, Sa G. Curcumin induces apoptosis in human breast cancer cells through p53-dependent Bax induction. FEBS Lett. 2002;512:334–40.
Nagaraju GP, Aliya S, Zafar SF, Basha R, Diaz R, El-Rayes BF. The impact of curcumin on breast cancer. Integr Biol. 2012;4:996–1007.
Trapnell C, Roberts A, Goff L, Pertea G, Kim D, Kelley DR, et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Protoc. 2012;7:562–78.
Da Wei Huang BTS, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2008;4:44–57.
Wang J, Duncan D, Shi Z, Zhang B. WEB-based gene set analysis toolkit (WebGestalt): update 2013. Nucleic Acids Res. 2013;41:439.
Franceschini A, Szklarczyk D, Frankild S, Kuhn M, Simonovic M, Roth A, et al. STRING v9. 1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 2013;41:D808–15.
Kohl M, Wiese S, Warscheid B. Cytoscape: software for visualization and analysis of biological networks. Data mining in proteomics. Berlin: Springer; 2011. p. 291–303.
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Investig. 2011;121:2750–67.
Asiedu MK, Ingle JN, Behrens MD, Radisky DC, Knutson KL. TGFβ/TNFα-mediated epithelial–mesenchymal transition generates breast cancer stem cells with a claudin-low phenotype. Can Res. 2011;71:4707–19.
Stephens PJ, Tarpey PS, Davies H, Van Loo P, Greenman C, Wedge DC, et al. The landscape of cancer genes and mutational processes in breast cancer. Nature. 2012;486:400–4.
Zhang B, Zhao Y, Zhu J. Global gene regulatory and protein interaction networks in breast cancer metastasis. Cancer Res. 2013;73:A81.
Izrailit J, Berman HK, Datti A, Wrana JL, Reedijk M. High throughput kinase inhibitor screens reveal TRB3 and MAPK-ERK/TGFβ pathways as fundamental Notch regulators in breast cancer. Proc Natl Acad Sci. 2013;110:1714–9.
Masoudi M, Saadat I, Omidvari S, Saadat M. Association between N142D genetic polymorphism of GSTO2 and susceptibility to colorectal cancer. Mol Biol Rep. 2011;38:4309.
Wang Z, Qu K, Huang Z, Xu X, Zhang J, Zhang L, et al. Glutathione S-transferase O2 gene rs157077 polymorphism predicts response to transarterial chemoembolization in hepatocellular carcinoma. Tumor Biol. 2015;36:6463–9.
Pongstaporn W, Rochanawutanon M, Wilailak S, Linasamita V, Weerakiat S, Petmitr S. Genetic alterations in chromosome 10q24. 3 and glutathione S-transferase omega 2 gene polymorphism in ovarian cancer. J Exp Clin Cancer Res CR. 2006;25:107.
Masoudi M, Saadat I, Omidvari S, Saadat M. Additive effects of genetic variations of xenobiotic detoxification enzymes and DNA repair gene XRCC1 on the susceptibility to breast cancer. Breast Cancer Res Treat. 2010;120:263–5.
Andonova IE, Justenhoven C, Winter S, Hamann U, Baisch C, Rabstein S, et al. No evidence for glutathione S-transferases GSTA2, GSTM2, GSTO1, GSTO2, and GSTZ1 in breast cancer risk. Breast Cancer Res Treat. 2010;121:497–502.
Huang C-C, Tu S-H, Lien H-H, Jeng J-Y, Huang C-S, Huang C-J, et al. Concurrent gene signatures for han chinese breast cancers. PLoS One. 2013;8:e76421.
Armstrong AJ, Marengo MS, Oltean S, Kemeny G, Bitting RL, Turnbull JD, et al. Circulating tumor cells from patients with advanced prostate and breast cancer display both epithelial and mesenchymal markers. Mol Cancer Res. 2011;9:997–1007.
Cheng C-W, Wang H-W, Chang C-W, Chu H-W, Chen C-Y, Yu J-C, et al. MicroRNA-30a inhibits cell migration and invasion by downregulating vimentin expression and is a potential prognostic marker in breast cancer. Breast Cancer Res Treat. 2012;134:1081–93.
Basu G, Van Vickle G, Ghazalpour A, Ashfaq R, Gatalica Z, Blevins R, et al. Frequency distribution of SPARC in triple-negative breast cancer patients. J Clin Oncol. 2011;29:s27.
Guillardoy T, Gorostiaga MA, Lanari C, Giulianelli S. FGF-2 stimulates breast cancer growth activating ER and PR. Mol Cancer Res. 2013;11:A006.
Acknowledgements
None.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interests to state.
About this article
Cite this article
Wang, R., Li, J., Zhao, Y. et al. Investigating the therapeutic potential and mechanism of curcumin in breast cancer based on RNA sequencing and bioinformatics analysis. Breast Cancer 25, 206–212 (2018). https://doi.org/10.1007/s12282-017-0816-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12282-017-0816-6