Abstract
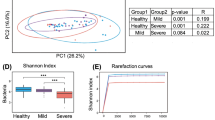
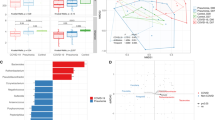
Several follow-up studies have found that COVID-19 (coronavirus disease 2019) patients had persistent symptoms after discharge. Gut microbiota play an important role in human health and immune responses. Therefore, this study investigated the gut microbiota of recovered COVID-19 patients and the correlations between gut microbiota and persistent symptoms after discharge. Stool samples were collected from 15 recovered healthcare workers (HCWs) with COVID-19 at three months after discharge, in addition, stool samples were collected from 14 healthy controls (HCs) to perform 16S rRNA gene sequencing between May and July 2020. Compared with HCs, recovered HCWs had reduced bacterial diversity at three months after discharge, with a significantly higher relative abundance of opportunistic pathogens, and a significantly lower relative abundance of beneficial bacteria. In addition, Escherichia unclassified was positively correlated with persistent symptoms at three months after discharge, including fatigue (r = 0.567, p = 0.028), chest tightness after activity (r = 0.687, p = 0.005), and myalgia (r = 0.523, p = 0.045). Intestinibacter bartlettii was positively correlated with anorexia (r = 0.629, p = 0.012) and fatigue (r = 0.545, p = 0.036). However, Faecalibacterium prausnitzii was negatively correlated with chest tightness after activity (r = -0.591, p = 0.02), and Intestinimonas butyriciproducens was negatively correlated with cough (r = -0.635, p = 0.011). In conclusion, the gut microbiota of recovered HCWs with COVID-19 at three months after discharge was different from that of HCs, and altered gut microbiota was correlated with persistent symptoms after discharge, highlighting that gut microbiota may play an important role in the recovery of patients with COVID-19.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Availability of Data and Materials
The sequences from our study were deposited in the NCBI Sequence Read Archive (PRJNA736160).
References
Arpaia, N., Campbell, C., Fan, X., Dikiy, S., van der Veeken, J., deRoos, P., Liu, H., Cross, J.R., Pfeffer, K., Coffer, P.J., et al. 2013. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455.
Belkaid, Y. and Hand, T.W. 2014. Role of the microbiota in immunity and inflammation. Cell 157, 121–141.
Bojović, K., Ignjatović, Đ., Soković Bajić, S., Vojnović Milutinović, D., Tomić, M., Golić, N., and Tolinaćki, M. 2020. Gut microbiota dysbiosis associated with altered production of short chain fatty acids in children with neurodevelopmental disorders. Front. Cell. Infect. Microbiol. 10, 223.
Bui, T.P., Shetty, S.A., Lagkouvardos, I., Ritari, J., Chamlagain, B., Douillard, F.P., Paulin, L., Piironen, V., Clavel, T., Plugge, C.M., et al. 2016. Comparative genomics and physiology of the butyrateproducing bacterium Intestinimonas butyriciproducens. Environ. Microbiol. Rep. 8, 1024–1037.
Chan, J.F.W., Yuan, S., Kok, K.H., To, K.K.W., Chu, H., Yang, J., Xing, F., Liu, J., Yip, C.C.Y., Poon, R.W.S., et al. 2020. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet 395, 514–523.
Chang, S.C., Shen, M.H., Liu, C.Y., Pu, C.M., Hu, J.M., and Huang, C.J. 2020. A gut butyrate-producing bacterium Butyricicoccus pullicaecorum regulates short-chain fatty acid transporter and receptor to reduce the progression of 1,2-dimethylhydrazineassociated colorectal cancer. Oncol. Lett. 20, 327.
Furet, J.P., Kong, L.C., Tap, J., Poitou, C., Basdevant, A., Bouillot, J.L., Mariat, D., Corthier, G., Doré, J., Henegar, C., et al. 2010. Differential adaptation of human gut microbiota to bariatric surgery-induced weight loss: links with metabolic and low-grade inflammation markers. Diabetes 59, 3049–3057.
Furusawa, Y., Obata, Y., Fukuda, S., Endo, T.A., Nakato, G., Takahashi, D., Nakanishi, Y., Uetake, C., Kato, K., Kato, T., et al. 2013. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450.
Galardini, M., Clermont, O., Baron, A., Busby, B., Dion, S., Schubert, S., Beltrao, P., and Denamur, E. 2020. Major role of iron uptake systems in the intrinsic extra-intestinal virulence of the genus Escherichia revealed by a genome-wide association study. PLoS Genet. 16, e1009065.
Gu, S., Chen, Y., Wu, Z., Chen, Y., Gao, H., Lv, L., Guo, F., Zhang, X., Luo, R., Huang, C., et al. 2020. Alterations of the gut microbiota in patients with coronavirus disease 2019 or H1N1 influenza. Clin. Infect. Dis. 71, 2669–2678.
Huang, C., Wang, Y., Li, X., Ren, L., Zhao, J., Hu, Y., Zhang, L., Fan, G., Xu, J., Gu, X., et al. 2020. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Lancet 395, 497–506.
Ichinohe, T., Pang, I.K., Kumamoto, Y., Peaper, D.R., Ho, J.H., Murray, T.S., and Iwasaki, A. 2011. Microbiota regulates immune defense against respiratory tract influenza A virus infection. Proc. Natl. Acad. Sci. USA 108, 5354–5359.
Jin, M., Kalainy, S., Baskota, N., Chiang, D., Deehan, E.C., McDougall, C., Tandon, P., Martínez, I., Cervera, C., Walter, J., et al. 2019. Faecal microbiota from patients with cirrhosis has a low capacity to ferment non-digestible carbohydrates into short-chain fatty acids. Liver Int. 39, 1437–1447.
Kuang, Y.S., Lu, J.H., Li, S.H., Li, J.H., Yuan, M.Y., He, J.R., Chen, N.N., Xiao, W.Q., Shen, S.Y., Qiu, L., et al. 2017. Connections between the human gut microbiome and gestational diabetes mellitus. GigaScience 6, 1–12.
Leylabadlo, H.E., Ghotaslou, R., Feizabadi, M.M., Farajnia, S., Moaddab, S.Y., Ganbarov, K., Khodadadi, E., Tanomand, A., Sheykhsaran, E., Yousefi, B., et al. 2020. The critical role of Faecalibacterium prausnitzii in human health: an overview. Microb. Pathog. 149, 104344.
Li, W., Sun, Y., Dai, L., Chen, H., Yi, B., Niu, J., Wang, L., Zhang, F., Luo, J., Wang, K., et al. 2021. Ecological and network analyses identify four microbial species with potential significance for the diagnosis/treatment of ulcerative colitis (UC). BMC Microbiol. 21, 138.
Liang, L., Yang, B., Jiang, N., Fu, W., He, X., Zhou, Y., Ma, W.L., and Wang, X. 2020. Three-month follow-up study of survivors of coronavirus disease 2019 after discharge. J. Korean Med. Sci. 35, e418.
Lopez-Siles, M., Duncan, S.H., Garcia-Gil, L.J., and Martinez-Medina, M. 2017. Faecalibacterium prausnitzii: from microbiology to diagnostics and prognostics. ISME J. 11, 841–852.
Martin-Gallausiaux, C., Marinelli, L., Blottière, H.M., Larraufie, P., and Lapaque, N. 2021. SCFA: mechanisms and functional importance in the gut. Proc. Nutr. Soc. 80, 37–49.
Miquel, S., Martín, R., Rossi, O., Bermúdez-Humarán, L.G., Chatel, J.M., Sokol, H., Thomas, M., Wells, J.M., and Langella, P. 2013. Faecalibacterium prausnitzii and human intestinal health. Curr. Opin. Microbiol. 16, 255–261.
Rosés, C., Cuevas-Sierra, A., Quintana, S., Riezu-Boj, J.I., Martínez, J.A., Milagro, F.I., and Barceló, A. 2021. Gut microbiota bacterial species associated with mediterranean diet-related food groups in a northern spanish population. Nutrients 13, 636.
Schirmer, M., Smeekens, S.P., Vlamakis, H., Jaeger, M., Oosting, M., Franzosa, E.A., Ter Horst, R., Jansen, T., Jacobs, L., Bonder, M.J., et al. 2016. Linking the human gut microbiome to inflammatory cytokine production capacity. Cell 167, 1125–1136.
Segain, J.P., de la Blétière, D.R., Bourreille, A., Leray, V., Gervois, N., Rosales, C., Ferrier, L., Bonnet, C., Blottière, H.M., and Galmiche, J.P. 2000. Butyrate inhibits inflammatory responses through NFκB inhibition: implications for Crohn’s disease. Gut 47, 397–403.
Shattock, P. and Whiteley, P. 2002. Biochemical aspects in autism spectrum disorders: updating the opioid-excess theory and presenting new opportunities for biomedical intervention. Expert Opin. Ther. Targets 6, 175–183.
Song, Y.L., Liu, C.X., McTeague, M., Summanen, P., and Finegold, S.M. 2004. Clostridium bartlettii sp. nov., isolated from human faeces. Anaerobe 10, 179–184.
Takahashi, K., Nishida, A., Fujimoto, T., Fujii, M., Shioya, M., Imaeda, H., Inatomi, O., Bamba, S., Sugimoto, M., and Andoh, A. 2016. Reduced abundance of butyrate-producing bacteria species in the fecal microbial community in Crohn’s disease. Digestion 93, 59–65.
Ticinesi, A., Mancabelli, L., Tagliaferri, S., Nouvenne, A., Milani, C., Del Rio, D., Lauretani, F., Maggio, M.G., Ventura, M., and Meschi, T. 2020. The gut-muscle axis in older subjects with low muscle mass and performance: a proof of concept study exploring fecal microbiota composition and function with shotgun metagenomics sequencing. Int. J. Mol. Sci. 21, 8946.
Wang, X., Zhou, Q., He, Y., Liu, L., Ma, X., Wei, X., Jiang, N., Liang, L., Zheng, Y., Ma, L., et al. 2020a. Nosocomial outbreak of COVID-19 pneumonia in Wuhan, China. Eur. Respir. J. 55, 2000544.
Wang, X., Zhou, Y., Jiang, N., Zhou, Q., and Ma, W.L. 2020b. Persistence of intestinal SARS-CoV-2 infection in patients with COVID-19 leads to re-admission after pneumonia resolved. Int. J. Infect. Dis. 95, 433–435.
Wei, Y., Li, Y., Yan, L., Sun, C., Miao, Q., Wang, Q., Xiao, X., Lian, M., Li, B., Chen, Y., et al. 2020. Alterations of gut microbiome in autoimmune hepatitis. Gut 69, 569–577.
Williams, O.M., Brazier, J., Peraino, V., and Goldstein, E.J. 2010. A review of three cases of Clostridium aldenense bacteremia. Anaerobe 16, 475–477.
Yeoh, Y.K., Zuo, T., Lui, G.C.Y., Zhang, F., Liu, Q., Li, A.Y.L., Chung, A.C.K., Cheung, C.P., Tso, E.Y.K., Fung, K.S.C., et al. 2021. Gut microbiota composition reflects disease severity and dysfunctional immune responses in patients with COVID-19. Gut 70, 698–706.
Yu, J., Feng, Q., Wong, S.H., Zhang, D., Liang, Q.Y., Qin, Y., Tang, L., Zhao, H., Stenvang, J., Li, Y., et al. 2017. Metagenomic analysis of faecal microbiome as a tool towards targeted non-invasive biomarkers for colorectal cancer. Gut 66, 70–78.
Zhou, Y., Shi, X., Fu, W., Xiang, F., He, X., Yang, B., Wang, X., and Ma, W.L. 2021. Gut microbiota dysbiosis correlates with abnormal immune response in moderate COVID-19 patients with fever. J. Inflamm. Res. 14, 2619–2631.
Zuo, T., Liu, Q., Zhang, F., Lui, G.C., Tso, E.Y., Yeoh, Y.K., Chen, Z., Boon, S.S., Chan, F.K., Chan, P.K., et al. 2021. Depicting SARS-CoV-2 faecal viral activity in association with gut microbiota composition in patients with COVID-19. Gut 70, 276–284.
Zuo, T., Zhan, H., Zhang, F., Liu, Q., Tso, E.Y.K., Lui, G.C.Y., Chen, N., Li, A., Lu, W., Chan, F.K.L., et al. 2020a. Alterations in fecal fungal microbiome of patients with COVID-19 during time of hospitalization until discharge. Gastroenterology 159, 1302–1310.
Zuo, T., Zhang, F., Lui, G.C.Y., Yeoh, Y.K., Li, A.Y.L., Zhan, H., Wan, Y., Chung, A.C.K., Cheung, C.P., Chen, N., et al. 2020b. Alterations in gut microbiota of patients with COVID-19 during time of hospitalization. Gastroenterology 159, 944–955.
Acknowledgements
This study gratefully acknowledges the patients who participated in the research. This work was supported by the Fundamental Research Funds for the Central Universities (No. 2020kfyXGYJ034) and (No. 2020kfyXGYJ009).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest The authors declare that there is no conflict of interest.
Ethical Statement The study was approved by the Ethics Committee of the Wuhan Union Hospital (2020-0149-02). All participants provided written informed consent prior to participation.
Additional information
Supplemental material for this article may be found at http://www.springerlink.com/content/120956.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Zhou, Y., Zhang, J., Zhang, D. et al. Linking the gut microbiota to persistent symptoms in survivors of COVID-19 after discharge. J Microbiol. 59, 941–948 (2021). https://doi.org/10.1007/s12275-021-1206-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-021-1206-5