Abstract
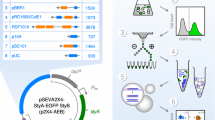
Plasmids are circular DNA molecules that are used as gene vectors in countless applications. They are easy to manipulate and transfer, but can also vary considerably in copy number between individual cells. In this article, a model expression system with a large amount of heterogeneity is presented. We used flow cytometry and digital PCR to count the number of plasmids in sorted sub-populations. Finally, heterogeneity could be attributed to the expression plasmid being either present or absent in bacterial sub-populations.
Similar content being viewed by others
Literatur
Friehs K (2004) Plasmid copy number and plasmid stability. Adv Biochem Eng Biotechnol 86:47–82
Kittleson JT, Cheung S, Anderson JC (2011) Rapid optimization of gene dosage in E. coli using DIAL strains. J Biol Eng 5:10
Kroll J, Klinter S, Schneider C et al. (2010) Plasmid addiction systems: perspectives and applications in biotechnology. Microb Biotechnol 3:634–657
Segura A, Molina L, Fillet S et al. (2012) Solvent tolerance in Gram-negative bacteria. Curr Opin Biotechnol 23:415–421
Jahn M, Vorpahl C, Türkowsky D et al. (2014) Accurate determination of plasmid copy number of flow-sorted cells using droplet digital PCR. Anal Chem 86:5969–5976
Jahn M, Seifert J, von Bergen M et al. (2013) Subpopulation-proteomics in prokaryotic populations. Curr Opin Biotechnol 24:79–87
Hindson BJ, Ness KD, Masquelier DA et al. (2011) Highthroughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem 83:8604–8610
Gürtler P, Gerdes L (2014) Digitale Polymerase kettenreaktion (dPCR). BIOspektrum 6:632–635
Whale AS, Huggett JF, Cowen S et al. (2012) Comparison of microfluidic digital PCR and conventional quantitative PCR for measuring copy number variation. Nucleic Acids Res 40:e82
Hindson CM, Chevillet JR, Briggs HA et al. (2013) Absolute quantification by droplet digital PCR versus analog real-time PCR. Nat Methods 10:1003–1005
Summers DK (1991) The kinetics of plasmid loss. Trends Biotechnol 9:273–278
Jahn M, Günther S, Müller S (2015) Non-random distribution of macromolecules as driving forces for phenotypic variation. Curr Opin Microbiol 25:49–55
Lindmeyer M, Jahn M, Vorpahl C et al. (2015) Variability in subpopulation formation propagates into biocatalytic variability of engineered Pseudomonas putida strains. Front Microbiol 6:1042
Author information
Authors and Affiliations
Corresponding author
Additional information
Michael Jahn, Carsten Vorpahl, Dominique Türkowsky, Susann Müller (v. l. n. r.)
Rights and permissions
About this article
Cite this article
Jahn, M., Vorpahl, C., Türkowsky, D. et al. Akkurate Bestimmung der Plasmidkopienzahl pro Zelle. Biospektrum 22, 211–213 (2016). https://doi.org/10.1007/s12268-016-0674-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12268-016-0674-3