Abstract
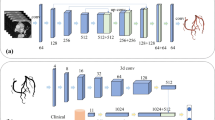
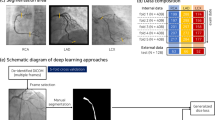
The visual inspection of coronary artery stenosis is known to be significantly affected by variation, due to the presence of other tissues, camera movements, and uneven illumination. More accurate and intelligent coronary angiography diagnostic models are necessary for improving the above problems. In this study, 2980 medical images from 949 patients are collected and a novel deep learning-based coronary angiography (DLCAG) diagnose system is proposed. Firstly, we design a module of coronary classification. Then, we introduce RetinaNet to balance positive and negative samples and improve the recognition accuracy. Additionally, DLCAG adopts instance segmentation to segment the stenosis of vessels and depict the degree of the stenosis vessels. Our DLCAG is available at http://101.132.120.184:8077/. When doctors use our system, all they need to do is login to the system, upload the coronary angiography videos. Then, a diagnose report is automatically generated.
Graphical Abstract






Similar content being viewed by others
Data Availability
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation, to any qualified researcher upon request.
References
Khera AV, Kathiresan S. Genetics of coronary artery disease: discovery, biology and clinical translation. Nat Rev Genet. 2017;18:331–44.
GBD 2015 Mortality and Causes of Death Collaborators. Global, regional, and national life expectancy, all-cause mortality, and cause-specific mortality for 249 causes of death, 1980–2015: a systematic analysis for the Global Burden of Disease Study 2015. Lancet. 2016;388:1459–1544.
Lemkes JS. Coronary angiography after cardiac arrest: a deep dive for PEARL. Circulation. 2020;142:2013–5.
Adjedj J, Xaplanteris P, Toth G, Ferrara A, Pellicano M, Ciccarelli G, Floré V, Barbato E and De Bruyne B. Visual and quantitative assessment of coronary stenoses at angiography versus fractional flow reserve: the impact of risk factors. Circ Cardiovasc Imaging. 2017;10(7):e006243.
Brilakis ES, Mashayekhi K, Tsuchikane E, Abi Rafeh N, Alaswad K, Araya M, Avran A, Azzalini L, Babunashvili AM, Bayani B, Bhindi R, Boudou N, Boukhris M, Božinović N, Bryniarski L, Bufe A, Buller CE, Burke MN, Büttner HJ, Cardoso P, Carlino M, Christiansen EH, Colombo A, Croce K, Damas de Los Santos F, De Martini T, Dens J, Di Mario C, Dou K, Egred M, ElGuindy AM, Escaned J, Furkalo S, Gagnor A, Galassi AR, Garbo R, Ge J, Goel PK, Goktekin O, Grancini L, Grantham JA, Hanratty C, Harb S, Harding SA, Henriques JPS, Hill JM, Jaffer FA, Jang Y, Jussila R, Kalnins A, Kalyanasundaram A, Kandzari DE, Kao HL, Karmpaliotis D, Kassem HH, Knaapen P, Kornowski R, Krestyaninov O, Kumar AVG, Laanmets P, Lamelas P, Lee SW, Lefevre T, Li Y, Lim ST, Lo S, Lombardi W, McEntegart M, Munawar M, Navarro Lecaro JA, Ngo HM, Nicholson W, Olivecrona GK, Padilla L, Postu M, Quadros A, Quesada FH, Prakasa Rao VS, Reifart N, Saghatelyan M, Santiago R, Sianos G, Smith E, J CS, Stone GW, Strange JW, Tammam K, Ungi I, Vo M, Vu VH, Walsh S, Werner GS, Wollmuth JR, Wu EB, Wyman RM, Xu B, Yamane M, Ybarra LF, Yeh RW, Zhang Q and Rinfret S. Guiding principles for chronic total occlusion percutaneous coronary intervention. Circulation. 2019;140:420–433.
Zhang H, Mu L, Hu S, Nallamothu BK, Lansky AJ, Xu B, Bouras G, Cohen DJ, Spertus JA, Masoudi FA, Curtis JP, Gao R, Ge J, Yang Y, Li J, Li X, Zheng X, Li Y, Krumholz HM, Jiang L. Comparison of physician visual assessment with quantitative coronary angiography in assessment of stenosis severity in China. JAMA Intern Med. 2018;178:239–47.
LeCun Y, Bengio Y, Hinton G. Deep learning. Nature. 2015;521:436–44.
Esteva A, Robicquet A, Ramsundar B, Kuleshov V, DePristo M, Chou K, Cui C, Corrado G, Thrun S, Dean J. A guide to deep learning in healthcare. Nat Med. 2019;25:24–9.
Isensee F, Jaeger PF, Kohl SAA, Petersen J, Maier-Hein KH. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods. 2021;18:203–11.
Milea D, Najjar RP, Jiang Z, Ting D, Vasseneix C, Xu X, AghsaeiFard M, Fonseca P, Vanikieti K, Lagrèze WA, La Morgia C, Cheung CY, Hamann S, Chiquet C, Sanda N, Yang H, Mejico LJ, Rougier M-B, Kho R, Tran THC, Singhal S, Gohier P, Clermont-Vignal C, Cheng C-Y, Jonas JB, Yu-Wai-Man P, Fraser CL, Chen JJ, Ambika S, Miller NR, Liu Y, Newman NJ, Wong TY, Biousse V. Artificial intelligence to detect papilledema from ocular fundus photographs. N Engl J Med. 2020;382:1687–95.
SippensGroenewegen A, Spekhorst H, Hemel NMv, Kingma JH, Hauer RNW, Barker JMTd, Grimbergen CA, Janse MJ and Dunning AJ. Value of body surface mapping in localizing the site of origin of ventricular tachycardia in patients with previous myocardial infarction. J Am Coll Cardiol. 1994;24:1708–1724.
Gong D, Wu L, Zhang J, Mu G, Shen L, Liu J, Wang Z, Zhou W, An P, Huang X, Jiang X, Li Y, Wan X, Hu S, Chen Y, Hu X, Xu Y, Zhu X, Li S, Yao L, He X, Chen D, Huang L, Wei X, Wang X, Yu H. Detection of colorectal adenomas with a real-time computer-aided system (ENDOANGEL): a randomised controlled study. Lancet Gastroenterol Hepatol. 2020;5:352–61.
He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. 2016 IEEE Conf Comput Vision Pattern Recogn (CVPR). 2016;6:770–778.
Zhu X, Li X, Ong K, Zhang W, Li W, Li L, Young D, Su Y, Shang B, Peng L, Xiong W, Liu Y, Liao W, Xu J, Wang F, Liao Q, Li S, Liao M, Li Y, Rao L, Lin J, Shi J, You Z, Zhong W, Liang X, Han H, Zhang Y, Tang N, Hu A, Gao H, Cheng Z, Liang L, Yu W, Ding Y. Hybrid AI-assistive diagnostic model permits rapid TBS classification of cervical liquid-based thin-layer cell smears. Nat Commun. 2021;12:3541.
Lin TY, Goyal P, Girshick R, He K, Dollar P. Focal loss for dense object detection. IEEE Trans Pattern Anal Mach Intell. 2020;42:318–27.
He K, Gkioxari G, Dollar P, Girshick R. Mask R-CNN. IEEE Trans Pattern Anal Mach Intell. 2020;42:386–97.
Li D, Bledsoe JR, Zeng Y, Liu W, Hu Y, Bi K, Liang A, Li S. A deep learning diagnostic platform for diffuse large B-cell lymphoma with high accuracy across multiple hospitals. Nat Commun. 2020;11:6004.
Huang K-C, Huang C-S, Su M-Y, Hung C-L, Tu Y-CE, Lin L-C, Hwang J-J. Artificial intelligence aids cardiac image quality assessment for improving precision in strain measurements. JACC Cardiovasc Imaging. 2021;14:335–45.
Moon JH, Lee DY, Cha WC, Chung MJ, Lee KS, Cho BH, Choi JH. Automatic stenosis recognition from coronary angiography using convolutional neural networks. Comput Methods Programs Biomed. 2021;198:105819.
Constantinescu L, Kim J, Feng DD. SparkMed: a framework for dynamic integration of multimedia medical data into distributed m-Health systems. IEEE Trans Inf Technol Biomed. 2012;16:40–52.
Cong C, Kato Y, Vasconcellos HD, Lima J and Venkatesh B. Automated stenosis detection and classification in X-ray angiography using deep neural network. 2019 IEEE Int Conf Bioinform Biomed (BIBM). 2019;11:18–21
Au B, Shaham U, Dhruva S, Bouras G, Krumholz H. Automated characterization of stenosis in invasive coronary angiography images with convolutional neural networks. 2018;7:1–12.
Quan L, Wu B, Mao S, Yang C, Li H. An instance segmentation-based method to obtain the leaf age and plant centre of weeds in complex field environments. Sensors (Basel). 2021;21(10):3389–3411.
Zreik M, van Hamersvelt RW, Wolterink JM, Leiner T, Viergever MA, Isgum I. A Recurrent CNN for automatic detection and classification of coronary artery plaque and stenosis in coronary CT angiography. IEEE Trans Med Imaging. 2019;38:1588–98.
Zreik M, van Hamersvelt RW, Wolterink JM, Leiner T, Viergever MA, Isgum I. Automatic detection and characterization of coronary artery plaque and stenosis using a recurrent convolutional neural network in coronary CT angiography. 2018;9:7–11.
Eberhard M, Nadarevic T, Cousin A, von Spiczak J, Hinzpeter R, Euler A, Morsbach F, Manka R, Keller DI, Alkadhi H. Machine learning-based CT fractional flow reserve assessment in acute chest pain: first experience. Cardiovasc Diagn Ther. 2020;10:820–30.
Funding
This work was supported by grants from the Department of Science and Technology of Jilin Province (No. 20200708112YY), the National Natural Science Foundation of China (No. 61972174 and 62172187).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no conflict of interest.
Additional information
Associate Editor Angela Taylor oversaw the review of this article
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ling, H., Chen, B., Guan, R. et al. Deep Learning Model for Coronary Angiography. J. of Cardiovasc. Trans. Res. 16, 896–904 (2023). https://doi.org/10.1007/s12265-023-10368-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12265-023-10368-8