Abstract
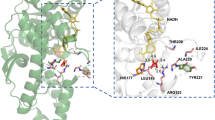
Native 3-hydroxybutyrate dehydrogenase from Alcaligenes faecalis can catalyze the reversible reduction of acetoacetate, a four carbon chain oxo acid. This enzyme has been engineered to enable the reduction of levulinic acid, with one carbon longer than acetoacetate. In this study, the native and engineered enzymes were subjected to the catalysis of oxo acids with a carbon chain length of 3 to 8, in order to examine the capability of the enzyme to work on various platform chemicals. The engineered enzyme could reduce the C7 and C8 oxo acids whereas the wild-type had no activity on these substrates. Docking simulation has indicated Tyr155 and Ser142 are key residues for the catalysis. In addition, stable hydrogen bond formation between Gln196 and the substrates affects the turnover rate. Mutation sites in the engineered enzyme were focused on creating larger active site volume for substrates with extended chain lengths. Both qualitative and quantitative structural basis for the enzyme substrate specificity on alpha, beta, gamma and omega hydroxy acids could be elucidated.
Similar content being viewed by others
References
Wohlgemuth, R. (2010) Biocatalysis-key to sustainable industrial chemistry. Curr. Opin. Biotechnol. 21: 713–724.
Privett, H. K., G. Kiss, T. M. Lee, R. Blomberg, R. A. Chica, L. M. Thomas, D. Hilvert, K. N. Houk, and S. L. Mayo (2012) Iterative approach to computational enzyme design. Proc. Natl. Acad. Sci. U.S.A. 109: 3790–3795.
Bornscheuer, U. T., G. W. Huisman, R. J. Kazlauskas, S. Lutz, J. C. Moore, and K. Robins (2012) Engineering the third wave of biocatalysis. Nature. 485: 185–194.
Greenhagen, B. T., P. E. O'maille, J. P. Noel, and J. Chappell (2006) Identifying and manipulating structural determinates linking catalytic specificities in terpene synthases. Proc. Natl. Acad. Sci. U.S.A. 103: 9826–9831.
Zhang, K., M. R. Sawaya, D. S. Eisenberg, and J. C. Liao (2008) Expanding metabolism for biosynthesis of nonnatural alcohols. Proc. Natl. Acad. Sci. U.S.A. 105: 20653–20658.
Savile, C. K., J. M. Janey, E. C. Mundorff, J. C. Moore, S. Tam, W. R. Jarvis, J. C. Colbeck, A. Krebber, F. J. Fleitz, J. Brands, P. N. Devine, G. W. Huisman, and G. J. Hughes (2010) Biocatalytic asymmetric synthesis of chiral amines from ketones applied to sitagliptin manufacture. Science. 329: 305–309.
Hoque, M. M., S. Shimizu, M. T. Hossain, T. Yamamoto, S. Imamura, K. Suzuki, M. Tsunoda, H. Amano, T. Sekiguchi, and A. Takenaka (2008) The structures of Alcaligenes faecalis D-3-hydroxybutyrate dehydrogenase before and after NAD+ and acetate binding suggest a dynamical reaction mechanism as a member of the SDR family. Acta Crystallogr. D Biol. Crystallogr. 64: 496–505.
Hoque, M. M., S. Shimizu, E. C. Juan, Y. Sato, M. T. Hossain, T. Yamamoto, S. Imamura, K. Suzuki, H. Amano, T. Sekiguchi, M. Tsunoda, and A. Takenaka (2009) Structure of D-3-hydroxybutyrate dehydrogenase prepared in the presence of the substrate D-3-hydroxybutyrate and NAD+. Acta Crystallogr. F Struct. Biol. Cryst. Commun. 65: 331–335.
Senior, P. J. and E. A. Dawes (1973) The regulation of poly-beta-hydroxybutyrate metabolism in Azotobacter beijerinckii. Biochem. J. 134: 225–238.
Lenz, R. W. and R. H. Marchessault (2005) Bacterial polyesters: Biosynthesis, biodegradable plastics and biotechnology. Biomacro-molecules. 6: 1–8.
Saadat, A., A. Behnamghader, S. Karbasi, D. Abedi, M. Soleimani, and A. Shafiee (2013) Comparison of acellular and cellular bioactivity of poly 3-hydroxybutyrate/hydroxyapatite nano-composite and poly 3-hydroxybutyrate scaffolds. Biotechnol. Bioprcess Eng. 18: 587–593.
Yeon, Y. J., H. Y. Park, and Y. J. Yoo (2013) Enzymatic reduction of levulinic acid by engineering the substrate specificity of 3-hydroxybutyrate dehydrogenase. Bioresource Technol. 134: 377–380.
Bozell, J. J., L. Moens, D. C. Elliott, Y. Wang, G. G. Neuenscwander, S. W. Fitzpatrick, R. J. Bilski, and J. L. Jarnefeld (2000) Production of levulinic acid and use as a platform chemical for derived products. Resour. Conserv. Recycl. 28: 227–239.
Bozell, J. J. and G. R. Petersen (2010) Technology development for the production of biobased products from biorefinery carbohydrates—the US department of energy's “top 10” revisited. Green Chem. 12: 539–554.
Zhang, Z. (2016) Synthesis of γ-valerolactone from carbohydrates and its applications. ChemSusChem. 9: 156–171.
Bond, J. Q., D. M. Alonso, D. Wang, R. M. West, and J. A. Dumesic (2010) Integrated catalytic conversion of gammavalerolactone to liquid alkenes for transportation fuels. Science. 327: 1110–1114.
Xiong, M., J. Deng, A. P. Woodruff, M. Zhu, J. Zhou, S. W. Park, H. Li, Y. Fu, and K. Zhang (2012) A bio-catalytic approach to aliphatic ketones. Sci Rep. 2: 311.
Kornhauser, A., S. G. Coelho, and V. J. Hearing (2010) Applications of hydroxy acids: Classification, mechanisms, and photoactivity. Clin. Cosmet. Investig. Dermatol. 3: 135–142.
Bhalla, T., V. Kumar, and S. Bhatia (2014) Hydroxy acids: production and applications. pp. 56–76. In: R. S. Singh, A. Pandey, and C. Larroche (eds.). Advances in Industrial Biotechnology. I. K. International Publishing House Pvt. Ltd., New Delhi, India
Mitra, A. K. and S. K. Samanta (2011) Drug conjugates. US Patent 12,999,149
Jestin, J.-L. and S. Vichier-Guerre (2005) How to broaden enzyme substrate specificity: Strategies, implications and applications. Res. Microbiol. 156: 961–966.
Wilks, H. M., D. J. Halsall, T. Atkinson, W. N. Chia, A. R. Clarke, and J. J. Holbrook (1990) Designs for a broad substrate specificity keto acid dehydrogenase. Biochemistry. 29: 8587–8591.
Yeon, Y. J., H. J. Park, H.-Y Park, and Y. J. Yoo (2014) Effect of his-tag location on the catalytic activity of 3-hydroxybutyrate dehydrogenase. Biotechnol. Bioprocess Eng. 19: 798–802.
Lee, H. S., J. Park, Y. J. Yoo, and Y. J. Yeon (2019) A novel D-2-hydroxy acid dehydrogenase with high substrate preference for phenylpyruvate originating from lactic acid bacteria: Structural analysis on the substrate specificity. Enzyme Microb. Technol. 125: 37–44.
Bradford, M. M. (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72: 248–254.
Schrödinger (2018) Release 2018-4: Prime, Schrödinger, LLC, New York, NY
Jacobson, M. P, D. L. Pincus, C. S. Rapp, T. J. Day, B. Honig, D. E. Shaw, and R. A. Friesner (2004) A hierarchical approach to all-atom protein loop prediction Proteins. 55: 351–367.
Schrödinger (2018) Release 2018-4: Ligprep, Schrödinger, LLC, New York, NY
Kenyon, V., I. Chorny, W. J. Carvajal, T. R. Holman, and M. P Jacobson (2006) Novel human lipoxygenase inhibitors discovered using virtual screening with homology models. J. Med. Chem. 49: 1356–1363.
Schrödinger (2018) Release 2018-4: Giide, Schrödinger, LLC, New York, NY
Friesner, R. A., R. B. Murphy, M. P Repasky, L. L. Frye, J. R. Greenwood, T. A. Halgren, P. C. Sanschagrin, and D. T. Mainz (2006) Extra precision glide: Docking and scoring incorporating a model of hydrophobic enclosure for protein-ligand complexes. J. Med. Chem. 49: 6177–6196.
Yeon, Y. J., H.-Y. Park, K. Park, H. J. Park, and Y. J. Yoo (2016) Structural basis for the substrate specificity of 3-hydroxybutyrate dehydrogenase. Biotechnol. Bioprocess Eng. 21: 364–372.
Hammes-Schiffer, S. (2002) Comparison of hydride, hydrogen atom, and proton-coupled electron transfer reactions. ChemPhysChem. 3: 33–42.
Acknowledgements
This research was supported by the C1 Gas Refinery Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Science, ICT & Future Planning (2015M3D3A1A01064929).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher’s Note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Lee, HS., Na, J.G., Lee, J. et al. Structure-based Mutational Studies of D-3-hydroxybutyrate Dehydrogenase for Substrate Recognition of Aliphatic Hydroxy Acids with a Variable Length of Carbon Chain. Biotechnol Bioproc E 24, 605–612 (2019). https://doi.org/10.1007/s12257-019-0135-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12257-019-0135-1