Abstract
Subtropical and tropical islands are undergoing rapid urbanization as the human population expands globally. Urbanization disrupts coastal ecosystems through several pathways—including the replacement of natural habitats with concrete structures that increase runoff pollution—but it remains difficult to isolate and characterize specific impacts of urbanization on marine ecosystems. The historical gradient in urbanization on the subtropical island of Okinawa, Japan, sets up a natural laboratory to study urbanization effects on nearshore ecosystems. Physicochemical parameters and bacterial community composition were assessed every 2 weeks for 1 year at two nearshore sites adjacent to watersheds with > 70% urban land use and two nearshore sites adjacent to watersheds with > 70% rural land use. Urbanization increased freshwater input and nutrient loading—indicated by decreased salinity and elevated nitrate + nitrite, ammonium, and phosphate at urban sites—despite the urban sites being more open to flushing due to land reclamation projects filling in the coral lagoon. Urbanization significantly altered microbial community composition by increasing diversity through the addition of fecal indicator and pathogenic bacteria—eight orders of bacteria were only detected in urban samples, whereas only Verrucomicrobiales was unique to rural samples. The change in microbial community composition at urban sites persisted throughout the seasonal cycle, suggesting a regime change or sustained disturbance. The altered physicochemical conditions and microbial communities at urban sites could degrade nearby coral reefs and their ecosystem services, highlighting the importance of coastal land management in marine conservation efforts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Human population density along coastlines is roughly three times the overall global average (Small and Nicholls 2003), and as human population growth continues, urbanization along coasts is accelerating (Seto et al. 2011; Bloom 2011). The intense concentration of humans in coastal areas disrupts adjacent ecosystems through three main pathways: resource exploitation, pollution, and ocean sprawl (Todd et al. 2019). Resource exploitation includes extraction of living (e.g., fishing, aquaculture) and non-living (e.g., dredging, mining, oil and gas extraction, water use for cooling or desalination) resources (Todd et al. 2019). Pollution near urban centers includes sediments, nutrients, plastic debris, chemicals, and pathogens, as well as increased light and noise (Todd et al. 2019; Carlson et al. 2019). Ocean sprawl includes the development of reclaimed land and artificial islands, artificial coastal defenses, ports, docks, marinas, oil infrastructure, and submarine cables and pipelines (Todd et al. 2019). Each pathway has its own impacts on coastal ecosystems, but they can also act together synergistically, leading to profoundly different ecological conditions near urban centers compared to pre-urbanization conditions (e.g., habitat and keystone species loss).
It is largely recognized that coastal urbanization negatively impacts coastal ecosystem health. However, many coastal cities have been human population centers since before ecosystem monitoring was initiated (Röthig et al. 2023), making it difficult to fully isolate and characterize the impact of urbanization on coastal ecosystems due to missing or shifting baselines (Roberts et al. 2017). Additional methodological issues, including the difficulty in identifying suitable whole-habitat replicates and lacking well-defined indices of urbanization across regions, further complicate measuring urbanization effects on ecosystems (Lee et al. 2006). The recent urbanization of many tropical and subtropical islands compared to continental coastlines allows for more direct measurement of the ecosystem effects of urbanization since more modern monitoring technologies were developed before or during urbanization and because island chains provide naturally replicated habitats in close geographic vicinity. For example, the relatively recent urbanization of the Caribbean Island of Roatan allowed for satellite data spanning the entire urbanization period to be analyzed to directly correlate urbanization with mangrove forest removal (Tuholske et al. 2017). In another study, biodiversity on urbanized atolls in the South Pacific was directly compared to nearby uninhabited atolls to demonstrate that urbanization significantly reduced biodiversity (Steibl et al. 2021).
While tropical and subtropical islands present unique opportunities to study urbanization effects on coastal ecosystems, they are also uniquely vulnerable to urbanization (Russell and Kueffer 2019). Oceanic islands are biodiversity hotspots but are simultaneously experiencing some of the highest rates of species loss due to anthropogenic pressure (Kier et al. 2009). Smaller overall area on islands means that more urban development is closer to the coast (Cao et al. 2017) and there is a higher likelihood of land reclamation projects being undertaken (Heery et al. 2018; Masucci and Reimer 2019). Islands also attract large volumes of tourists that can overwhelm local facilities and lead to more pollution and habitat degradation (DeGeorges et al. 2010; Baum et al. 2015). However, since urbanization is ongoing in many island systems, it may still be possible to minimize impacts on ecosystem functioning through research-informed sustainable development (Ghina 2003).
Like many tropical and subtropical islands, Okinawa Island (Japan) has undergone urbanization and population growth relatively recently (Kobayashi 2022; Uehara et al. 2019), with rapid urbanization commencing after 1972 when administrative control of the island was returned to Japan from the U.S.A. (Li et al. 2018). Sprawling high-density population centers now fill the southern third of the island, while the northern third of the island remains rural (i.e., composed of forests, agricultural operations, and small villages). Alongside urbanization, Okinawa’s nearshore marine ecosystems have been subjected to resource exploitation (Kobayashi 2022; Uehara et al. 2019), pollution (Ares et al. 2020; Li et al. 2018), and ocean sprawl (Masucci and Reimer 2019)—the three main pathways through which urbanization impacts coastal ecosystems (Todd et al. 2019). Okinawa has a strong land-sea connection due to frequent and intense rains associated with spring monsoons and summer/fall typhoons (Singh et al. 2022). Consequently, ongoing environmental challenges related to urbanization—particularly stormwater runoff and associated pollutants, such as wastewater and sediment pollution (red soil)—may be especially relevant in Okinawa (Ares et al. 2020). In contrast, the oceanic setting of Okinawa may dampen urbanization effects on nearshore ecosystems due to strong coastal currents and dilution effects (Ares et al. 2020; Rintoul et al. 2022).
In this study, we leveraged the natural laboratory set up by Okinawa’s urbanization pattern to investigate the impact of intense urbanization on the nearshore ecosystem of a subtropical oceanic island. We assessed microbial community composition (i.e., 16S rRNA gene metabarcode analysis) and physicochemical environmental parameters every 2 weeks for 1 year at four sites with varying levels of urbanization based on land cover classification from satellite imagery. We focused on microbial community composition since microbes respond rapidly to changing physical and chemical conditions, leading to shifts in microbial community dynamics being detectable before responses in economically important macro-organisms—such as corals, mollusks, fish, or aquacultured seaweeds (McLellan et al. 2015). Moreover, specific microbes are indicators of pollution (e.g., fecal indicator bacteria) and some microbes are considered pollutants themselves (e.g., the coral pathogen Serratia marcescens; Sutherland et al. 2011). Thus, shifting microbial communities and the presence of specific microbes can indicate or predict degradation of nearshore marine ecosystems (Becker et al. 2023). By analyzing the microbial community at urban and rural sites across a relatively high-resolution time series, this study will disentangle seasonal variability from anthropogenic influences, allowing for better understanding of how urbanization changes coastal microbial community dynamics. Persistent changes in microbial community dynamics at urban sites would suggest that increased coastal management is needed in Okinawa, as well as in other urbanizing island systems.
Methods
Sampling Area Description
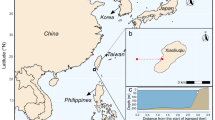
Nearshore seawater samples were collected along the west coast of Okinawa Island in the Ryukyu archipelago, South Japan. The island shows well-defined dominant land uses, with concentrated urbanized watersheds primarily situated in the southern third of the island (Fig. 1). As an exception, Nago City, a major city on the island, lies in the northern part of the island. Four primary sampling sites were selected based on watershed size and the percent of the watershed with land cover classified as urban. Watersheds were delineated and their land areas were calculated using a digital elevation model (DEM) interpolated at 30 m based on a 2008 10-m LiDAR survey provided by the Geospatial Information Authority of Japan (https://fgd.gsi.go.jp/download/menu.php). Land cover classification was based on a Jan 4, 2015, Landsat 8 Operational Land Imager image obtained from the US Geological Survey (Scene: LC81130422015004LGN00) following methods described in Ross et al. (2018). Land cover classes were defined as agriculture, dominated by sugarcane and other crops at various stages; forest, dense tree stands with closed canopy; scrub, short woody vegetation without tree canopy; grass, short trimmed vegetation found in golf courses and airfields; rock/dirt; sand; urban, a complex mix of man-made surface materials such as concrete and asphalt with limited vegetation; water, including fresh and marine bodies of water; and unclassified, pixels that could not be classified (Fig. 1).
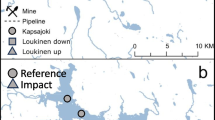
Geographic setting of sampling sites and subsites on the west coast of Okinawa Island, Japan. Left panel: Map of Okinawa Island with land cover classifications based on satellite data. Thin black lines indicate watershed boundaries based on a LiDAR digital elevation model. Primary sampling sites are highlighted by color coded boxes (purple, urban sites; blue, rural sites) with points demarcating the subsites where samples were collected. Right panels: Detailed maps of the four primary sampling sites with the subsites demarcated as points. Point color indicates the subsites—blue, south (S); red, central (C); yellow, north (N)—where water samples and measurements were taken. Satellite true-color imagery is from Google Earth
The four primary sampling sites (Urban 1 (U1) - Ginowan, Urban 2 (U2) - Nago, Rural 1 (R1) - Tancha, and Rural 2 (R2) - Ogimi) are located in watersheds of similar size (407,002–993,281 m2) and are classified as urban or rural based on the percent of the watershed with land cover classified as urban. Specifically, urban primary sites U1 and U2 are in watersheds with > 70% of land cover classified as urban and rural primary sites R1 and R2 are in watersheds with > 70% land cover classified as agriculture, forest, scrub, grass, rock/dirt, and sand. In addition, both urban sites have artificial shorelines created from land reclamation projects that filled in the coral lagoon (Masucci and Reimer 2019). Primary sampling sites were centered on the freshwater outflow point into the ocean from the study watershed. Three subsites at each primary site were designated for sample collection based on their proximity to the point source of freshwater outflow: central (C) subsites are perpendicularly offshore to the outflow and North (N) and South (S) subsites were ~ 200 m North and South of the outflows, respectively. Subsites were sampled synoptically every other week for 1 year, from September 17, 2020, to September 2, 2021, leading to 25 sampling events in total.
Seawater Sampling
Nearshore surface seawater was collected for DNA metabarcoding by submerging acid-cleaned 500-mL Nalgene bottles below the sea surface in the uppermost 20–50 cm of the water column. Seawater for dissolved macronutrient analysis was collected in acid-cleaned 50-mL Falcon tubes. A total of 300 of each sample type was collected: four primary sampling sites, three subsites per primary site, and 25 collection events. Samples were transported to the lab on ice and in the dark. After transport, seawater samples for metabarcoding were immediately filtered through 0.2-µm pore-size Polytetrafluoroethylene filters (Millipore) under gentle vacuum pressure, and filters were stored at − 80 °C for later DNA extraction. Samples for dissolved macronutrient analysis were syringe-filtered and stored at − 20 °C until chemical analysis. Physicochemical properties—dissolved oxygen (DO), salinity, sea surface temperature (SST), turbidity, and chlorophyll a fluorescence (Chla a)—were measured at each subsite with a RINKO conductivity, temperature, and depth (CTD) probe (JFE Advantech, Japan).
Nutrient Analyses
Nutrient concentrations—including nitrate (NO3−), nitrite (NO2−), ammonium (NH4+), phosphate (PO43−), and silica (SiO2)—were determined on a QuAAtro39 Continuous Segmented Flow Analyzer (SEAL Analytical) following manufacturer guidelines. Final concentrations were calculated through AACE software (SEAL Analytical). Nutrient analysis was carried out at the Okinawa Prefecture Fisheries and Ocean Technology Center.
DNA Extraction and Metabarcode Sequencing
DNA was extracted from frozen filters following the manufacturer’s protocol for the DNeasy Power Water Kit (Qiagen), including the optional heating step. Metabarcode sequencing libraries were prepared for the V3–V4 region of the bacterial 16S ribosomal RNA gene following Illumina’s 16S Metagenomic Sequencing Library Preparation manual without any modifications. The V3–V4 region of the 16S rRNA gene provides good taxonomic resolution for environmental bacterial communities and is widely sequenced for such studies, increasing the intercomparability of study results (Klindworth et al. 2013). Sequencing was performed by the Okinawa Institute of Science and Technology Sequencing Center using 2 × 300-bp v3 chemistry on the Illumina MiSeq platform. Out of 300 samples, 291 produced sequencing results that passed all quality filters. Overall, 53 million sequencing reads were generated, with 62,656–612,998 sequencing reads per sample (mean = 182,814). Sequencing data are available from the NCBI Sequencing Read Archive (SRA) under the accession PRJNA1044524.
Bioinformatic and Statistical Analyses
Sequencing reads were denoised using the Divisive Amplicon Denoising Algorithm (Callahan et al. 2016) with the DADA2 plug-in for QIIME 2 (Bolyen et al. 2019). Taxonomy was assigned to representative amplicon sequence variants (ASVs) using a naive Bayes classifier trained on the SILVA 99% consensus taxonomy (version 132; Quast et al. 2013) with the QIIME 2 feature-classifier plug-in (Bokulich et al. 2018). The results were imported into the R statistical environment (R Core Team 2018) for further analysis with the phyloseq (Mcmurdie and Holmes 2013) and vegan (Oksanen et al. 2019) R packages. Rarefaction sampling was performed and plotted with the ggrare function, and all samples reached richness saturation within their total sample size (Fig. S1). Alpha diversity estimates (richness and Shannon index) were determined with the breakaway R package (Willis et al. 2017), and the statistical significance of differences in mean alpha diversity between urban and rural sites each month was tested with pairwise Wilcox tests. To minimize compositional bias inherent in metabarcoding data, we used the Aitchison distance between samples, which includes a centered log-ratio transformation to normalize data (Gloor et al. 2017), for principal coordinates analyses (PCoA). Permutational multivariate analyses of variance (PERMANOVA) on Aitchison distances were performed with the adonis2 function (999 permutations) in the vegan R package to test whether shifts in community composition between sample types were statistically significant (Oksanen et al. 2019). The influence of environmental parameters in shaping bacterial community composition was investigated through redundancy analysis (RDA) and variance partitioning using functions from the vegan R package. Lastly, the SPIEC-EASI R package was used for network analysis and to infer co-occurrence patterns between taxa (Kurtz et al. 2015). Intermediate data files and the code necessary to replicate analyses are available in a GitHub repository (https://github.com/maggimars/UrbanOki) where an interactive HTML document (https://maggimars.github.io/UrbanOki/Amplicons.html) can also be found.
Results
Land Cover Effects on Physicochemical Parameters in Nearshore Ecosystems
The annual pattern in SST was nearly identical across sites and subsites (Fig. 2). In contrast, salinity was lower and more variable at the urban sites compared to rural sites, especially at the central and north subsites for both urban sampling sites (Fig. 2). Turbidity was variable at all locations and has multiple causes–including phytoplankton growth, sediment resuspension, and soil pollution (Fig. 2). The percent saturation of dissolved oxygen (DO) was highest at the northern rural site (R2 - Ogimi; all subsites) compared to the other three study sites (Fig. 2), potentially caused by more wave action in that region. Nitrate + nitrite, ammonium, and phosphate concentrations were low at rural sites, whereas these macronutrients were elevated at urban sites—particularly at the central and north subsites of the U1 - Ginowan sampling area (Fig. 3). Silica concentrations were variable at all sites (Fig. 3). Decreases in salinity were strongly correlated with high nitrate + nitrite concentrations at urban sites (R = − 0.65, p = 0) and were also correlated with increased phosphate (R = − 0.454, p = 0) and Chl a (R = − 0.29, p = 0; Fig. 4). Nitrate + nitrite concentrations were also strongly correlated with phosphate concentrations at urban sites (R = 0.76, p = 0) and chlorophyll a was positively correlated with nitrate + nitrite (R = 0.34, p = 0), ammonium (R = 0.27, p = 0.001), and phosphate (R = 0.26, p = 0.001) at urban sites but not rural sites (Fig. 4). Overall, chlorophyll a fluorescence was higher at urban sites (U1 = 0.47 ± 0.03 RFU, U2 = 0.43 ± 0.05 RFU) compared to rural sites (R1 = 0.34 ± 0.03 RFU, R2 = 0.18 ± 0.01 RFU), particularly at central and north subsites (Fig. S2). At rural sites, salinity was significantly negatively correlated with silica concentrations (R = − 0.36, p = 0; Fig. 3).
Time series of physical parameters measured at nearshore urban and rural sites along the west coast of Okinawa Island, Japan. Sea surface temperature (SST), salinity, turbidity, and dissolved oxygen (DO) were measured at each subsite (south, central, and north) within each primary sampling site (U1, U2, R1, and R2) using a RINKO CTD probe. Plots are faceted by primary sampling site (vertically) and parameter (horizontally). Point and line color represent subsites—blue, south (S); red, central (C); yellow, north (N)—where water samples and measurements were taken. The annual pattern in SST was nearly identical across sites and subsites. Salinity was lower and more variable at the urban sites, especially at central and north subsites. Turbidity was variable at all locations. The percent saturation of DO was highest at the northern rural site (R2 - Ogimi)
Time series of chemical parameters measured at nearshore urban and rural areas along the west coast of Okinawa Island, Japan. Nitrate + nitrite (NO2 + NO3), ammonium (NH4), phosphate (PO4), and silica (SiO2) were measured in water samples collected from subsites (south, central, and north) within each primary sampling site (U1, U2, R1, and R2) with a QuAAtro39 Continuous Segmented Flow Analyzer. Plots are faceted by primary sampling site (vertically) and parameter (horizontally). Point and line color represents subsites—blue, south (S); red, central (C); yellow, north (N)—where water samples and measurements were taken. Nitrate + nitrite, ammonium, and phosphate concentrations were rarely above the detection limit at rural sites, while these macronutrients were elevated at urban sites—particularly the central and north subsites of the U1 - Ginowan primary sampling site. Silica was variable at all sites
Correlation between environmental parameters at urban and rural sites on the west coast of Okinawa Island, Japan. Values for all parameters were z-scaled and the Pearson correlation coefficients were calculated for all parameters at the two urban primary sites (U1 - Ginowan, U2 - Nago) and the two rural primary sites (R1 - Tancha, R2 - Ogimi). Correlation coefficients are visualized for all comparisons by both point size and color: darker blue colors are more positively correlated, darker red colors are more negatively correlated, and point size reflects the absolute value of the correlation coefficient. The statistical significance of each correlation coefficient was evaluated with the cor.mtest function in the corrplot R package. Correlation coefficients were considered statistically significant if the p-value was ≤ 0.01 and significant correlations are marked with an asterisk (*) on the plot
Effects of Land Cover on Nearshore Bacterial Communities
We used metabarcode analysis with the 16S rRNA gene to investigate the effect of land cover (urban or rural) on nearshore bacterial community composition across a biweekly time series spanning a full year. Alpha diversity metrics (ASV richness and Shannon index) were lower in warmer months (May–August) compared to cooler months at both urban and rural sites (Fig. S3). However, the mean richness and Shannon indices were higher at urban sites compared to rural sites across all months (Fig. S3), with the difference in richness statistically significant in six out of twelve months (Jan, March, June, July, Oct, and Dec; Table S1) and the difference in Shannon index statistically significant in eight out of twelve months (Jan, March June, July, Aug, Sept, Oct, Dec; Table S1). The bacterial community compositions in nearshore waters of urban sites were significantly different from the communities in the nearshore waters of rural sites (PERMANOVA, 999 permutations, F = 8.5, p = 0.001; Fig. S4). PCoA plots for nearshore bacterial communities in urban and rural regions showed contrasting patterns (Fig. 5). The warmer months (May through the beginning of August) formed tight clusters in the PCoA plots for both rural (Fig. 5A) and urban sites (Fig. 5B). In the PCoA plot for rural sites, samples from seasons cluster so that they are separated roughly by quadrant, with winter samples in the top right, spring and early summer samples in the top left, samples from late summer in the bottom left, and autumn samples in the bottom right. In the PCoA plot for urban sites, samples separate based on which urban site they originated from, with more samples from U1 - Ginowan in the bottom half of the plot and more samples from U2 - Nago in the top half of the plot. These differences in clustering patterns suggest that the seasonal succession cycle is disrupted at urban sites compared to rural sites. However, results from PERMANOVA analyses on the season and primary site variables were significant for both rural and urban sites (Table 1). The subsite variable (i.e., the proximity to a freshwater outlet) was only significant for urban sites (F = 3.63, p = 0.001; Table 1).
Principal coordinates analysis (PCoA) plots of Aitchison distances between bacterial communities through time at two nearshore sites adjacent to rural areas (A) and two nearshore sites adjacent to urban areas (B). Shape indicates sampling sites, with circles representing the more southern site (site 1) of both the rural and urban sites, and triangles representing the more northern sites (site 2). The color indicates the month of the year, with blues representing winter months, purples representing spring months, greens representing summer, and yellow–red representing autumn. Samples from late spring and early summer form tight clusters in both plots. Samples from rural sites (A) separate into quadrants based on the season (winter; top right, autumn; bottom right, late summer; bottom left, spring and early summer; top left), whereas samples from the two urban sites (B) separate by site location on the secondary (y) axis and the seasonal cycle is not visible
We performed a redundancy analysis (RDA) to visualize which environmental variables contributed to the clustering observed in PCoA ordination plots (Fig. S5). An analysis of variance (ANOVA) was run on the RDA to determine if model results were significant and an ANOVA by term was used to test which variables’ contributions were statistically significant (Table S2). Variance partitioning was calculated for all significant variables. SST significantly affected community composition clustering among samples from both urban (F = 3.6, p = 0.004, variance partition = 0.7%) and rural sites (F = 4.2, p = 0.012, variance partition = 2.2%). Dissolved oxygen (F = 13.4, p = 0.001, variance partition = 7.4%), phosphate (F = 6.9, p = 0.004, variance partition = 3.1%), and salinity (F = 2.83, p = 0.045, variance partition = 0.6%) were also significant determinants of community composition in rural samples. In contrast, nitrate + nitrite (F = 5.4, p = 0.001, variance partition = 1.5%) was the only significant determinant of community composition in urban samples after SST (Fig. S5, Table S2).
Plotting the relative abundance of ASVs grouped by bacterial order showed clear differentiation between samples collected at urban and rural sites (Figs. 6 and S6). Eight bacterial orders were present in three or more urban samples but absent in all rural samples (Fig. 6; highlighted in red) and nine orders were prevalent among urban samples but rare in rural samples (Fig. 6; highlighted in orange). The cumulative relative abundance of these orders ranged from 1.3 to 63.7% (mean = 19.6%) at subsites in Ginowan (U1) and 1.1–57.9% (mean = 15.4%) at subsites in Nago (U2), whereas the cumulative relative abundance of these orders never exceeded 9.8% at rural sites (Fig. 7). Members of these orders were more abundant at the central and north subsites of the two urban sampling sites than at the south subsites (Fig. 7), which are also the subsites that appear more impacted by runoff and freshwater input based on physicochemical parameters (Figs. 2 and 3). Interestingly, seven of these orders were shown to covary in urban samples through network analysis (Fig. 8a; Pseudomonadales, Clostridiales, Saccharimonadales, Campylobacterales, Bacteroidales, Betaproteobacteriales, Thiotrichales). The covariance of these orders suggests they may share a common source or that their growth is supported by shared resources associated with urbanization.
Relative abundance of major bacterial orders in each sample. Color represents the relative abundance of each order and was determined by summing the relative abundance of each ASV classified as belonging to the order. Samples are grouped by primary site and columns represent subsites (S; southern subsite, C; central subsite, and N; northern subsite). Order names are highlighted as follows: Orange; orders that are prevalent among samples from urban sites but rarely found in rural samples, Red; orders that are prevalent among or present in ≥ 3 samples from urban sites but not found in any rural samples, Blue; orders that are prevalent among or present in ≥ 3 samples from rural sites but not found in any urban samples
Relative abundance of bacterial orders identified as either prevalent among urban samples and rare in rural samples or prevalent among/present in ≥ 3 urban samples and absent in all rural samples. Samples are grouped by site and subsite (S; southern subsite, C; central subsite, and N; northern subsite) and stacked bar chart fill colors indicate bacterial order. All orders highlighted in red or orange in Fig. 6 are included. Orders that are rare at rural sites make up a large proportion of communities in urban samples, particularly at the central and northern subsites
Covariance networks for bacterial orders detected at urban and rural sites. Orange nodes represent orders that are prevalent among samples from urban sites but were rarely observed in rural samples. Red nodes represent orders that are prevalent among or present in ≥ 3 samples from urban sites but not found in any rural samples. Green edges connecting nodes indicate significantly positive covariance between node orders (no negative relationships could be detected). There is a cluster composed solely of “red” (Pseudomonadales, Clostridiales, Saccharimonadales) and “orange” (Campylobacterales, Bacteroidales, Betaproteobacteriales, Thiotrichales) orders in the urban site network, indicating that these orders co-occur across samples collected from the urban sites
Discussion
Coastlines around the world are rapidly being urbanized, with widespread consequences for adjacent marine ecosystems. Despite the apparent impacts of urbanization in many nearshore areas, it can be difficult to fully assess the effects due to shifting or absent baselines and methodological challenges. In this study, we investigated the impacts of urbanization on the physicochemical conditions and microbial communities in nearshore marine ecosystems. Studying microbial responses to urbanization is a good first step in assessing the impacts of urbanization because changes in microbial communities can indicate disruption to the ecosystem more broadly (Nogales et al. 2011). Okinawa Island has urbanized relatively recently compared to many other coastal urban centers and the pattern of urbanization—with high-density urban centers in the south and rural regions in the north—allows for ecological effects of urbanization to be investigated more directly. We detected an altered physicochemical setting in nearshore waters along urban coastlines compared to rural settings, with increased nutrient concentrations at urban sites. Microbial communities at urban sites were significantly different from communities at rural sites in all seasons. The differences largely reflected increased microbial diversity at urban sites due to the presence of bacterial taxa that were absent or rare at rural sites. Many of the additional taxa found at urban sites are associated with anthropogenic sources, such sewage treatment facilities or stormwater runoff. Our results demonstrate that urbanization is changing the surrounding marine ecosystem and that mitigation approaches that include terrestrial management—such as restoring natural habitats along coastlines or improving wastewater treatment—may be more effective than approaches solely focused on marine regulations.
Urbanization Alters the Physicochemical Characteristics of Nearshore Ecosystems
Urbanization had a dramatic effect on salinity and nutrient conditions in Okinawa’s nearshore waters. Urban sites exhibited highly variable salinity, with salinities nearing zero on several dates. Dips in salinity were most pronounced at the central and northern urban subsites, which were more likely to to be influenced by the freshwater outflow. Subsite, and thus proximity to freshwater outflows, had less effect on salinity at rural sites. While watershed size—which influences freshwater input volume—was similar across sites, differences in geomorphology at urban and rural sites may influence residence times for inflowing freshwater and suspended sediments. The rural sites are semi-enclosed coral lagoons, leading to longer residence times for inflowing freshwater and suspended sediments (Sakamaki et al. 2022). In contrast, urban sites underwent substantial land reclamation that completely filled in the coral lagoons (Masucci and Reimer 2019), which should decrease the residence times for terrigenous inputs (Sakamaki et al. 2022). While the residence times for the sites in this study are undetermined, the projected geomorphology-driven variability in residence time is incongruent with our observations. Consequently, the difference in salinity between urban and rural sites is most likely due instead to the variation in impervious land cover (concretization) between sites. Urban watersheds in this study have scarce natural or permeable ground cover, which increases flooding and runoff to coastal areas (Blum et al. 2020). Wide variations in salinity—like those seen at the urban sites in this study—can have severe biological implications. Decreased salinity from runoff reduces coral holobiont respiration and photosynthetic rates, heightens coral bleaching and mortality risk, disrupts symbiotic relationships between the coral, microbiome, and zooxanthellae, and reduces coral larval and gamete survival (Röthig et al. 2023). Moreover, salinity is a major driver for bacterioplankton community composition (Jurdzinski et al. 2023; Lozupone and Knight 2007), and reduced salinity in nearshore regions can promote the growth of pathogenic bacteria (Bordalo et al. 2002; Burge et al. 2014; Randa et al. 2004) and increase bacterial pathogenicity (Barca et al. 2023).
The major macronutrients—nitrogen and phosphorus—were enriched throughout the annual cycle at urban sites compared to rural sites. Both urban and rural land uses contribute to nutrient loading with nutrient pollution deriving from sewage, fertilizers, detergents, and pet waste in urban areas (Hobbie et al. 2017) and from agricultural fertilizers, manure application, and livestock waste in rural areas (Del Rossi et al. 2023). However, nutrient export is positively correlated with impervious land cover, making nutrient pollution more likely to reach the coast in urban areas (Duan et al. 2012). Nitrogen and phosphorus were consistently elevated at urban sites compared to rural sites, but were also highly variable. Inorganic nutrient concentrations are temporally variable due to rapid uptake by microorganisms and short-term hydrodynamic processes (Viana and Bode 2013) but sampling across a time series allowed the overall elevation at urban sites to be detected. Nutrient loading increases primary production (Howarth et al. 2021) and chlorophyll a was significantly positively correlated with nitrogen and phosphorus at urban sites. Excess nutrient-induced primary production increases heterotrophic bacterial growth and respiration that can lead to hypoxia when combined with stratification (Obenour et al. 2012), and dissolved oxygen was significantly negatively correlated with nitrate and nitrite at urban sites. Nutrient loading can harm coral reefs by fueling the growth of fleshy macroalgae that compete with corals for light and space and increase bioerosion (Silbiger et al. 2018), and by amplifying the negative effects of ocean acidification and rising temperatures (DeCarlo et al. 2015). Together the altered physicochemical conditions associated with urbanization have potential to degrade the coral reef ecosystems that are important to Okinawa for fisheries and tourism.
Urbanization Changes Nearshore Bacterioplankton Community Composition
There were significantly different bacterial communities in nearshore waters adjacent to urban areas compared to those adjacent to rural areas, which is reflected in elevated bacterial diversity at urban sites. In addition to a shared core bacterial community of ubiquitous marine bacteria, urban sites hosted additional taxa that were rare or absent at rural sites. Bacteria in the orders Cloacimonadales, Clostridiales, Lactobacillales, Pseudomonadales, Saccharimonadales, Selenomonadales, Sphingobacteriales, and Thiomicrospirales were present in at least three urban samples but absent in all rural samples. Bacteria in the orders Aeromonadales, Alteromonadales, Bacteroidales, Betaproteobacteriales, Campylobacterales, Chitinophagales, Sphingomonadales, Thiotrichales, and Vibrionales were prevalent among urban samples but were rare in rural samples. The majority of these orders contain bacteria known to be associated with anthropogenic sources, including some that are considered fecal indicator bacteria (FIB) and some that are potentially pathogenic to humans and marine life (Verburg et al. 2021). In contrast, Verrucomicrobiales was the only order detected in multiple rural samples but absent in all urban samples. Bacteria in the order Verrucomicrobiales are ubiquitous in soils (Bergmann et al. 2011), which may explain their absence in urbanized regions where there is little natural soil exposed.
Among bacterial groups present in urban regions but absent in rural areas, there are several of particular concern due to the inclusion of fecal indicator bacteria or known pathogens. Specifically, Clostridiales and Pseudomonadales were prevalent at both urban sites and contain known human pathogens. At the genus level, most of the Clostridiales were classified as Blautia spp., Clostridium sensu stricto 1, and several other genera in the Ruminococcaceae family. Blautia spp. and Ruminococcaceae bacteria are both common components of microbiomes in humans and other mammals, contributing upwards of 50% of mammalian intestinal microbiome communities (Liu et al. 2021). Clostridium sensu stricto 1 includes the “true” Clostridium species, such as Clostridium perfringens that is a reliable indicator of human fecal contamination due to its specificity to sewage and ability to form environmentally stable cysts (Stelma 2018). While Pseudomonadales includes several known pathogens in the genera Pseudomonas and Acinetobacter, the Pseudomonadales ASVs abundant at urban sites were Pseudomonas hussainii and Acinetobacter indicus, which are not known human or animal pathogens. However, A. indicus was originally isolated for its chitinase activity, which may allow it to act as an invertebrate pathogen in reef systems (Akram et al. 2022). Cloacimonadales, Lactobacillales, and Thiomocrospirales are often associated with wastewater treatment plants, sewage sludge, and wastewater effluent (Shakeri Yekta et al. 2019; Verburg et al. 2021; Meyer et al. 2016). High abundances of these microbes in urban ecosystems may be due to their presence in wastewater reaching the coastal ecosystem.
Bacteria in orders that were prevalent in urban samples, but rare in rural samples, also included groups of concern (i.e., Vibrionales, Campylobacterales, Betaproteobacteriales, and Bacteroidales). The increased prevalence of Vibrionales at urban sites is alarming due to a recent surge in Vibrio spp. infections leading to human fatalities and limb amputations in developed coastal areas (Archer et al. 2023). Severe vibriosis cases are most often caused by V. vulnificans, but multiple Vibrio species can cause serious illness in humans (Baker-Austin et al. 2018) and marine organisms (Grimes 2020). Vibrio fortis and Vibrio chagasii were abundant in Ginowan (Urban Site 1), Photobacterium leiognathi subsp. mandapamensis was the major Vibrionales species found in Nago (Urban Site 2), and Photobacterium damselae subsp. damselae was detected at both sites. V. fortis, V. chagasii, and P. damselae are marine pathogens—V. fortis causes coral bleaching (Sun et al. 2023) and enteritis in fishes (Wang et al. 2016), V. chagasii infects molluscs (Urtubia et al. 2023), and P. damselae causes necrotizing fasciitis in humans and fish (Rivas et al. 2013)—while P. mandapamensis is not pathogenic (Urbanczyk et al. 2011). Urbanization-associated changes in physicochemical conditions could be responsible for increased abundance of Vibrionales at urban sites as several marine Vibrio species become more prevalent and infectious when salinity is reduced to 5–25 PSU (Sullivan and Neigel 2018) and salinity was often within this range at urban sites.
Campylobacterales, Betaproteobacteriales, and Bacteroidales were the most abundant groups prevalent among urban samples and rare in rural samples. Campylobacterales includes clinical pathogens (Man 2011) and environmental bacteria (Fera et al. 2004), and can be abundant in urban road-runoff samples (Liguori et al. 2021). Most Campylobacterales ASVs abundant at urban sites were classified as Arcobacter spp. Three of the five described Arcobacter species are associated with human disease and are found in human diarrhea and sewage samples, while the other two species are found in a variety of environmental settings—including rivers, salt marshes, and nearshore marine waters (Fera et al. 2004). Betaproteobacteriales is a large and diverse taxonomic group including many species found in wastewater treatment plants (Zhang et al. 2017). We detected 102 separate Betaproteobacteriales genera, but C39 (family: Rhodocyclaceae) was the most abundant and is associated with pollution and wastewater input in coastal ecosystems (Kopprio et al. 2020; Nascimento et al. 2018). Bacteroidales include dominant members of mammalian gut microbiomes (Magne et al. 2020) and sewage sludge communities (Shakeri Yekta et al. 2019), and have been found in road runoff (Liguori et al. 2021). The majority of Bacteroidales at urban sites belong to the genera Bacteroides and Prevotella, which are major mammalian gut bacteria and likely originate from sewage (Wu et al. 2011).
Overall, our results showcase a clear anthropogenic impact on microbial communities in nearshore environments adjacent to heavily urbanized watersheds in Okinawa. The urban ecosystems were consistently altered rather than exhibiting episodic or seasonal disturbances. Previous work focusing on the Tancha (R1) sampling site showed that extreme runoff events associated with typhoons caused short-lived perturbation of nearshore microbial communities, with the bacterial community composition returning to baseline only 48–72 h after storms passed (Ares et al. 2020). The quick return to pre-typhoon conditions highlights the quick-flushing of Okinawa’s coral lagoons, as well as the overall resiliency of nearshore ecosystems in more rural regions of Okinawa’s coast. In contrast, the consistently altered state observed at urban sites may indicate that these ecosystems are so altered that they have reached a tipping point and experienced a regime change. This regime change may interact with other components of the system (e.g., corals, algae, and other invertebrates) and create feedback loops that further degrade the ecosystem and its functioning (Becker et al. 2023; Qin et al. 2020). Alternatively, the consistently altered microbial communities observed at urban sites may reflect sustained, ongoing disturbances that would likely also have downstream ecological consequences (Ruprecht et al. 2021).
Conclusions and Future Directions
Biweekly observations made in nearshore waters adjacent to urban and rural watersheds revealed the profound influence of urbanization on salinity, macronutrient concentrations, and microbial communities in a recently urbanized subtropical island system. Expanding this work to include metatranscriptomics would allow for better characterization of the active metabolisms of microbes in urban and rural areas and could shed light on microbial nutrient uptake, as well as pathogenicity. Nonetheless, the observed changes in physicochemical parameters and microbial communities have likely contributed to degrading nearby coral reefs, as coral disease, death, and loss of diversity have been attributed to reduced salinity, nutrient loading, and introduced pathogenic bacteria in other systems. Coral reefs around Okinawa and other subtropical and tropical islands provide myriad ecosystem services, including supporting tourism and fisheries as well as protecting islands from waves and erosion, but the ability of reefs to perform these ecosystem services declines as reef health declines. To protect these key cultural, economic, and ecological resources, wholistic conservation measures that encompass both the reef ecosystem and the adjacent watershed are needed. Our results highlight the need for comprehensive conservation efforts that include land use management and coastal rehabilitation to ultimately protect important nearshore ecosystems, such as coral reefs. Moreover, more extensive wastewater treatment and moving outfalls further offshore can significantly reduce nutrient and pathogen loading on reefs. Such mitigation efforts may become more critical as cyclonic tropical storms increase in both frequency and intensity due to global climate change. Tropical cyclones deliver large amounts of precipitation that trigger extreme runoff events. Rural sites in Okinawa have proven to be resilient to extreme storms, but more study is needed to understand how urbanization influences ecosystem resiliency in the face of climate change.
Data Availability Statement
All raw sequencing reads generated for this project are publicly available from NCBI with accession number PRJNA1044524. Intermediate data files (including CTD and nutrient measurements) and the code necessary to replicate analyses are available in a GitHub repository (https://github.com/maggimars/UrbanOki) and as an interactive HTML document (https://maggimars.github.io/UrbanOki/Amplicons.html).
References
Akram, Fatima, Ikram ul Haq, Ayesha Roohi, and Rabia Akram. 2022. Acinetobacter indicus CCS-12: a new bacterial source for the production and biochemical characterization of thermostable chitinase with promising antifungal activity. Waste and Biomass Valorization 13 (7): 3371–3388.
Archer, Elizabeth J., Craig Baker-Austin, Timothy J. Osborn, Natalia R. Jones, Jaime Martínez-Urtaza, Joaquín Trinanes, James D. Oliver, Felipe J. Colón González, and Iain R. Lake. 2023. Climate warming and increasing Vibrio vulnificus infections in North America. Scientific Reports 13 (1): 3893.
Ares, Ángela., Margaret Mars Brisbin, Kirk N. Sato, Juan P. Martín, Yoshiteru Iinuma, and Satoshi Satoshi. 2020. Extreme storms cause rapid but short-lived shifts in nearshore subtropical bacterial communities. Environmental Microbiology 22 (11): 4571–4588.
Baker-Austin, Craig, James D. Oliver, Munirul Alam, Afsar Ali, Matthew K. Waldor, Firdausi Qadri, and Jaime Martinez-Urtaza. 2018. Vibrio spp. infections. Nature Reviews Disease Primers 4 (1): 8.
Barca, Alba V., Ana Vences, Mateus S. Terceti, Ana do Vale, and Carlos R. Osorio. 2023. Low salinity activates a virulence program in the generalist marine pathogen Photobacterium damselae subsp. damselae. Msystems 8 (3): e0125322.
Baum, Gunilla, Hedi I. Januar, Sebastian CA. Ferse, and Andreas Kunzmann. 2015. Local and regional impacts of pollution on coral reefs along the Thousand Islands north of the megacity Jakarta, Indonesia. PloS One 10 (9): e0138271.
Becker, Cynthia C., Laura Weber, Brian Zgliczynski, Chris Sullivan, Stuart Sandin, Erinn Muller, Abigail S. Clark, et al. 2023. Microorganisms and dissolved metabolites distinguish Florida’s coral reef habitats. PNAS Nexus 2 (9): gad287.
Bergmann, Gaddy T., Scott T. Bates, Kathryn G. Eilers, Christian L. Lauber, J Gregory Caporaso, William A. Walters, Rob Knight, and Noah Fierer. 2011. The under-recognized dominance of Verrucomicrobia in soil bacterial communities. Soil Biology & Biochemistry 43 (7): 1450–1455.
Bloom, David E. 2011. 7 billion and counting. Science 333(6042): 562–569.
Blum, Annalise G., Paul J. Ferraro, Stacey A. Archfield, and Karen R. Ryberg. 2020. Causal effect of impervious cover on annual flood magnitude for the United States. Geophysical Research Letters 47: e2019GL086480.
Bokulich, Nicholas A., Benjamin D. Kaehler, Jai Ram Rideout, Matthew Dillon, Evan Bolyen, Rob Knight, Gavin A. Huttley, and J Gregory Caporaso. 2018. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6 (1): 90.
Bolyen, Evan, Jai Ram Rideout, Matthew R. Dillon, Nicholas A. Bokulich, Christian C. Abnet, Gabriel A. Al-Ghalith, Harriet Alexander, et al. 2019. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nature Biotechnology. https://doi.org/10.1038/s41587-019-0209-9.
Bordalo, A.A., R. Onrassami, and C. Dechsakulwatana. 2002. Survival of faecal indicator bacteria in tropical estuarine waters (Bangpakong River, Thailand). Journal of Applied Microbiology 93 (5): 864–871.
Burge, Colleen A., Mark C. Eakin, Carolyn S. Friedman, Brett Froelich, Paul K. Hershberger, Eileen E. Hofmann, Laura E. Petes, et al. 2014. Climate change influences on marine infectious diseases: implications for management and society. Annual Review of Marine Science 6: 249–277.
Callahan, Benjamin J., Paul J. McMurdie, Michael J. Rosen, Andrew W. Han, Amy Jo A. Johnson, and Susan P. Holmes. 2016. DADA2: high-resolution sample inference from Illumina amplicon data. Nature Methods 13 (7): 581–583.
Cao, Wenting, Rui Li, Xiaoli Chi, Ninghua Chen, Jianyu Chen, and Huaguo Zhang. 2017. Island urbanization and its ecological consequences: a case study in the Zhoushan Island, East China. Ecological Indicators 76 (May): 1–14.
Carlson, Rachel R., Shawna A. Foo, and Gregory P. Asner. 2019. Land use impacts on coral reef health: a ridge-to-reef perspective. Frontiers in Marine Science 6: 00562.
DeCarlo, Thomas M., Anne L. Cohen, Hannah C. Barkley, Quinn Cobban, Charles Young, Kathryn E. Shamberger, Russell E. Brainard, and Yimnang Golbuu. 2015. Coral macrobioerosion is accelerated by ocean acidification and nutrients. Geology 43 (1): 7–10.
DeGeorges, Andre, Thomas J. Goreau, and Brian Reilly. 2010. Land-sourced pollution with an emphasis on domestic sewage: lessons from the Caribbean and implications for coastal development on Indian Ocean and Pacific coral reefs. Sustainability: Science Practice and Policy 2 (9): 2919–2949.
Del Rossi, Gemma, Mohammad Mainul Hoque, Y. Ji, and Catherine L. Kling. 2023. The economics of nutrient pollution from agriculture. Annual Review of Resource Economics. https://doi.org/10.1146/annurev-resource-111820-021317.
Duan, S., S.S. Kaushal, P.M. Groffman, L.E. Band, and K.T. Belt. 2012. Phosphorus export across an urban to rural gradient in the Chesapeake Bay watershed. JGR Biogeosciences 117 (G1): G01025 .
Fera, M.T., T.L. Maugeri, C. Gugliandolo, C. Beninati, M. Giannone, E. La Camera, and M. Carbone. 2004. Detection of Arcobacter spp in the coastal environment of the Mediterranean Sea. Applied and Environmental Microbiology 70 (3): 1271–1276.
Ghina, Fathimath. 2003. Sustainable development in small island developing states. Environment Development and Sustainability 5 (1): 139–165.
Gloor, Gregory B., Jean M. Macklaim, Vera Pawlowsky-Glahn, and Juan J. Egozcue. 2017. Microbiome datasets are compositional: and this is not optional. Frontiers in Microbiology 8 (November): 2224.
Grimes, DJay. 2020. The vibrios: scavengers, symbionts, and pathogens from the sea. Microbial Ecology 80 (3): 501–506.
Heery, Eliza C., Bert W. Hoeksema, Nicola K. Browne, James D. Reimer, Put O. Ang, Danwei Huang, Daniel A. Friess, et al. 2018. Urban coral reefs: degradation and resilience of hard coral assemblages in coastal cities of East and Southeast Asia. Marine Pollution Bulletin 135 (October): 654–681.
Hobbie, Sarah E., Jacques C. Finlay, Benjamin D. Janke, Daniel A. Nidzgorski, Dylan B. Millet, and Lawrence A. Baker. 2017. Contrasting nitrogen and phosphorus budgets in urban watersheds and implications for managing urban water pollution. Proceedings of the National Academy of Sciences of the United States of America 114 (16): 4177–4182.
Howarth, R.W., F. Chan, D.P. Swaney, R.M. Marino, and M. Hayn. 2021. Role of external inputs of nutrients to aquatic ecosystems in determining prevalence of nitrogen vs. phosphorus limitation of net primary productivity. Biogeochemistry 154 (2): 293–306.
Jurdzinski, Krzysztof T., Maliheh Mehrshad, Luis Fernando Delgado, Ziling Deng, Stefan Bertilsson, and Anders F. Andersson. 2023. Large-scale phylogenomics of aquatic bacteria reveal molecular mechanisms for adaptation to salinity. Science Advances 9 (21): eadg2059.
Kier, Gerold, Holger Kreft, Tien Ming Lee, Walter Jetz, Pierre L. Ibisch, Christoph Nowicki, Jens Mutke, and Wilhelm Barthlott. 2009. A global assessment of endemism and species richness across island and mainland regions. Proceedings of the National Academy of Sciences of the United States of America 106 (23): 9322–9327.
Klindworth, Anna, Elmar Pruesse, Timmy Schweer, Jörg. Peplies, Christian Quast, Matthias Horn, and Frank Oliver Glöckner. 2013. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Research 41 (1): e1.
Kobayashi, Masanori. 2022. The COVID-19 impacts and challenges to achieving sustainability in Japan’s fisheries and aquaculture. Marine Policy 143 (September): 105161.
Kopprio, Germán A., Sucharit B. Neogi, Harunur Rashid, Cecilia Alonso, Shinji Yamasaki, Boris P. Koch, Astrid Gärdes, and Rubén J. Lara. 2020. Vibrio and bacterial communities across a pollution gradient in the Bay of Bengal: unraveling their biogeochemical drivers. Frontiers in Microbiology 11 (April): 594.
Kurtz, Zachary D., Christian L. Müller, Emily R. Miraldi, Dan R. Littman, Martin J. Blaser, and Richard A. Bonneau. 2015. Sparse and compositionally robust inference of microbial ecological networks. PLoS Computational Biology 11 (5): e1004226.
Lee, S.Y., R.J.K. Dunn, R.A. Young, R.M. Connolly, P.E.R. Dale, R. Dehayr, C.J. Lemckert, et al. 2006. Impact of urbanization on coastal wetland structure and function. Austral Ecology 31 (2): 149–163.
Li, Yinji, Xiaobo Lou, and Sachiko Harada. 2018. Social responses to a fishery-tourism conflict in Onna Village, Okinawa, Japan. In Global change in marine systems: societal and governing responses, vol. 19, ed. Patrice Guillotreau, Alida Bundy, and Ian R. Perry. Routledge.
Liguori, Renato, Steffen H. Rommel, Johan Bengtsson-Palme, Brigitte Helmreich, and Christian Wurzbacher. 2021. Microbial retention and resistances in stormwater quality improvement devices treating road runoff. bioRxiv. https://doi.org/10.1101/2021.01.12.426166.
Liu, Xuemei, Bingyong Mao, Jiayu Gu, Jiaying Wu, Shumao Cui, Gang Wang, Jianxin Zhao, Hao Zhang, and Wei Chen. 2021. Blautia—a new functional genus with potential probiotic properties? Gut Microbes 13 (1): 1–21.
Lozupone, Catherine A., and Rob Knight. 2007. Global patterns in bacterial diversity. Proceedings of the National Academy of Sciences of the United States of America 104 (27): 11436–11440.
Magne, Fabien, Martin Gotteland, Lea Gauthier, Alejandra Zazueta, Susana Pesoa, Paola Navarrete, and Ramadass Balamurugan. 2020. The firmicutes/bacteroidetes ratio: a relevant marker of gut dysbiosis in obese patients? Nutrients 12 (5): 1474.
Man, Si Ming. 2011. The clinical importance of emerging Campylobacter species. Nature Reviews Gastroenterology & Hepatology 8(12): 669–685.
Masucci, Giovanni Diego, and James D. Reimer. 2019. Expanding walls and shrinking beaches: loss of natural coastline in Okinawa Island, Japan. PeerJ 7 (September): e7520.
McLellan, Sandra L., Jenny C. Fisher, and Ryan J. Newton. 2015. The microbiome of urban waters. International Microbiology: The Official Journal of the Spanish Society for Microbiology 18 (3): 141–149.
Mcmurdie, Paul J and Susan Holmes. 2013. Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLOS ONE 8 (4): e61217.
Meyer, Daniel Derrossi, Pedro Avelino Maia. de Andrade, Ademir Durrer, Fernando Dini Andreote, Gertrudes Corção, and Adriano Brandelli. 2016. Bacterial communities involved in sulfur transformations in wastewater treatment plants. Applied Microbiology and Biotechnology 100 (23): 10125–35.
Nascimento, Altina Lacerda, Adijailton Jose Lacerda, Pedro Avelino Maia. Andrade, Fernando Dini Andreote, Aline Renée Coscione, Fernando Carvalho Oliveira, and Jussara Borges Regitano. 2018. Sewage sludge microbial structures and relations to their sources, treatments, and chemical attributes. Frontiers in Microbiology 9 (July): 1462.
Nogales, Balbina, Mariana P. Lanfranconi, Juana M. Piña-Villalonga, and Rafael Bosch. 2011. Anthropogenic perturbations in marine microbial communities. FEMS Microbiology Reviews 35 (2): 275–298.
Obenour, Daniel R., Anna M. Michalak, Yuntao Zhou, and Donald Scavia. 2012. Quantifying the impacts of stratification and nutrient loading on hypoxia in the Northern Gulf of Mexico. Environmental Science & Technology 46 (10): 5489–5496.
Oksanen, Jari, F. Guillaume, Michael Blanchet, Roeland Friendly, Pierre Kindt, Dan Legendre, Peter R. McGlinn, and Minchin et al. 2019. Vegan: community ecology package. R Package Version 2.5-4. https://CRAN.R-project.org/package=vegan.
Qin, Zhenjun, Kefu Yu, Jiayuan Liang, Qiucui Yao, and Biao Chen. 2020. Significant changes in microbial communities associated with reef corals in the southern South China sea during the 2015/2016 global-scale coral bleaching event. Journal of Geophysical Research: Oceans 125: e2019JC015579.
Quast, Christian, Elmar Pruesse, Pelin Yilmaz, Jan Gerken, Timmy Schweer, Frank Oliver Glo, Pablo Yarza, Jörg Peplies, and Frank Oliver Glöckner. 2013. The SILVA ribosomal RNA gene database project : improved data processing and web-based tools. Nucleic Acids Research 41 (D1): 590–96.
R Core Team. 2018. R: a language and environment for statistical computing. https://www.R-project.org/.
Randa, Mark A., Martin F. Polz, and Eelin Lim. 2004. Effects of temperature and salinity on Vibrio vulnificus population dynamics as assessed by quantitative PCR. Applied and Environmental Microbiology 70 (9): 5469–5476.
Rintoul, M.S., T.A. Courtney, J.L. Dohner, S.N. Giddings, S.A.H. Kekuewa, S. Mitarai, S.G. Monismith, A.K. Pezner, and A.J. Andersson. 2022. The effects of light intensity and flow speed on biogeochemical variability within a fringing coral reef in Onna-son, Okinawa, Japan. Journal of Geophysical Research: Oceans 127: e2021JC018369.
Rivas, Amable J., Manuel L. Lemos, and Carlos R. Osorio. 2013. Photobacterium damselae subsp. damselae, a bacterium pathogenic for marine animals and humans. Frontiers in Microbiology 4 (September): 283.
Roberts, Michaela, Nick Hanley, Sam Williams, and Will Cresswell. 2017. Terrestrial degradation impacts on coral reef health: evidence from the Caribbean. Ocean & Coastal Management 149 (November): 52–68.
Ross, Samuel RP-J., Nicholas R. Friedman, Kenneth L. Dudley, Masashi Yoshimura, Takuma Yoshida, and Evan P. Economo. 2018. Listening to ecosystems: data-rich acoustic monitoring through landscape‐scale sensor networks. Ecological Research 33 (1): 135–147.
Röthig, Till, Stacey M. Trevathan-Tackett, Christian R. Voolstra, Cliff Ross, Samuel Chaffron, Paul J. Durack, Laura M. Warmuth, and Michael Sweet. 2023. Human-Induced Salinity Changes Impact Marine organisms and ecosystems. Global Change Biology 29 (17): 4731–4749.
Ruprecht, J.E., S.C. Birrer, K.A. Dafforn, S.M. Mitrovic, S.L. Crane, E.L. Johnston, F. Wemheuer, et al. 2021. Wastewater effluents cause microbial community shifts and change trophic status. Water Research 200 (July): 117206.
Russell, James C., and Christoph Kueffer. 2019. Island biodiversity in the anthropocene. Annual Review of Environment and Resources 44 (1): 31–60.
Sakamaki, Takashi, Akiko Morita, Shouji Touyama, Yasushi Watanabe, and Shouhei Suzuki. 2022. Effects of watershed land use on coastal marine environments: a multiscale exploratory analysis with multiple biogeochemical indicators in fringing coral reefs of Okinawa Island. Marine Pollution Bulletin 183 (October): 114054.
Seto, Karen C., Michail Fragkias, Burak Güneralp, and Michael K. Reilly. 2011. A meta-analysis of global urban land expansion. PloS One 6 (8): e23777.
Shakeri Yekta, Sepehr, Tong Liu, Mette Axelsson Bjerg, Luka Šafarič, Anna Karlsson, Annika Björn, and Anna Schnürer. 2019. Sulfide level in municipal sludge digesters affects microbial community response to long-chain fatty acid loads. Biotechnology for Biofuels 12 (November): 259.
Silbiger, Nyssa J., Craig E. Nelson, Kristina Remple, Jessica K. Sevilla, Zachary A. Quinlan, Hollie M. Putnam, Michael D. Fox, and Megan J. Donahue. 2018. Nutrient pollution disrupts key ecosystem functions on coral reefs. Proceedings Biological Sciences 285 (1880): 20172718.
Singh, Tanya, Frederic Sinniger, Yoshikatsu Nakano, Shigeo Nakamura, Shouhei Kadena, Mori Jinza, and Hiroyuki Fujimura. 2022. Long-term trends and seasonal variations in environmental conditions in Sesoko Island, Okinawa, Japan. Galaxea Journal of Coral Reef Studies 24 (1): 121–133.
Small, Christopher, and Robert J. Nicholls. 2003. A global analysis of human settlement in coastal zones. Journal of Coastal Research 19 (3): 584–599.
Steibl, Sebastian, Jonas Franke, and Christian Laforsch. 2021. Tourism and urban development as drivers for invertebrate diversity loss on tropical islands. Royal Society Open Science 8 (10): 210411.
Stelma, Gerard N. 2018. Use of bacterial spores in monitoring water quality and treatment. Journal of Water and Health 16 (4): 491–500.
Sullivan, Timothy J., and Joseph E. Neigel. 2018. Effects of temperature and salinity on prevalence and intensity of infection of blue crabs, Callinectes sapidus, by Vibrio cholerae, V. parahaemolyticus, and V. vulnificus in Louisiana. Journal of Invertebrate Pathology 151 (January): 82–90.
Sun, Xiaohui, Yan Li, Qian Yang, Han Zhang, Nuo Xu, Zheng Tang, Shishi Wu, et al. 2023. Identification of quorum sensing-regulated Vibrio fortis as potential pathogenic bacteria for coral bleaching and the effects on the microbial shift. Frontiers in Microbiology 14 (February): 1116737.
Sutherland, Kathryn Patterson, Sameera Shaban, Jessica L. Joyner, James W. Porter, and Erin K. Lipp. 2011. Human pathogen shown to cause disease in the threatened eklhorn coral Acropora palmata. PloS One 6 (8): e23468.
Todd, Peter A., Eliza C. Heery, Lynette HL. Loke, Ruth H. Thurstan, D Johan Kotze, and Christopher Swan. 2019. Towards an urban marine ecology: characterizing the drivers, patterns and processes of marine ecosystems in coastal cities. Oikos 128 (9): 1215–1242.
Tuholske, Cascade, Zachary Tane, David López-Carr, Dar Roberts, and Susan Cassels. 2017. Thirty years of land use/cover change in the Caribbean: assessing the relationship between urbanization and mangrove loss in Roatán, Honduras. Applied Geography 88 (November): 84–93.
Uehara, Masato, Akihiko Ebisawa, Itaru Ohta, and Yoshimasa Aonuma. 2019. Effectiveness of deepwater marine protected areas: implication for Okinawan demersal fisheries management. Fisheries Research 215 (July): 123–130.
Urbanczyk, Henryk, Jennifer C. Ast, and Paul V. Dunlap. 2011. Phylogeny, genomics, and symbiosis of photobacterium. FEMS Microbiology Reviews 35 (2): 324–342.
Urtubia, Rocío, Claudio D. Miranda, Sergio Rodríguez, Javier Dubert, Juan L. Barja, and Rodrigo Rojas. 2023. First report, characterization and pathogenicity of Vibrio chagasii isolated from diseased reared larvae of Chilean scallop, Argopecten purpuratus (Lamarck, 1819). Pathogens 12 (2): 183.
Verburg, Ilse, H. Pieter J. van Veelen, Karola Waar, John WA. Rossen, Alex W. Friedrich, Lucia Hernández Leal, Silvia García-Cobos, and Heike Schmitt. 2021. Effects of clinical wastewater on the bacterial community structure from sewage to the environment. Microorganisms 9 (4): 718.
Viana, Inés. G., and Antonio Bode. 2013. Stable nitrogen isotopes in coastal macroalgae: geographic and anthropogenic variability. The Science of the Total Environment 443 (January): 887–895.
Wang, X., Y. Zhang, G. Qin, W. Luo, and Q. Lin. 2016. A novel pathogenic bacteria (Vibrio fortis) causing enteritis in cultured seahorses, Hippocampus Erectus Perry, 1810. Journal of Fish Diseases 39 (6): 765–769.
Willis, Amy, John Bunge, and Thea Whitman. 2017. Improved detection of changes in species richness in high diversity microbial communities. Journal of the Royal Statistical Society: Series C (Applied Statistics) 66 (5): 963–977.
Wu, Gary D., Jun Chen, Christian Hoffmann, Kyle Bittinger, Ying-Yu. Chen, Sue A. Keilbaugh, Meenakshi Bewtra, et al. 2011. Linking long-term dietary patterns with gut microbial enterotypes. Science 334 (6052): 105–108.
Zhang, Bo., Xiangyang Xu, and Liang Zhu. 2017. Structure and function of the microbial consortia of activated sludge in typical municipal wastewater treatment plants in winter. Scientific Reports 7 (1): 17930.
Acknowledgements
We acknowledge the Incorporated Foundation Okinawa Environment Science Centre for collecting samples, and Ayse Oshima and Pradeep Palanichamy for assisting in filtering water samples and extracting DNA. The Okinawa Institute of Science and Technology (OIST) Sequencing Center (SQC) performed sequencing library preparation and sequencing. Research was funded by the OIST Marine Biophysics Unit and a Japan Society for the Promotion for Science (JSPS) Kakenhi award to AA (award 20K19986). MMB was supported by a Simons Foundation Postdoctoral Fellowship in Marine Microbiology (award 874439).
Funding
Open access funding provided by the Okinawa Institute of Science and Technology Graduate University (OIST). Research was funded by the OIST Marine Biophysics Unit and a Japan Society for the Promotion for Science (JSPS) Kakenhi award to AA (award 20K19986). MMB was supported by a Simons Foundation Postdoctoral Fellowship in Marine Microbiology (award 874439).
Author information
Authors and Affiliations
Contributions
AA conceived of the project. AA, SM, and MMB planned the research. AA and YY performed the research. AA, MMB, and KD analyzed the data. MMB wrote the manuscript. All authors edited drafts and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Bongkeun Song
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mars Brisbin, M., Dudley, K.L., Yonashiro, Y. et al. Urbanization of a Subtropical Island (Okinawa, Japan) Alters Physicochemical Characteristics and Disrupts Microbial Community Dynamics in Nearshore Ecosystems. Estuaries and Coasts 47, 1266–1281 (2024). https://doi.org/10.1007/s12237-024-01366-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12237-024-01366-3