Abstract
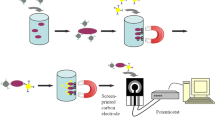
The application of rapid, specific, and sensitive methods for pathogen detection and quantification is very advantageous in diagnosis of human pathogens in several applications, including food analysis. The aim of this study was the evaluation of a method for the multiplexed detection and quantification of three significant foodborne pathogenic species (Escherichia coli O157, Salmonella spp., and Listeria monocytogenes). The assay combines specific DNA extraction by multiplex magnetic capture hybridization (mMCH) with multiplex real-time PCR. The amplification assay showed linearity in the range 106–10 genomic units (GU)/PCR for each co-amplified species. The sensitivity corresponded to 1 GU/PCR for E. coli O157 and L. monocytogenes, and 10 GU/PCR for Salmonella spp. The immobilization process and the hybrid capture of the MCH showed good efficiency and reproducibility for all targets, allowing the combination in equal amounts of the different nanoparticle types in mMCH. MCH and mMCH efficiencies were similar. The detection limit of the method was 10 CFU in samples with individual pathogens and 102 CFU in samples with combination of the three pathogens in unequal amounts (amount’s differences of 2 or 3 log). In conclusion, this multiplex molecular platform can be applied to determine the presence of target species in food samples after culture enrichment. In this way, this method could be a time-saving and sensitive tool to be used in routine diagnosis.



Similar content being viewed by others
References
Algaba IG, Geerts M, Jennes M, Coucke W, Opsteegh M, Cox E, Dorny P, Dierick K, De Craeye S (2017) A more sensitive, efficient and ISO 17025 validated magnetic capture real time PCR method for the detection of the archetypal Toxoplasma gondii strains in meat. Int J Parasitol doi:https://doi.org/10.1016/j.ijpara.2017.05.005
Amagliani G, Omiccioli E, Campo A, Bruce IJ, Brandi G, Magnani M (2006) Development of a magnetic capture hybridization-PCR assay for Listeria monocytogenes direct detection in milk samples. J Appl Microbiol 100:375–383. https://doi.org/10.1111/j.1365-2672.2005.02761.x
Amagliani G, Omiccioli E, Brandi G, Bruce IJ, Magnani M (2010) A multiplex magnetic capture hybridisation and multiplex real-time PCR protocol for pathogen detection in seafood. Food Microbiol 27:580–585. https://doi.org/10.1016/j.fm.2010.01.007
Amagliani G, Petruzzelli A, Carloni E, Tonucci F, Foglini M, Micci E, Ricci M, Di Lullo S, Rotundo L, Brandi G (2016) Presence of Escherichia coli O157, Salmonella spp., and Listeria monocytogenes in raw ovine milk destined for cheese production and evaluation of the equivalence between the analytical methods applied. Foodborne Pathog Dis 13:626–632. https://doi.org/10.1089/fpd.2016.2159
Brusa V, Galli L, Linares LH, Ortega EE, Lirón JP, Leotta GA (2015) Development and validation of two SYBR green PCR assays and a multiplex real-time PCR for the detection of Shiga toxin-producing Escherichia coli in meat. J Microbiol Methods 119:10–17. https://doi.org/10.1016/j.mimet.2015.09.013
Bustin SA (2000) Absolute quantification of mRNA using real-time reverse transcription polymerase chain reaction assays. J Mol Endocrinol 25:169–193
Carloni E, Amagliani G, Omiccioli E, Ceppetelli V, Del Mastro M, Rotundo L, Brandi G, Magnani M (2017) Validation and application of a quantitative real-time PCR assay to detect common wheat adulteration of durum wheat for pasta production. Food Chem 224:86–91. https://doi.org/10.1016/j.foodchem.2016.12.053
Cocolin L, Rantsiou K (2012) Quantitative polymerase chain reaction in food microbiology. In: Filion M (ed) Quantitative real-time PCR in applied microbiology. Department of Biology, Universitè de Moncton, Moncton, NB, Canada, pp 149–160
Cocolin L, Rajkovic A, Rantsiou K, Uyttendaele M (2011) The challenge of merging food safety diagnostic needs with quantitative PCR platforms. Trends Food Sci Technol 22:S30–S38. https://doi.org/10.1016/j.tifs.2011.02.009.
D’Urso OF, Poltronieri P, Marsigliante S, Storelli C, Hernández M, Rodríguez-Lázaro D (2009) A filtration-based real-time PCR method for the quantitative detection of viable Salmonella enterica and Listeria monocytogenes in food samples. Food Microbiol 26:311–316. https://doi.org/10.1016/j.fm.2008.12.006
EU (2005) Commission Regulation (EC) No 2073/2005 of 15 November 2005 on microbiological criteria for foodstuffs. Available at: http://eur-lex.europa.eu/legal-content/EN/ALL/?uri=CELEX%3A32005R2073
Forghani F, Wei S, Oh D-H (2016) A rapid multiplex real-time PCR high-resolution melt curve assay for the simultaneous detection of Bacillus cereus, Listeria monocytogenes, and Staphylococcus aureus in food. J Food Prot 79:810–815. https://doi.org/10.4315/0362-028X.JFP-15-428
Garrido A, Chapela M-J, Román B, Fajardo P, Vieites JM, Cabado AG (2013) In-house validation of a multiplex real-time PCR method for simultaneous detection of Salmonella spp., Escherichia coli O157 and Listeria monocytogenes. Int J Food Microbiol 164:92–98. https://doi.org/10.1016/j.ijfoodmicro.2013.03.024
Guo Y, Sheng S, Nie B, Tu Z (2015) Development of magnetic capture hybridization and quantitative polymerase chain reaction for hepatitis B virus covalently closed circular DNA. Hepat Mon 15:e23729. https://doi.org/10.5812/hepatmon.23729
Harada T, Iguchi A, Iyoda S, Seto K, Taguchi M, Kumeda Y (2015) Multiplex real-time PCR assays for screening of Shiga toxin 1 and 2 genes, including all known subtypes, and Escherichia coli O26-, O111-, and O157-specific genes in beef and sprout enrichment cultures. J Food Prot 78:1800–1811. https://doi.org/10.4315/0362-028X.JFP-15-050
Johnson KL, Zheng D, Kaewnum S, Reid CL, Burr T (2013) Development of a magnetic capture hybridization real-time PCR assay for detection of tumorigenic Agrobacterium vitis in grapevines. Phytopathology 103:633–640. https://doi.org/10.1094/PHYTO-10-12-0267-R
Kawasaki S, Fratamico PM, Horikoshi N, Okada Y, Takeshita K, Sameshima T, Kawamoto S (2010) Multiplex real-time polymerase chain reaction assay for simultaneous detection and quantification of Salmonella species, Listeria monocytogenes, and Escherichia coli O157:H7 in ground pork samples. Foodborne Pathog Dis 7:549–554. https://doi.org/10.1089/fpd.2009.0465
Löfström C, Schelin J, Norling B, Vigre H, Hoorfar J, Rådström P (2011) Culture-independent quantification of Salmonella enterica in carcass gauze swabs by flotation prior to real-time PCR. Int J Food Microbiol 145:S103–S109. https://doi.org/10.1016/j.ijfoodmicro.2010.03.042.
Nadkarni MA, Martin FE, Jacques NA, Hunter N (2002) Determination of bacterial load by real-time PCR using a broad-range (universal) probe and primers set. Microbiol 148:257–266. https://doi.org/10.1099/00221287-148-1-257
Nordstrom JL, Vickery MCL, Blackstone GM, Murray SL, DePaola A (2007) Development of a multiplex real-time PCR assay with an internal amplification control for the detection of total and pathogenic Vibrio parahaemolyticus bacteria in oysters. Appl Environ Microbiol 73:5840–5847. https://doi.org/10.1128/AEM.00460-07
Omiccioli E, Amagliani G, Brandi G, Magnani M (2009) A new platform for real-time PCR detection of Salmonella spp., Listeria monocytogenes and Escherichia coli O157 in milk. Food Microbiol 26:615–622. https://doi.org/10.1016/j.fm.2009.04.008
Rantsiou K, Alessandria V, Urso R, Dolci P, Cocolin L (2008) Detection, quantification and vitality of Listeria monocytogenes in food as determined by quantitative PCR. Int J Food Microbiol 121:99–105. https://doi.org/10.1016/j.ijfoodmicro.2007.11.006
Reischl U, Kochanowski B (1995) Quantitative PCR. A survey of the present technology. Mol Biotechnol 3:55–71. https://doi.org/10.1007/BF02821335.
Reiter M, Kirchner B, Müller H, Holzhauer C, Mann W, Pfaffl MW (2011) Quantification noise in single cell experiments. Nucleic Acids Res 39:e124. https://doi.org/10.1093/nar/gkr505
Rodríguez-Lázaro D, Jofré A, Aymerich T, Garriga M, Pla M (2005) Rapid quantitative detection of Listeria monocytogenes in salmon products: evaluation of pre-real-time PCR strategies. J Food Prot 68:1467–1471
Rowan NJ (2004) Viable but non-culturable forms of food and waterborne bacteria: quo vadis? Trends Food Sci Technol 15:462–467. https://doi.org/10.1016/j.tifs.2004.02.009
Singh J, Batish VK, Grover S (2009) A molecular beacon-based duplex real-time polymerase chain reaction assay for simultaneous detection of Escherichia coli O157:H7 and Listeria monocytogenes in milk and milk products. Foodborne Pathog Dis 6:1195–1201. https://doi.org/10.1089/fpd.2009.0310
Suo B, He Y, Tu S-I, Shi X (2010) A multiplex real-time polymerase chain reaction for simultaneous detection of Salmonella spp., Escherichia coli O157, and Listeria monocytogenes in meat products. Foodborne Pathog Dis 7:619–628. https://doi.org/10.1089/fpd.2009.0430
Wang L, Li Y, Mustaphai A (2007) Rapid and simultaneous quantitation of Escherichia coli 0157:H7, Salmonella, and Shigella in ground beef by multiplex real-time PCR and immunomagnetic separation. J Food Pro 70:1366–1372
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(DOCX 16.7 kb)
Rights and permissions
About this article
Cite this article
Carloni, E., Rotundo, L., Brandi, G. et al. Rapid and simultaneous detection of Salmonella spp., Escherichia coli O157, and Listeria monocytogenes by magnetic capture hybridization and multiplex real-time PCR. Folia Microbiol 63, 735–742 (2018). https://doi.org/10.1007/s12223-018-0617-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-018-0617-0