Abstract
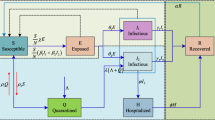
The outbreak of coronavirus disease 2019 (COVID-19) has aroused a global alert. To release social panic and guide future schedules, this article proposes a novel mathematical model, the Delay Differential Epidemic Analyzer (D2EA), to analyze the dynamics of epidemic and forecast its future trends. Based on the traditional Susceptible-Exposed-Infectious-Recovered (SEIR) model, the D2EA model innovatively introduces a set of quarantine states and applies both ordinary differential equations and delay differential equations to describe the transition between two states. Potential variations of practical factors are further considered to reveal the true epidemic picture. In the experiment part, we use the D2EA model to simulate the epidemic in Hubei Province. Fitting to the collected real data as non-linear optimization, the D2EA model forecasts that the accumulated confirmed infected cases in Hubei Province will reach the peak at the end of February and then steady down. We also evaluate the effectiveness of the quarantine measures and schedule the date to reopen Hubei Province.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- D(t):
-
Deaths of disease, person
- E(t):
-
Free exposed population (also infectious), person
- I(t):
-
Free infectious population, person
- N(t):
-
Total of free populations, person
- QE(t):
-
Quarantined exposed population, person
- QI(t):
-
Quarantined confirmed infected cases, person
- QN(t):
-
Quarantined suspected but negative cases, person
- QP(t):
-
Quarantined suspected and positive cases, person
- QS(t):
-
Quarantined healthy susceptible population, person
- r(t):
-
Number of contact with one person per unit time, person/d
- R(t):
-
Recovered population (also free), person
- S(t):
-
Free susceptible population, person
- t :
-
Time, d
- T :
-
Time needed to confirm one suspected case, d
- α :
-
Death rate of QI (t), d−1
- α I :
-
Death rate of I(t), d−1
- β E :
-
Probability of transmission per contact with E(t)
- β I :
-
Probability of transmission per contact with I(t)
- γ(t):
-
Recovery rate of QI (t), d−1
- γ E :
-
Recovery rate of E(t), d−1
- γ I :
-
Recovery rate of I(t), d−1
- δ(t):
-
Quarantine rate of I(t), d−1
- ε :
-
Rate of progression from exposed state to infectious state, d−1
- η :
-
Ratio of common cold sufferers to all susceptibles
- τ :
-
Quarantine duration, d
References
CDC. 2019 novel coronavirus symptoms [EB/OL]. (2019-01-31) [2020-02-28]. https://www.cdc.gov/coronavirus/2019-ncov/about/symptoms.html.
CHAN J F W, YUAN S F, KOK K H, et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster [J]. The Lancet, 2020, 395(10223): 514–523.
WHO. Coronavirus disease 2019 (COVID-19): Situation report-24 [EB/OL]. (2020-02-13) [2020-02-28]. https://www.who.int/emergencies/diseases/novelcoronavirus-2019/situation-reports.
ROTHE C, SCHUNK M, SOTHMANN P, et al. Transmission of 2019-ncov infection from an asymptomatic contact in Germany [J]. The New England Journal of Medicine, 2020, 382: 970–971.
LI Q, GUAN X H, WU P, et al. Early transmission dynamics in Wuhan, China, of novel coronavirus-infected pneumonia [J]. The New England Journal of Medicine, 2020. https://doi.org/10.1056/NEJMoa2001316 (published online).
WHO. Pneumonia of unknown cause-China [EB/OL]. (2020-01-05) [2020-02-28]. https://www.who.int/csr/don/05-january-2020-pneumonia-of-unkown-causechina/en/.
DONG N, YANG X, YE L, et al. Genomic and protein structure modelling analysis depicts the origin and infectivity of 2019-nCoV, a new coronavirus which caused a pneumonia outbreak in Wuhan, China [EB/OL]. (2020-01-22) [2020-02-28]. https://doi.org/10.1101/2020.01.20.913368.
LETKO M, MARZI A, MUNSTER V. Functional assessment of cell entry and receptor usage for SARS-CoV-2 and other lineage B betacoronaviruses [J]. Nature Microbiology, 2020. https://doi.org/10.1038/s41564-020-0688-y (published online).
CORMAN V M, LANDT O, KAISER M, et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR [J]. Eurosurveillance, 2020, 25(3): 2000045.
BEAL J, MITCHELL T, WYSCHOGROD D, et al. Highly distinguished amino acid sequences of 2019-nCoV (Wuhan coronavirus) [EB/OL]. (2020-02-02) [2020-02-28]. https://doi.org/10.1101/2020.01.31.929497.
JU J, KUMAR S, LI X, et al. Nucleotide analogues as inhibitors of viral polymerases [EB/OL]. (2020-01-31) [2020-02-28]. https://doi.org/10.1101/2020.01.30.927574.
GUO Q, LI M, WANG C, et al. Host and infectivity prediction of Wuhan 2019 novel coronavirus using deep learning algorithm [EB/OL]. (2020-02-02) [2020-02-28]. https://doi.org/10.1101/2020.01.21.914044.
ZHOU P, YANG X L, WANG X G, et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin [J]. Nature, 2020, 579: 270–273.
LU H. Drug treatment options for the 2019-new coronavirus (2019-nCoV) [J]. BioScience Trends, 2020, 14(1): 69–71.
RAMAIAH A, ARUMUGASWAMI V. Insights into cross-species evolution of novel human coronavirus 2019-nCoV and defining immune determinants for vaccine development [EB/OL]. (2020-02-04) [2020-02-28]. https://doi.org/10.1101/2020.01.29.925867.
QUILTY B, CLIFFORD S, FLASCHE S, et al. Effectiveness of airport screening at detecting travellers infected with 2019-nCoV [EB/OL]. (2020-02-02) [2020-02-28]. https://doi.org/10.1101/2020.01.31.20019265.
ZHAO S, LIN Q, RAN J, et al. Preliminary estimation of the basic reproduction number of novel coronavirus (2019-nCoV) in China, from 2019 to 2020: A datadriven analysis in the early phase of the outbreak [J]. International Journal of Infectious Diseases, 2020, 92: 214–217.
WU J T, LEUNG K, LEUNG G M. Nowcasting and forecasting the potential domestic and international spread of the 2019-nCoV outbreak originating in Wuhan, China: A modelling study [J]. The Lancet, 2020, 395(10225): 689–697.
RIOU J, ALTHAUS C L. Pattern of early human-tohuman transmission of Wuhan 2019 novel coronavirus (2019-nCoV), December 2019 to January 2020 [J]. Eurosurveillance, 2020, 25(4): 2000058.
READ J M, BRIDGEN J R E, CUMMINGS D A T, et al. Novel coronavirus 2019-nCoV: Early estimation of epidemiological parameters and epidemic predictions [EB/OL]. (2020-01-28) [2020-02-28]. https://doi.org/10.1101/2020.01.23.20018549.
MA Z E, ZHOU Y C, WANG W D, et al. Mathematical modeling and research of infectious disease dynamics [M]. Beijing: Science Press, 2004 (in Chinese).
DIEKMANN O, HEESTERBEEK H, BRITTON T. Mathematical tools for understanding infectious disease dynamics [M]. Princeton: Princeton University Press, 2012.
HETHCOTE H W. The mathematics of infectious diseases [J]. SIAM Review, 2000, 42(4): 599–653.
KEELING M J, ROHANI P. Modeling infectious diseases in humans and animals [M]. Princeton: Princeton University Press, 2008.
FUNK S, CIGLENECKI I, TIFFANY A, et al. The impact of control strategies and behavioural changes on the elimination of Ebola from Lofa County, Liberia [J]. Philosophical Transactions of the Royal Society B: Biological Sciences, 2017, 372(1721): 20160302.
WANG W D, RUAN S G. Simulating the SARS outbreak in Beijing with limited data [J]. Journal of Theoretical Biology, 2004, 227(3): 369–379.
GONZÁLEZ-PARRA G, ARENAS A J, ARANDA D F, et al. Modeling the epidemic waves of AH1N1/09 influenza around the world [J]. Spatial and Spatiotemporal Epidemiology, 2011, 2(4): 219–226.
Author information
Authors and Affiliations
Corresponding author
Additional information
Foundation item: the National Key Research and Development Program of China (No. 2018YFB1004700), the National Natural Science Foundation of China (Nos. 61872238 and 61972254), the Shanghai Science and Technology Fund (No. 17510740200), and the CCFHuawei Database System Innovation Research Plan (No. CCF-Huawei DBIR2019002A)
Rights and permissions
About this article
Cite this article
Liu, C., Zhao, J., Liu, G. et al. D2EA: Depict the Epidemic Picture of COVID-19. J. Shanghai Jiaotong Univ. (Sci.) 25, 165–176 (2020). https://doi.org/10.1007/s12204-020-2170-7
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12204-020-2170-7
Key words
- coronavirus disease 2019 (COVID-19)
- epidemic model
- quarantine states
- Susceptible-Exposed-Infectious-Recovered (SEIR)
- delay differential equation
- non-linear optimization