Abstract
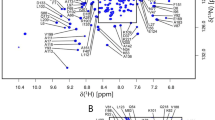
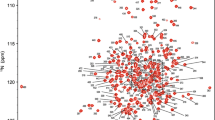
Adenylate kinase reversibly catalyzes the conversion of ATP plus AMP to two ADPs. This essential catalyst is present in every cell, and the Escherichia coli protein is often employed as a model enzyme. Our aim is to use the E. coli enzyme to understand dry protein structure and protection. Here, we report the expression, purification, steady-state assay, NMR conditions and 1H, 13C, 15N backbone resonance NMR assignments of its C77S variant. These data will also help others utilize this prototypical enzyme.

Similar content being viewed by others
Data availability
The assignments have been deposited to the BMRB under the accession code 51,973. The plasmid map for pet28b(+)/ADK C77S, and the plasmid itself, is available from Addgene as plasmid # 201,409.
References
Ådén J, Verma A, Schug A, Wolf-Watz M (2012) Modulation of a pre-existing conformational equilibrium tunes adenylate kinase activity. J Am Chem Soc 134(40):16562–16570. https://doi.org/10.1021/ja3032482
Brom JA, Petrikis RG, Pielak GJ (2023) How sugars protect dry protein structure. Biochemistry 62(5):1044–1052. https://doi.org/10.1021/acs.biochem.2c00692
Burlacu-Miron S, Gilles A-M, Popescu A, BaÃrzu O, Craescu CT (1999a) Multinuclear magnetic resonance studies of Escherichia coli adenylate kinase in free and bound forms: resonance assignment, secondary structure and ligand binding. Eur J Biochem 264:765–774. https://doi.org/10.1046/j.1432-1327.1999.00633.x
Burlacu-Miron S, Perrier V, Gilles AM, Mispelter J, Bârzu O, Craescu CT (1999b) Letter to the editor: 1H, 13 C and 15 N backbone resonance assignment of Escherichia coli adenylate kinase, a 23.6 kDa protein. J Biomol NMR 13(1):93–94
Crilly CJ, Brom JA, Kowalewski ME, Piszkiewicz S, Pielak GJ (2021a) Dried protein structure revealed at the residue level by liquid-observed vapor exchange NMR. Biochemistry 60(2):152–159. https://doi.org/10.1021/acs.biochem.0c00863
Crilly CJ, Eicher JE, Warmuth O, Atkin JM, Pielak GJ (2021b) Water’s variable role in protein stability uncovered by liquid-observed vapor exchange NMR. Biochemistry. https://doi.org/10.1021/acs.biochem.1c00552
Crilly CJ, Brom JA, Warmuth O, Esterly HJ, Pielak GJ (2022) Protection by desiccation-tolerance proteins probed at the residue level. Protein Sci 31(2):396–406. https://doi.org/10.1002/pro.4231
Duggan B (2022) Biomolecular NMR tools: Isotopic labelling. 2023, from http://sopnmr.ucsd.edu/biomol-tools.htm
Good NE, Winget GD, Winter W, Connolly TN, Izawa S, Singh RMM (1966) Hydrogen ion buffers for biological research. Biochemistry 5(2):467–477. https://doi.org/10.1021/bi00866a011
Hyberts SG, Takeuchi K, Wagner G (2010) Poisson-gap sampling and forward maximum entropy reconstruction for enhancing the resolution and sensitivity of protein NMR data. J Am Chem Soc 132(7):2145–2147. https://doi.org/10.1021/ja908004w
Kerns SJ, Agafonov RV, Cho YJ, Pontiggia F, Otten R, Pachov DV, Kutter S, Phung LA, Murphy PN, Thai V, Alber T, Hagan MF, Kern D (2015) The energy landscape of adenylate kinase during catalysis. Nat Struct Mol Biol 22(2):124–131. https://doi.org/10.1038/nsmb.2941
Miklos AC, Li C, Pielak GJ (2009) Using NMR-detected backbone amide 1H exchange to assess macromolecular crowding effects on globular-protein stability. Methods Enzymol 466:1–18. https://doi.org/10.1016/s0076-6879(09)66001-8
Miljenović TM, Jia X, Mobli M (2017) Nonuniform sampling in biomolecular NMR. In: Webb, G. (eds) Mod Magn Reson. Springer International Publishing, Cham, pp 1–21
Munier-Lehmann H, Burlacu-Miron S, Craescu CT, Mantsch HH, Schultz CP (1999) A new subfamily of short bacterial adenylate kinases with the Mycobacterium tuberculosis enzyme as a model: a predictive and experimental study. Proteins Struct Funct Genet 36(2):238–248. https://doi.org/10.1002/(SICI)1097-0134(19990801)36:2%3C238::AID-PROT9%3E3.0.CO;2-K
Ojeda-May P, Mushtaq AU, Rogne P, Verma A, Ovchinnikov V, Grundstrom C, Dulko-Smith B, Sauer UH, Wolf-Watz M, Nam K (2021) Dynamic connection between enzymatic catalysis and collective protein motions. Biochemistry https://doi.org/10.1021/acs.biochem.1c00221
Pielak GJ (2021) Buffers, especially the good kind. Biochemistry 60:3436–3440. https://doi.org/10.1021/acs.biochem.1c00200
Saint Girons I, Gilles AM, Margarita D, Michelson S, Monnot M, Fermandjian S, Danchin A, Bârzu O (1987) Structural and catalytic characteristics of Escherichia coli adenylate kinase. J Biol Chem 262(2):622–629. https://doi.org/10.1016/s0021-9258(19)75829-3
Sambrook J (2001) In: Russell D (ed) Molecular cloning: a laboratory manual, vol A22. Cold Spring Harbor Laboratory Press, New York
Sarkar M, Lu J, Pielak GJ (2014) Protein crowder charge and protein stability. Biochemistry 53(10):1601–1606. https://doi.org/10.1021/bi4016346
Wolf-Watz M, Thai V, Henzler-Wildman K, Hadjipavlou G, Eisenmesser EZ, Kern D (2004) Linkage between dynamics and catalysis in a thermophilic-mesophilic enzyme pair. Nat Struct Mol Biol 11(10):945–949. https://doi.org/10.1038/nsmb821
Ying J, Delaglio F, Torchia DA, Bax A (2017) Sparse multidimensional iterative lineshape-enhanced (SMILE) reconstruction of both non-uniformly sampled and conventional NMR data. J Biomol NMR 68(2):101–118. https://doi.org/10.1007/s10858-016-0072-7
Acknowledgements
The authors thank Brandie Ehrmann from the UNC Chemistry Mass Spectrometry Core Laboratory for equipment maintenance and advice, the UNC Biomolecular NMR Laboratory for spectrometer maintenance and advice, Elizabeth Pielak for comments on the manuscript, and the Pielak lab for helpful discussion.
Funding
The UNC Chemistry Mass Spectrometry Core Laboratory is supported by the National Science Foundation (CHE-1726291) and the UNC Biomolecular NMR Laboratory receives funding from the National Cancer Institute of the National Institutes of Health (P30CA016086). This research was supported by the National Institutes of Health (R01GM127291) and the National Science Foundation (CHE-2203505).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. JAB: material preparation, data collection, data processing, data analysis, figure preparation, initial draft of the manuscript. SS: material preparation, data collection, data processing, data analysis. RGP: material preparation, data collection. SP: data processing, data analysis. GJP: funding, data analysis. All the authors edited the manuscript. All the authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Ethical standards
The experiments comply with the current laws of the United States.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Brom, J.A., Samsri, S., Petrikis, R.G. et al. 1H, 13C, 15N backbone resonance assignment of Escherichia coli adenylate kinase. Biomol NMR Assign 17, 235–238 (2023). https://doi.org/10.1007/s12104-023-10147-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-023-10147-1