Abstract
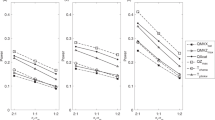
The ‘large p, small n’ problem in genomewide association studies (GWAS) is an important subject in genetic studies. Many approaches have been proposed for this issue, but none of them successfully combine the Haseman–Elston (H–E) regression with sliding-window scan approaches in GWAS. In this article, we extended H–E regression to GWAS, and replaced original data with different measurements of phenotype of sib pairs. Meanwhile, we also applied hidden Markov model to infer identity by state. Using subsequent simulation studies, we found that it had higher statistical power than the corresponding single-marker association studies. The advantage of the H–E regression was also sufficient to capture about 48.01% of the quantitative trait locus (QTL). Meanwhile, the results show that the power decreases with the increase in the number of QTLs, and the power of H–E regression is sensitive to heritability.



Similar content being viewed by others
References
Atwell S., Huang Y. S., Vilhjalmsson B. J., Willems G., Horton M., Li Y. et al. 2010 Genome-wide association study of 107 phenotypes in Arabidopsis thaliana inbred lines. Nature 465, 627–631.
Barber M. J., Cordell H. J., MacGregor A. J. and Andrew T. 2004 Gamma regression improves Haseman–Elston and variance components linkage analysis for sib-pairs. Genet. Epidemiol. 26, 97–107.
Bercovici S., Meek C., Wexler Y. and Geiger D. 2010 Estimating genome-wide IBD sharing from SNP data via an efficient hidden Markov model of LD with application to gene mapping. Bioinformatics 26, i175–i182.
Chen G. B. 2014 Estimating heritability of complex traits from genome-wide association studies using IBS-based Haseman–Elston regression. Front. Genet. 5, 107.
Daetwyler H. D., Calus M. P., Pong-Wong R., de Los Campos G. and Hickey J. M. 2013 Genomic prediction in animals and plants: simulation of data, validation, reporting, and benchmarking. Genetics 193, 347–365.
de Los Campos G., Hickey J. M., Pong-Wong R., Daetwyler H. D. and Calus M. P. 2013 Whole-genome regression and prediction methods applied to plant and animal breeding. Genetics 193, 327–345.
DeFries J. C. 2010 Haseman and Elston sib-pair linkage analysis: a brief historical note. Behav. Genet. 40, 1–2.
Diao G. and Vidyashankar A. N. 2013 Assessing genome-wide statistical significance for large p small n problems. Genetics 194, 781–783.
Drigalenko E. 1999 Matrix representation of the Haseman–Elston method. Theor. Popul. Biol. 55, 157–165.
Elston R. C., Buxbaum S., Jacobs K. B. and Olson J. M. 2000 Haseman and Elston revisited. Genet. Epidemiol. 19, 1–17.
Etzel C. J., Shete S., Beasley T. M., Fernandez J. R., Allison D. B. and Amos C. I. 2003 Effect of Box–Cox transformation on power of Haseman–Elston and maximum-likelihood variance components tests to detect quantitative trait loci. Hum. Hered. 55, 108–116.
Forrest W. F. 2001 Weighting improves the new Haseman–Elston method. Hum. Hered. 52, 47–54.
Franke D., Kleensang A., Elston R. C. and Ziegler A. 2005 Haseman–Elston weighted by marker informativity. BMC Genet. 6 suppl 1, S50.
Garner C. P. 2002 Nonparametric linkage analysis. I. Haseman–Elston. Methods Mol. Biol. 195, 37–60.
Gerhard D. and Hothorn L. A. 2010 Rank transformation in Haseman–Elston regression using scores for location-scale alternatives. Hum. Hered. 69, 143–151.
Hadicke O., Pahlke F. and Ziegler A. 2008 A general approach for sample size and power calculations based on the Haseman–Elston method. Biom. J. 50, 257–269.
Legarra A. and Misztal I. 2008 Technical note: computing strategies in genome-wide selection. J. Dairy. Sci. 91, 360–366.
Meuwissen T. H., Hayes B. J. and Goddard M. E. 2001 Prediction of total genetic value using genome-wide dense marker maps. Genetics 157, 1819–1829.
Sham P. C. and Purcell S. 2001 Equivalence between Haseman–Elston and variance-components linkage analyses for sib pairs. Am. J. Hum. Genet. 68, 1527–1532.
Shen X., Alam M., Fikse F. and Ronnegard L. 2013 A novel generalized ridge regression method for quantitative genetics. Genetics 193, 1255–1268.
Shete S., Jacobs K. B. and Elston R. C. 2003 Adding further power to the Haseman and Elston method for detecting linkage in larger sibships: weighting sums and differences. Hum. Hered. 55, 79–85.
Single R. M. and Finch S. J. 1995 Gain in efficiency from using generalized least squares in the Haseman–Elston test. Genet. Epidemiol. 12, 889–894.
Solberg Woods L. C., Holl K., Tschannen M. and Valdar W. 2010 Fine-mapping a locus for glucose tolerance using heterogeneous stock rats. Physiol. Genomics 41, 102–108.
Stoesz M. R., Cohen J. C., Mooser V, Marcovina S. and Guerra R. 1997 Extension of the Haseman–Elston method to multiple alleles and multiple loci: theory and practice for candidate genes. Ann. Hum. Genet. 61, 263–274.
Valdar W., Solberg L. C., Gauguier D., Burnett S., Klenerman P., Cookson W. O. et al. 2006 Genome-wide genetic association of complex traits in heterogeneous stock mice. Nat. Genet. 38, 879–887.
Wang T. and Elston R. C. 2005 Two-level Haseman–Elston regression for general pedigree data analysis. Genet. Epidemiol. 29, 12–22.
Weeks D. E. and Harby L. D. 1995 The affected-pedigree-member method: power to detect linkage. Hum. Hered. 45, 13–24.
Won S., Elston R. C. and Park T. 2006 Extension of the Haseman–Elston regression model to longitudinal data. Hum. Hered. 61, 111–119.
Xu X., Weiss S., Xu X. and Wei L. J. 2000 A unified Haseman–Elston method for testing linkage with quantitative traits. Am. J. Hum. Genet. 67, 1025–1028.
Yoon S., Suh Y. J., Mendell N. R. and Ye K. Q. 2005 A Bayesian approach for applying Haseman–Elston methods. BMC Genet. 6 suppl 1, S39.
Yu T., Ye H., Sun W., Li K. C., Chen Z., Jacobs S. et al. 2007 A forward–backward fragment assembling algorithm for the identification of genomic amplification and deletion breakpoints using high-density single nucleotide polymorphism (SNP) array. BMC Bioinformatics 8, 145.
Zhang Y. M., Lu H. Y. and Yao L. L. 2008 Multiple quantitative trait loci Haseman–Elston regression using all markers on the entire genome. Theor. Appl. Genet. 117, 683–690.
Ziegler A., Boddeker I. R. and Geller F. 2001 A bivariate Haseman–Elston method and application to the analysis of asthma-related phenotypes on chromosome 5q. Genet. Epidemiol. 21 suppl 1, S216–S221.
Acknowledgements
We thank the editor and referees for helpful comments. This work was supported by the National Natural Science Foundation of China (grant no. 31460594), China Scholarship Council (grant no. 201308155140), Hetao College teaching and research project (grant no. HTXYJZ14005).
Author information
Authors and Affiliations
Corresponding author
Additional information
Corresponding editor: Rajiva Raman
Bujun Mei initiated the idea, developed the theory and derived the equations; and wrote the paper. Zhihua Wang conducted the simulation studies and obtained the analytical results. All authors approved the final version of the paper.
[Mei B. and Wang Z. 2016 An efficient method to handle the ‘large p, small n’ problem for genomewide association studies using Haseman–Elston regression. J. Genet. 95, xx–xx]
Rights and permissions
About this article
Cite this article
MEI, B., WANG, Z. An efficient method to handle the ‘large p, small n’ problem for genomewide association studies using Haseman–Elston regression. J Genet 95, 847–852 (2016). https://doi.org/10.1007/s12041-016-0705-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12041-016-0705-3