Abstract
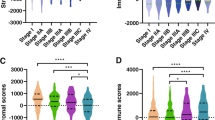
The development and progression of breast cancer (BC) depend heavily on the tumor microenvironment (TME), especially tumor infiltration leukocytes (TILs). TME-based classifications in BC remain largely unknown and need to be clarified. Using the bioinformatic analysis, we attempted to construct a prognostic nomogram based on clinical features and TME-related differentially expressed genes (DEGs). We also tried to investigate the association between the prognostic nomogram and clinical characteristics, TILs, possible signaling pathways, and response to immunotherapy in BC patients. DEGs for BC patients were identified from The Cancer Genome Atlas Breast Invasive Carcinoma database. TME-related genes were downloaded from the Immunology Database and Analysis Portal. After intersecting DEGs and TME-related genes, 3985 overlapping TME-related DEGs were selected for non-negative matrix factorization clustering, microenvironment cell populations-counter (MCP-counter), LASSO Cox regression, tumor immune dysfunction, and exclusion (TIDE) algorithm analyses. BC patients were divided into three clusters based on the TME-related DEGs and survival data, in which cluster 3 had the best overall survival (OS). Of note, cluster 3 exhibited the highest infiltration or lowest infiltration of CD3+ T-cells, CD8+ T-cells, cytotoxic lymphocytes, B-lymphocytes, monocytic lineage, and myeloid dendritic cells (MDCs). A total of 33 TME-related DEGs were identified as a prognostic gene signature by the LASSO regression analysis. The prognostic gene signature separated BC patients into low- and high-risk groups with significant differences in OS (p<0.01) and demonstrated powerful effectiveness (TCGA all group: 1-year area under the curve [AUC] = 0.773, 3-year AUC = 0.770, 5-year AUC = 0.792). By integrating demographic features, tumor-node metastasis (TNM) stages, and prognostic gene signature, we constructed a nomogram with better predictive value than other clinical features alone. TME-related DEGs in the low-risk BC patients (with better OS) were enriched in chemokine, cytokine–cytokine receptor interaction, and JAK-STAT and Toll-like receptor signaling pathways. BC patients in the low-risk group exhibited higher TIDE scores associated with worse immune checkpoint blockade response. A prognostic nomogram based on TME-related DEGs and clinical characteristics could predict prognosis and guide immunotherapy in BC patients.









Similar content being viewed by others
Data availability statement
The gene expression data, corresponding clinicopathological information, somatic mutation information, and the tumor mutation burden of The Cancer Genome Atlas Breast Invasive Carcinoma (TCGA-BRCA) were downloaded from the Genomic Data Commons portal (https://portal.gdc.cancer.gov). The GSE159956 dataset used for validation were retrieved from the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/). TME-related genes were downloaded from the Immunology Database and Analysis Portal (ImmPort) website (https://www.immport.org).
References
Asati V, Mahapatra DK and Bharti SK 2016 PI3K/Akt/mTOR and Ras/Raf/MEK/ERK signaling pathways inhibitors as anticancer agents: Structural and pharmacological perspectives. Eur. J. Med. Chem. 109 314–341
Baptista MZ, Sarian LO, Derchain SF, et al. 2016 Prognostic significance of PD-L1 and PD-L2 in breast cancer. Hum. Pathol. 47 78–84
Becht E, Giraldo NA, Lacroix L, et al. 2016 Estimating the population abundance of tissue-infiltrating immune and stromal cell populations using gene expression. Genome Biol. 17 218
Benelli R, Morini M, Carrozzino F, et al. 2002 Neutrophils as a key cellular target for angiostatin: implications for regulation of angiogenesis and inflammation. FASEB J. 16 267–269
DeNardo DG, Brennan DJ, Rexhepaj E, et al. 2011 Leukocyte complexity predicts breast cancer survival and functionally regulates response to chemotherapy. Cancer Discov. 1 54–67
Denkert C, Loibl S, Noske A, et al. 2010 Tumor-associated lymphocytes as an independent predictor of response to neoadjuvant chemotherapy in breast cancer. J. Clin. Oncol. 28 105–113
Denkert C, Liedtke C, Tutt A, et al. 2017 Molecular alterations in triple-negative breast cancer-the road to new treatment strategies. Lancet 389 2430–2442
Denkert C, von Minckwitz G, Darb-Esfahani S, et al. 2018 Tumour-infiltrating lymphocytes and prognosis in different subtypes of breast cancer: a pooled analysis of 3771 patients treated with neoadjuvant therapy. Lancet Oncol. 19 40–50
Dieci MV, Miglietta F and Guarneri V 2021 Immune infiltrates in breast cancer: Recent updates and clinical implications. Cells 10 223
Friedman J, Hastie T and Tibshirani R 2010 Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33 1–22
Garaud S, Buisseret L, Solinas C, et al. 2019 Tumor infiltrating B-cells signal functional humoral immune responses in breast cancer. JCI Insight 5 e129641
Gaujoux R and Seoighe C 2010 A flexible R package for nonnegative matrix factorization. BMC Bioinform. 11 367
Goel MK, Khanna P and Kishore J 2010 Understanding survival analysis: Kaplan-Meier estimate. Int. J. Ayurveda Res. 1 274–278
Guo E, Ishii Y, Mueller J, et al. 2020 FEN1 endonuclease as a therapeutic target for human cancers with defects in homologous recombination. Proc. Natl. Acad. Sci. USA 117 19415–19424
Hanahan D 2022 Hallmarks of cancer: New dimensions. Cancer Discov. 12 31–46
Hanahan D and Coussens LM 2012 Accessories to the crime: functions of cells recruited to the tumor microenvironment. Cancer Cell 21 309–322
Hanahan D and Weinberg RA 2011 Hallmarks of cancer: the next generation. Cell 144 646–674
Henriques B, Mendes F and Martins D 2021 Immunotherapy in breast cancer: When, how, and what challenges? Biomedicines 9 1687
Hao R, Liu Y, Du Q, et al. 2019 Transgelin-2 expression in breast cancer and its relationships with clinicopathological features and patient outcome. Breast Cancer 26 776–783
Iasonos A, Schrag D, Raj GV, et al. 2008 How to build and interpret a nomogram for cancer prognosis. J. Clin. Oncol. 26 1364–1370
Jardim DL, Goodman A, de Melo Gagliato D, et al. 2021 The challenges of tumor mutational burden as an immunotherapy biomarker. Cancer Cell 39 154–173
Jiang P, Gu S, Pan D, et al. 2018 Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat. Med. 24 1550–1558
Joyce JA and Fearon DT 2015 T cell exclusion, immune privilege, and the tumor microenvironment. Science 348 74–80
Liu H, Yang Z, Lu W, et al. 2020 Chemokines and chemokine receptors: A new strategy for breast cancer therapy. Cancer Med. 9 3786–3799
Mierke CT 2019 The matrix environmental and cell mechanical properties regulate cell migration and contribute to the invasive phenotype of cancer cells. Rep. Prog. Phys. 82 064602
Newman AM, Liu CL, Green MR, et al. 2015 Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12 453–457
Possemato R, Marks KM, Shaul YD, et al. 2011 Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature 476 346–350
Riggio AI, Varley KE and Welm AL 2021 The lingering mysteries of metastatic recurrence in breast cancer. Br. J. Cancer 124 13–26
Ritchie ME, Phipson B, Wu D, et al. 2015 limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43 e47
Rupp T, Genest L, Babin D, et al. 2022 Anti-CTLA-4 and anti-PD-1 immunotherapies repress tumor progression in preclinical breast and colon model with independent regulatory T cells response. Transl. Oncol. 20 101405
Sauerbrei W, Royston P and Binder H 2007 Selection of important variables and determination of functional form for continuous predictors in multivariable model building. Stat. Med. 26 5512–5528
Seo AN, Lee HJ, Kim EJ, et al. 2013 Tumour-infiltrating CD8+ lymphocytes as an independent predictive factor for pathological complete response to primary systemic therapy in breast cancer. Br. J. Cancer 109 2705–2713
Shao G, Fan X, Zhang P, et al. 2021 Methylation-dependent MCM6 repression induced by LINC00472 inhibits triple-negative breast cancer metastasis by disturbing the MEK/ERK signaling pathway. Aging 13 4962–4975
Shaul ME and Fridlender ZG 2019 Tumour-associated neutrophils in patients with cancer. Nat. Rev. Clin. Oncol. 16 601–620
Siegel RL, Miller KD and Jemal A 2020 Cancer statistics, 2020. CA Cancer J. Clin. 70 7–30
Subramanian A, Tamayo P, Mootha VK, et al. 2005 Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102 15545–15550
Sung H, Ferlay J, Siegel RL, et al. 2021 Global Cancer Statistics 2020: GLOBOCAN Estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71 209–249
Vickers AJ and Elkin EB 2006 Decision curve analysis: a novel method for evaluating prediction models. Med. Decis. Making 26 565–574
West NR, Milne K, Truong PT, et al. 2011 Tumor-infiltrating lymphocytes predict response to anthracycline-based chemotherapy in estrogen receptor-negative breast cancer. Breast Cancer Res. 13 R126
Yu G, Wang LG, Han Y, et al. 2012 clusterProfiler: an R package for comparing biological themes among gene clusters. Omics 16 284–287
Zhao Y, Rahmy S, Liu Z, et al. 2020 Rational targeting of immunosuppressive neutrophils in cancer. Pharmacol. Ther. 212 107556
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
YY designed the study. LY and HX performed analysis. YY and LY were involved in data discussion, drafting, and editing the analysis. All authors contributed to the article and approved the final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare.
Ethics statement
This study was conducted following the Helsinki Declaration II and was approved by the Institutional Review Boards of Chongqing University Cancer Hospital.
Additional information
Corresponding editor: Ramray Bhat
Supplementary Information
Below is the link to the electronic supplementary material.
12038_2023_386_MOESM1_ESM.tif
Supplementary file1 Supplementary Figure 1. Heatmap for 2-10 clusters based on TME-related DEGs and survival data (TIF 3954 KB)
12038_2023_386_MOESM2_ESM.tif
Supplementary file2 Supplementary Figure 2. General distribution of different immune cells in each sample (TIF 11765 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liu, Y., He, X. & Yang, Y. Tumor immune microenvironment-based clusters in predicting prognosis and guiding immunotherapy in breast cancer. J Biosci 49, 19 (2024). https://doi.org/10.1007/s12038-023-00386-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12038-023-00386-8