Abstract
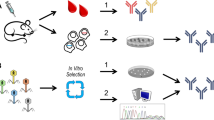
Synthetic antibodies (Abs) represent a category of engineered proteins meticulously crafted to replicate the functions of their natural counterparts. Such Abs are generated in vitro, enabling advanced molecular alterations associated with antigen recognition, paratope site engineering, and biochemical refinements. In a parallel realm, deep sequencing has brought about a paradigm shift in molecular biology. It facilitates the prompt and cost-effective high-throughput sequencing of DNA and RNA molecules, enabling the comprehensive big data analysis of Ab transcriptomes, including specific regions of interest. Significantly, the integration of artificial intelligence (AI), based on machine- and deep- learning approaches, has fundamentally transformed our capacity to discern patterns hidden within deep sequencing big data, including distinctive Ab features and protein folding free energy landscapes. Ultimately, current AI advances can generate approximations of the most stable Ab structural configurations, enabling the prediction of de novo synthetic Abs. As a result, this manuscript comprehensively examines the latest and relevant literature concerning the intersection of deep sequencing big data and AI methodologies for the design and development of synthetic Abs. Together, these advancements have accelerated the exploration of antibody repertoires, contributing to the refinement of synthetic Ab engineering and optimizations, and facilitating advancements in the lead identification process.


Similar content being viewed by others
Data Availability
Not applicable.
Abbreviations
- Abs:
-
Antibodies
- AI:
-
Artificial intelligence
- NGS:
-
Next-generation sequencing
- CDR:
-
Complementary determining region
- BCR:
-
B-cell receptor
- HER2:
-
Human epidermal growth factor receptor 2
- AbLang:
-
Antibody language
- AbRep:
-
Antibody representation
- AbHead:
-
Antibody head model
- AntBO:
-
Antibody design based on a Bayesian Optimization framework
- AbMap:
-
Antibody binding epitope mapping
- AntiBERTa:
-
Antibody-specific Bidirectional Encoder Representation from Transformers
- AbMap:
-
Antibody binding epitope Mapping
- OptMAVEn:
-
Optimal Method for Antibody Variable region Engineering
- OptCDR:
-
Optimal CDR
- AbDesign:
-
Antibody design
- RAbD:
-
Rosetta Ab design
- 2D:
-
Two-dimensional
- 3D:
-
Three-dimensional
References
Bradbury, A. R. M., Sidhu, S. S., Dübel, S., McCafferty, J., & Dubel, S. (2011). Beyond natural antibodies: The power of in vitro display technologies. Nature biotechnology, 29(3), 245–254. https://doi.org/10.1038/nbt.1791
Adams, J. J., & Sidhu, S. S. (2014). Synthetic antibody technologies. Current Opinion in Structural Biology, 24(1), 1–9. https://doi.org/10.1016/j.sbi.2013.11.003
Chen, G., & Sidhu, S. S. (2014). Design and generation of synthetic antibody libraries for phage display. Methods in Molecular Biology (Clifton, N.J.), 1131, 113–131. https://doi.org/10.1007/978-1-62703-992-5_8
Chen, G., Gorelik, L., Simon, K. J., Pavlenco, A., Cheung, A., Brickelmaier, M., & Sidhu, S. S. (2015). Synthetic antibodies and peptides recognizing progressive multifocal leukoencephalopathyspecific point mutations in polyomavirus JC capsid viral protein 1. MAbs, 7(4), 681–692. https://doi.org/10.1080/19420862.2015.1038447
Miersch, S., & Sidhu, S. S. (2012). Synthetic antibodies: Concepts, potential and practical considerations. Methods, 57(4), 486–498. https://doi.org/10.1016/j.ymeth.2012.06.012
Sidhu, S. S., & Fellouse, Fa. (2006). Synthetic therapeutic antibodies. Nature Chemical Biology, 2(12), 682–688. https://doi.org/10.1038/nchembio843
Fuh, G. (2007). Synthetic antibodies as therapeutics. Expert Opinion on Biological Therapy, 7(1), 73–87. https://doi.org/10.1517/14712598.7.1.73
Metzker, M. L. (2009). Sequencing technologies — the next generation. Nature Reviews Genetics, 11(1), 31–46. https://doi.org/10.1038/nrg2626
Goodwin, S., McPherson, J. D., & McCombie, W. R. (2016). Coming of age: Ten years of next-generation sequencing technologies. Nature Reviews Genetics, 17(6), 333–351. https://doi.org/10.1038/nrg.2016.49
Zhang, J., Chiodini, R., Badr, A., & Zhang, G. (2011). The impact of next-generation sequencing on genomics. Journal of Genetics and Genomics, 38(3), 95–109. https://doi.org/10.1016/J.JGG.2011.02.003
Mardis, E. R. (2013). Next-generation sequencing platforms. Annual Review of Analytical Chemistry, 6(1), 287–303. https://doi.org/10.1146/annurev-anchem-062012-092628
Shendure, J., & Aiden, E. L. (2012). The expanding scope of DNA sequencing. Nature Biotechnology, 30(11), 1084–1094. https://doi.org/10.1038/nbt.2421
Meaburn, E., & Schulz, R. (2012). Next generation sequencing in epigenetics: Insights and challenges. Seminars in Cell & Developmental Biology, 23(2), 192–199. https://doi.org/10.1016/J.SEMCDB.2011.10.010
Raoult, D. (2008). Obesity pandemics and the modification of digestive bacterial flora. European Journal of Clinical Microbiology and Infectious Diseases. https://doi.org/10.1007/s10096-008-0490-x
Jünemann, S., Kleinbölting, N., Jaenicke, S., Henke, C., Hassa, J., Nelkner, J., & Stoye, J. (2017). Bioinformatics for NGS-based metagenomics and the application to biogas research. Journal of Biotechnology, 261, 10–23. https://doi.org/10.1016/J.JBIOTEC.2017.08.012
Dong, Z. C., & Chen, Y. (2013). Transcriptomics: Advances and approaches. Science China Life Sciences, 56(10), 960–967. https://doi.org/10.1007/S11427-013-4557-2/METRICS
Morganti, S., Tarantino, P., Ferraro, E., D’Amico, P., Duso, B. A., & Curigliano, G. (2019). Next generation sequencing (NGS): A revolutionary technology in pharmacogenomics and personalized medicine in cancer. Advances in Experimental Medicine and Biology, 1168, 9–30. https://doi.org/10.1007/978-3-030-24100-1_2
Frese, K. S., Katus, H. A., & Meder, B. (2013). Next-generation sequencing: From understanding biology to personalized medicine. Biology, 2(1), 378–398. https://doi.org/10.3390/BIOLOGY2010378
Rabbani, B., Nakaoka, H., Akhondzadeh, S., Tekin, M., & Mahdieh, N. (2016). Next generation sequencing: Implications in personalized medicine and pharmacogenomics. Molecular BioSystems, 12(6), 1818–1830. https://doi.org/10.1039/C6MB00115G
Hong, H., Zhang, W., Su, Z., Shen, J., Ge, W., Ning, B., & Tong, W. (2013). Next-generation sequencing (NGS): A revolutionary technology in pharmacogenomics and personalized medicine. Omics for personalized medicine (pp. 39–61). New Delhi: Springer. https://doi.org/10.1007/978-81-322-1184-6_3
Quail, M. A., Smith, M., Coupland, P., Otto, T. D., Harris, S. R., Connor, T. R., & Gu, Y. (2012). A tale of three next generation sequencing platforms: comparison of Ion Torrent, Pacific Biosciences and Illumina MiSeq sequencers. BMC Genomics, 13(1), 341. https://doi.org/10.1186/1471-2164-13-341
Meng, W., Zhang, B., Schwartz, G. W., Rosenfeld, A. M., Ren, D., Thome, J. J. C., & Luning Prak, E. T. (2017). An atlas of B-cell clonal distribution in the human body. Nature Biotechnology, 35(9), 879–884. https://doi.org/10.1038/nbt.3942
Georgiou, G., Ippolito, G. C., Beausang, J., Busse, C. E., Wardemann, H., & Quake, S. R. (2014). The promise and challenge of high-throughput sequencing of the antibody repertoire. Nature Biotechnology, 32(2), 158–168. https://doi.org/10.1038/nbt.2782
Robins, H. (2013). Immunosequencing: Applications of immune repertoire deep sequencing. Current Opinion in Immunology, 25(5), 646–652. https://doi.org/10.1016/J.COI.2013.09.017
Waight, A. B., Prihoda, D., Shrestha, R., Metcalf, K., Bailly, M., Ancona, M., & Fayadat-Dilman, L. (2023). A machine learning strategy for the identification of key in silico descriptors and prediction models for IgG monoclonal antibody developability properties. MAbs. https://doi.org/10.1080/19420862.2023.2248671
Mathonet, P., & Ullman, C. G. (2013). The application of next generation sequencing to the understanding of antibody repertoires. Frontiers in Immunology, 4(SEP), 56179. https://doi.org/10.3389/fimmu.2013.00265
Glanville, J., D’Angelo, S., Khan, T. A., Reddy, S. T., Naranjo, L., Ferrara, F., & Bradbury, A. R. M. (2015). Deep sequencing in library selection projects: What insight does it bring? Current Opinion in Structural Biology, 33, 146–160. https://doi.org/10.1016/J.SBI.2015.09.001
Rouet, R., Jackson, K. J. L., Langley, D. B., & Christ, D. (2018). Next-generation sequencing of antibody display repertoires. Frontiers in Immunology, 9(FEB), 334733. https://doi.org/10.3389/fimmu.2018.00118
Vaisman-Mentesh, A., & Wine, Y. (2018). Monitoring phage biopanning by next-generation sequencing. Methods in molecular biology (Clifton, N.J.), 1701, 463–473. https://doi.org/10.1007/978-1-4939-7447-4_26
Friedensohn, S., Neumeier, D., Khan, T. A., Csepregi, L., Parola, C., de Vries, A. R. G., & Reddy, S. T. (2020). Convergent selection in antibody repertoires is revealed by deep learning. bioRxiv. https://doi.org/10.1101/2020.02.25.965673
Lim, Y. W., Adler, A. S., & Johnson, D. S. (2022). Predicting antibody binders and generating synthetic antibodies using deep learning. MAbs. https://doi.org/10.1080/19420862.2022.2069075
Lecun, Y., Bengio, Y., & Hinton, G. (2015). Deep learning. Nature, 521(7553), 436–444. https://doi.org/10.1038/NATURE14539
Hsu, H. J., Lee, K. H., Jian, J. W., Chang, H. J., Yu, C. M., Lee, Y. C., & Yang, A. S. (2014). Antibody variable domain interface and framework sequence requirements for stability and function by high-throughput experiments. Structure, 22(1), 22–34. https://doi.org/10.1016/j.str.2013.10.006
Fischman, S., & Ofran, Y. (2018). Computational design of antibodies. Current opinion in structural biology, 51, 156–162. https://doi.org/10.1016/J.SBI.2018.04.007
Huang, P. S., Boyken, S. E., & Baker, D. (2016). The coming of age of de novo protein design. Nature, 537(7620), 320–327. https://doi.org/10.1038/nature19946
Sormanni, P., Aprile, F. A., & Vendruscolo, M. (2018). Third generation antibody discovery methods: In silico rational design. Chemical Society Reviews, 47(24), 9137–9157. https://doi.org/10.1039/C8CS00523K
Cao, H., Wang, J., He, L., Qi, Y., & Zhang, J. Z. (2019). DeepDDG: Predicting the stability change of protein point mutations using neural networks. Journal of Chemical Information and Modeling, 59(4), 1508–1514. https://doi.org/10.1021/ACS.JCIM.8B00697/SUPPL_FILE/CI8B00697_SI_003.XLSX
Dehouck, Y., Kwasigroch, J. M., Gilis, D., & Rooman, M. (2011). PoPMuSiC 2.1: A web server for the estimation of protein stability changes upon mutation and sequence optimality. BMC Bioinformatics, 12(1), 151. https://doi.org/10.1186/1471-2105-12-151
Folkman, L., Stantic, B., Sattar, A., & Zhou, Y. (2016). EASE-MM: Sequence-based prediction of mutation-induced stability changes with feature-based multiple models. Journal of Molecular Biology, 428(6), 1394–1405. https://doi.org/10.1016/J.JMB.2016.01.012
Pires, D. E. V., Ascher, D. B., & Blundell, T. L. (2014). mCSM: Predicting the effects of mutations in proteins using graph-based signatures. Bioinformatics, 30(3), 335–342. https://doi.org/10.1093/BIOINFORMATICS/BTT691
Capriotti, E., Fariselli, P., Rossi, I., & Casadio, R. (2008). A three-state prediction of single point mutations on protein stability changes. BMC Bioinformatics. https://doi.org/10.1186/1471-2105-9-S2-S6
Quan, L., Lv, Q., & Zhang, Y. (2016). STRUM: Structure-based prediction of protein stability changes upon single-point mutation. Bioinformatics, 32(19), 2936–2946. https://doi.org/10.1093/BIOINFORMATICS/BTW361
Cheng, J., Randall, A., & Baldi, P. (2006). Prediction of protein stability changes for single-site mutations using support vector machines. Proteins: Structure, Function, and Bioinformatics, 62(4), 1125–1132. https://doi.org/10.1002/PROT.20810
Krawczyk, K., Baker, T., Shi, J., & Deane, C. M. (2013). Antibody i-Patch prediction of the antibody binding site improves rigid local antibody-antigen docking. Protein Engineering, Design & Selection : PEDS, 26(10), 621–629. https://doi.org/10.1093/PROTEIN/GZT043
Kunik, V., Ashkenazi, S., & Ofran, Y. (2012). Paratome: An online tool for systematic identification of antigen-binding regions in antibodies based on sequence or structure. Nucleic Acids Research, 40(W1), W521–W524. https://doi.org/10.1093/NAR/GKS480
Olimpieri, P. P., Chailyan, A., Tramontano, A., & Marcatili, P. (2013). Prediction of site-specific interactions in antibody-antigen complexes: The proABC method and server. Bioinformatics (Oxford, England), 29(18), 2285–2291. https://doi.org/10.1093/BIOINFORMATICS/BTT369
Mason, D. M., Friedensohn, S., Weber, C. R., Jordi, C., Wagner, B., Meng, S. M., & Reddy, S. T. (2021). Optimization of therapeutic antibodies by predicting antigen specificity from antibody sequence via deep learning. Nature Biomedical Engineering, 5(6), 600–612. https://doi.org/10.1038/S41551-021-00699-9
Deac, A., VeliČković, P., & Sormanni, P. (2019). Attentive cross-modal paratope prediction. Journal of Computational Biology : A Journal of Computational Molecular Cell Biology, 26(6), 536–545. https://doi.org/10.1089/cmb.2018.0175
Daberdaku, S., & Ferrari, C. (2019). Antibody interface prediction with 3D Zernike descriptors and SVM. Bioinformatics (Oxford, England), 35(11), 1870–1876. https://doi.org/10.1093/bioinformatics/bty918
Liberis, E., Velickovic, P., Sormanni, P., Vendruscolo, M., & Lio, P. (2018). Parapred: Antibody paratope prediction using convolutional and recurrent neural networks. Bioinformatics (Oxford, England), 34(17), 2944–2950. https://doi.org/10.1093/BIOINFORMATICS/BTY305
Ambrosetti, F., Olsen, T. H., Olimpieri, P. P., Jiménez-García, B., Milanetti, E., Marcatilli, P., & Bonvin, A. M. J. J. (2020). proABC-2: PRediction of AntiBody contacts v2 and its application to information-driven docking. Bioinformatics (Oxford, England), 36(20), 5107–5108. https://doi.org/10.1093/BIOINFORMATICS/BTAA644
Qi, H., Ma, M., Hu, C., Xu, Z. W., Wu, F. L., Wang, N., & Tao, S. C. (2021). Antibody binding epitope mapping (AbMap) of hundred antibodies in a single run. Molecular and Cellular Proteomics, 20, 100059. https://doi.org/10.1074/MCP.RA120.002314
Van Blarcom, T., Rossi, A., Foletti, D., Sundar, P., Pitts, S., Melton, Z., & Chaparro-Riggers, J. (2018). Epitope mapping using yeast display and next generation sequencing. Methods in molecular biology (Clifton, N.J.), 1785, 89–118. https://doi.org/10.1007/978-1-4939-7841-0_7
Ibsen, K. N., & Daugherty, P. S. (2017). Prediction of antibody structural epitopes via random peptide library screening and next generation sequencing. Journal of immunological methods, 451, 28–36. https://doi.org/10.1016/J.JIM.2017.08.004
Rebollo, I. R., Sabisz, M., Baeriswyl, V., & Heinis, C. (2014). Identification of target-binding peptide motifs by high-throughput sequencing of phage-selected peptides. Nucleic Acids Research. https://doi.org/10.1093/NAR/GKU940
Ionov, Y., & Rogovskyy, A. S. (2020). Comparison of motif-based and whole-unique-sequence-based analyses of phage display library datasets generated by biopanning of anti-Borrelia burgdorferi immune sera. PLoS ONE, 15(1), e0226378. https://doi.org/10.1371/journal.pone.0226378
Huang, W., Soeung, V., Boragine, D. M., Hu, L., Prasad, B. V. V., Estes, M. K., & Palzkill, T. (2020). High-resolution mapping of human norovirus antigens via genomic phage display library selections and deep sequencing. Journal of Virology. https://doi.org/10.1128/JVI.01495-20
Paull, M. L., Johnston, T., Ibsen, K. N., Bozekowski, J. D., & Daugherty, P. S. (2019). A general approach for predicting protein epitopes targeted by antibody repertoires using whole proteomes. PLoS ONE. https://doi.org/10.1371/JOURNAL.PONE.0217668
Paull, M. L., Bozekowski, J. D., & Daugherty, P. S. (2021). Mapping antibody binding using multiplexed epitope substitution analysis. Journal of Immunological Methods. https://doi.org/10.1016/J.JIM.2021.113178
Norman, R. A., Ambrosetti, F., Bonvin, A. M. J. J., Colwell, L. J., Kelm, S., Kumar, S., & Krawczyk, K. (2020). Computational approaches to therapeutic antibody design: Established methods and emerging trends. Briefings in Bioinformatics, 21(5), 1549–1567. https://doi.org/10.1093/BIB/BBZ095
Buck, P. M., Kumar, S., Wang, X., Agrawal, N. J., Trout, B. L., & Singh, S. K. (2012). Computational methods to predict therapeutic protein aggregation. Methods in Molecular Biology (Clifton, N.J.), 899, 425–451. https://doi.org/10.1007/978-1-61779-921-1_26
Agrawal, N. J., Kumar, S., Wang, X., Helk, B., Singh, S. K., & Trout, B. L. (2011). Aggregation in protein-based biotherapeutics: Computational studies and tools to identify aggregation-prone regions. Journal of pharmaceutical sciences, 100(12), 5081–5095. https://doi.org/10.1002/JPS.22705
Wang, X., Das, T. K., Singh, S. K., & Kumar, S. (2009). Potential aggregation prone regions in biotherapeutics: A survey of commercial monoclonal antibodies. MAbs, 1(3), 254–267. https://doi.org/10.4161/MABS.1.3.8035
Hebditch, M., Carballo-Amador, M. A., Charonis, S., Curtis, R., & Warwicker, J. (2017). Protein-Sol: A web tool for predicting protein solubility from sequence. Bioinformatics (Oxford, England), 33(19), 3098–3100. https://doi.org/10.1093/BIOINFORMATICS/BTX345
Roberts, C. J. (2014). Therapeutic protein aggregation: Mechanisms, design, and control. Trends in biotechnology, 32(7), 372. https://doi.org/10.1016/J.TIBTECH.2014.05.005
Rawat, P., Kumar, S., & Michael Gromiha, M. (2018). An in-silico method for identifying aggregation rate enhancer and mitigator mutations in proteins. International Journal of Biological Macromolecules, 118, 1157–1167. https://doi.org/10.1016/J.IJBIOMAC.2018.06.102
Obrezanova, O., Arnell, A., De La Cuesta, R. G., Berthelot, M. E., Gallagher, T. R. A., Zurdo, J., & Stallwood, Y. (2015). Aggregation risk prediction for antibodies and its application to biotherapeutic development. MAbs, 7(2), 352–363. https://doi.org/10.1080/19420862.2015.1007828
Kuroda, D., & Tsumoto, K. (2020). Engineering stability, viscosity, and immunogenicity of antibodies by computational design. Journal of Pharmaceutical Sciences, 109(5), 1631–1651. https://doi.org/10.1016/J.XPHS.2020.01.011
Chen, Z., Wang, X., Chen, X., Huang, J., Wang, C., Wang, J., & Wang, Z. (2023). Accelerating therapeutic protein design with computational approaches toward the clinical stage. Computational and Structural Biotechnology Journal, 21, 2909–2926. https://doi.org/10.1016/J.CSBJ.2023.04.027
Lippow, S. M., Wittrup, K. D., & Tidor, B. (2007). Computational design of antibody affinity improvement beyond in vivo maturation. Nature biotechnology, 25(10), 1171. https://doi.org/10.1038/NBT1336
Olsen, T. H., Moal, I. H., & Deane, C. M. (2022). AbLang: An antibody language model for completing antibody sequences. Bioinformatics Advances. https://doi.org/10.1093/BIOADV/VBAC046
Kovaltsuk, A., Leem, J., Kelm, S., Snowden, J., Deane, C. M., & Krawczyk, K. (2018). Observed antibody space: A resource for data mining next-generation sequencing of antibody repertoires. The Journal of Immunology, 201(8), 2502–2509. https://doi.org/10.4049/JIMMUNOL.1800708
Leem, J., Mitchell, L. S., Farmery, J. H. R., Barton, J., & Galson, J. D. (2022). Deciphering the language of antibodies using self-supervised learning. Patterns, 3(7), 100513. https://doi.org/10.1016/J.PATTER.2022.100513
Potocnakova, L., Bhide, M., & Pulzova, L. B. (2016). An introduction to B-cell epitope mapping and in silico epitope prediction. Journal of Immunology Research. https://doi.org/10.1155/2016/6760830
Peri, C., Gagni, P., Combi, F., Gori, A., Chiari, M., Longhi, R., & Colombo, G. (2013). Rational epitope design for protein targeting. ACS chemical biology, 8(2), 397–404. https://doi.org/10.1021/CB300487U
Van Blarcom, T., Rossi, A., Foletti, D., Sundar, P., Pitts, S., Bee, C., & Rajpal, A. (2015). Precise and efficient antibody epitope determination through library design, yeast display and next-generation sequencing. Journal of Molecular Biology, 427(6), 1513–1534. https://doi.org/10.1016/J.JMB.2014.09.020
Zhou, P., Yang, X. L., Wang, X. G., Hu, B., Zhang, L., Zhang, W., & Shi, Z. L. (2020). A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature, 579(7798), 270–273. https://doi.org/10.1038/S41586-020-2012-7
Ripoll, D. R., Chaudhury, S., & Wallqvist, A. (2021). Using the antibody-antigen binding interface to train image-based deep neural networks for antibody-epitope classification. PLoS Computational Biology. https://doi.org/10.1371/JOURNAL.PCBI.1008864
Schneidman-Duhovny, D., Inbar, Y., Nussinov, R., & Wolfson, H. J. (2005). PatchDock and SymmDock: Servers for rigid and symmetric docking. Nucleic Acids Research. https://doi.org/10.1093/nar/gki481
Torchala, M., Moal, I. H., Chaleil, R. A. G., Fernandez-Recio, J., & Bates, P. A. (2013). SwarmDock: A server for flexible protein-protein docking. Bioinformatics (Oxford, England), 29(6), 807–809. https://doi.org/10.1093/BIOINFORMATICS/BTT038
Dominguez, C., Boelens, R., & Bonvin, A. M. J. J. (2003). HADDOCK: A protein-protein docking approach based on biochemical or biophysical information. Journal of the American Chemical Society, 125(7), 1731–1737. https://doi.org/10.1021/JA026939X
De Vries, S. J., Schindler, C. E. M., Chauvot De Beauchêne, I., & Zacharias, M. (2015). A web interface for easy flexible protein-protein docking with ATTRACT. Biophysical journal, 108(3), 462–465. https://doi.org/10.1016/J.BPJ.2014.12.015
Jiménez-García, B., Pons, C., & Fernández-Recio, J. (2013). pyDockWEB: A web server for rigid-body protein-protein docking using electrostatics and desolvation scoring. Bioinformatics (Oxford, England), 29(13), 1698–1699. https://doi.org/10.1093/BIOINFORMATICS/BTT262
Ghanbarpour, A., Jiang, M., Foster, D., & Chai, Q. (2023). Structure-free antibody paratope similarity prediction for in silico epitope binning via protein language models. iScience. https://doi.org/10.1016/J.ISCI.2023.106036
Wong, W. K., Robinson, S. A., Bujotzek, A., Georges, G., Lewis, A. P., Shi, J., & Deane, C. M. (2021). Ab-Ligity: Identifying sequence-dissimilar antibodies that bind to the same epitope. MAbs. https://doi.org/10.1080/19420862.2021.1873478
Ruffolo, J. A., Chu, L. S., Mahajan, S. P., & Gray, J. J. (2023). Fast, accurate antibody structure prediction from deep learning on massive set of natural antibodies. Nature Communications. https://doi.org/10.1038/S41467-023-38063-X
Akbar, R., Robert, P. A., Pavlović, M., Jeliazkov, J. R., Snapkov, I., Slabodkin, A., & Greiff, V. (2021). A compact vocabulary of paratope-epitope interactions enables predictability of antibody-antigen binding. Cell Reports, 34(11), 108856. https://doi.org/10.1016/j.celrep.2021.108856
Li, T., Pantazes, R. J., & Maranas, C. D. (2014). OptMAVEn – A new framework for the de novo design of antibody variable region models targeting specific antigen epitopes. PLoS ONE, 9(8), e105954. https://doi.org/10.1371/JOURNAL.PONE.0105954
Poosarla, V. G., Li, T., Goh, B. C., Schulten, K., Wood, T. K., & Maranas, C. D. (2017). Computational de novo design of antibodies binding to a peptide with high affinity. Biotechnology and Bioengineering, 114(6), 1331–1342. https://doi.org/10.1002/BIT.26244
Pantazes, R. J., & Maranas, C. D. (2010). OptCDR: A general computational method for the design of antibody complementarity determining regions for targeted epitope binding. Protein Engineering, Design and Selection, 23(11), 849–858. https://doi.org/10.1093/PROTEIN/GZQ061
Lapidoth, G. D., Baran, D., Pszolla, G. M., Norn, C., Alon, A., Tyka, M. D., & Fleishman, S. J. (2015). AbDesign: An algorithm for combinatorial backbone design guided by natural conformations and sequences. Proteins: Structure, Function, and Bioinformatics, 83(8), 1385–1406. https://doi.org/10.1002/PROT.24779
Adolf-Bryfogle, J., Kalyuzhniy, O., Kubitz, M., Weitzner, B. D., Hu, X., Adachi, Y., & Dunbrack, R. L. (2018). RosettaAntibodyDesign (RAbD): A general framework for computational antibody design. PLOS Computational Biology, 14(4), e1006112. https://doi.org/10.1371/JOURNAL.PCBI.1006112
Kuroda, D., Shirai, H., Jacobson, M. P., & Nakamura, H. (2012). Computer-aided antibody design. Protein Engineering, Design and Selection, 25(10), 507–521. https://doi.org/10.1093/protein/gzs024
Liu, X., Taylor, R. D., Griffin, L., Coker, S. F., Adams, R., Ceska, T., & Baker, T. (2017). Computational design of an epitope-specific Keap1 binding antibody using hotspot residues grafting and CDR loop swapping. Scientific Reports, 7(1), 1–11. https://doi.org/10.1038/srep41306
Moroncini, G., Kanu, N., Solforosi, L., Abalos, G., Telling, G. C., Head, M., & Williamson, R. A. (2004). Motif-grafted antibodies containing the replicative interface of cellular PrP are specific for PrPSc. Proceedings of the National Academy of Sciences of the United States of America, 101(28), 10404–10409. https://doi.org/10.1073/pnas.0403522101
Robinson, L. N., Tharakaraman, K., Rowley, K. J., Costa, V. V., Chan, K. R., Wong, Y. H., & Sasisekharan, R. (2015). Structure-guided design of an anti-dengue antibody directed to a non-immunodominant epitope. Cell, 162(3), 493–504. https://doi.org/10.1016/J.CELL.2015.06.057
Barthelemy, P. A., Raab, H., Appleton, B. A., Bond, C. J., Wu, P., Wiesmann, C., & Sidhu, S. S. (2008). Comprehensive analysis of the factors contributing to the stability and solubility of autonomous human VH domains. Journal of Biological Chemistry, 283(6), 3639–3654. https://doi.org/10.1074/jbc.M708536200
Ladiwala, A. R. A., Bhattacharya, M., Perchiacca, J. M., Cao, P., Raleigh, D. P., Abedini, A., & Tessier, P. M. (2012). Rational design of potent domain antibody inhibitors of amyloid fibril assembly. Proceedings of the National Academy of Sciences of the United States of America, 109(49), 19965–19970. https://doi.org/10.1073/pnas.1208797109
Ruffolo, J. A., Sulam, J., & Gray, J. J. (2022). Antibody structure prediction using interpretable deep learning. Patterns. https://doi.org/10.1016/J.PATTER.2021.100406
Akbar, R., Robert, P. A., Weber, C. R., Widrich, M., Frank, R., Pavlović, M., & Greiff, V. (2022). In silico proof of principle of machine learning-based antibody design at unconstrained scale. MAbs. https://doi.org/10.1080/19420862.2022.2031482
Greiff, V., Menzel, U., Miho, E., Weber, C., Riedel, R., Cook, S., & Reddy, S. T. (2017). Systems analysis reveals high genetic and antigen-driven predetermination of antibody repertoires throughout B cell development. Cell Reports, 19(7), 1467–1478. https://doi.org/10.1016/J.CELREP.2017.04.054
Khan, A., Cowen-Rivers, A. I., Grosnit, A., Deik, D. G. X., Robert, P. A., Greiff, V., & Bou-Ammar, H. (2023). Toward real-world automated antibody design with combinatorial Bayesian optimization. Cell Reports Methods. https://doi.org/10.1016/J.CRMETH.2022.100374
Nimrod, G., Fischman, S., Austin, M., Herman, A., Keyes, F., Leiderman, O., & Ofran, Y. (2018). Computational design of epitope-specific functional antibodies. Cell Reports, 25(8), 2121-2131.e5. https://doi.org/10.1016/J.CELREP.2018.10.081
Acknowledgements
The author would like to thank Peter Olson and Janet Stewart for reviewing of this article and providing detailed suggestions.
Funding
This work was partially supported by RevivAb Educational Advancement Grant MT-021.
Author information
Authors and Affiliations
Contributions
Single author—Not applicable.
Corresponding author
Ethics declarations
Conflict of interest
The author declares no conflict of interest.
Ethical Approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gallo, E. Revolutionizing Synthetic Antibody Design: Harnessing Artificial Intelligence and Deep Sequencing Big Data for Unprecedented Advances. Mol Biotechnol (2024). https://doi.org/10.1007/s12033-024-01064-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12033-024-01064-2