Abstract
Brucellosis is one of the neglected endemic zoonoses in the world. Vaccination appears to be a promising health strategy to prevent it. This study used advanced computational techniques to develop a potent multi-epitope vaccine for human brucellosis. Seven epitopes from four main brucella species that infect humans were selected. They had significant potential to induce cellular and humoral responses. They showed high antigenic ability without the allergenic characteristic. In order to improve its immunogenicity, suitable adjuvants were also added to the structure of the vaccine. The physicochemical and immunological properties of the vaccine were evaluated. Then its two and three-dimensional structure was predicted. The vaccine was docked with toll-like receptor4 to assess its ability to stimulate innate immune responses. For successful expression of the vaccine protein in Escherichia coli, in silico cloning, codon optimization, and mRNA stability were evaluated. The immune simulation was performed to reveal the immune response profile of the vaccine after injection. The designed vaccine showed the high ability to induce immune response, especially cellular responses to human brucellosis. It showed the appropriate physicochemical properties, a high-quality structure, and a high potential for expression in a prokaryotic system.
Graphical Abstract

Similar content being viewed by others
Introduction
Brucellosis also known as Malta fever is a worldwide zoonotic disease. According to the World Health Organization (WHO), it is one of the most neglected endemic zoonoses in the world [1]. It is one of the five zoonotic infections caused by gram-negative aerobic non-motile facultative intracellular coccobacillus brucella spp. Four species are the main causes of infection in humans including B. abortus, B. canis, B. melitensis, and B. suis. Among them, B. melitensis is the main cause of human chronic brucellosis [2,3,4]. It is transmitted to humans by consumption of unpasteurized foods and dairy products, inhalation of infected aerosols, or occupational contact [5]. The disease causes clinical manifestations, including fever, weakness, malaise and weight loss, lymphadenopathy, arthralgia/arthritis, hepatosplenomegaly, and hearing loss [6]. Currently, there are no brucella vaccines for humans, and the designed vaccines for animals show many side effects, such as infection. In addition, since killed or attenuated bacterial vaccines have various side effects, the use of subunit vaccines will be more effective than a whole bacteria cell. Subunit vaccines are usually developed using laboratory techniques such as cloning and expression of immune-stimulating antigens in vitro and the ability to stimulate the immune system of living organisms [2]. These experimental methods are often time-consuming and highly expensive. Designing vaccines with bioinformatic approaches is a very useful way because it can identify effective epitopes and suggest more efficient vaccines [7]. Several candidate vaccines have been reported based on bioinformatics approaches such as the effective vaccines for human papillomavirus (HPV) [8, 9], Ebola [10, 11], Zika [12, 13], SARS-COV-2 [7, 14,15,16], Helicobacter pylori [17], and several types of cancer [18,19,20,21,22]. There were a few reports of bioinformatics vaccines against brucellosis [23,24,25,26,27,28,29]. They contained multiple cytotoxic T lymphocytes (CTL) and B cell epitopes against several proteins of the bacteria. However, these vaccines did not cover most of the predominant immune proteins or they were not immunogenic against all pathogenic strains of brucellosis. In addition, some of these studies did not use all bioinformatics approaches to design the vaccine. Therefore, we designed a multi-epitope peptide-based vaccine against brucellosis to overcome these limitations. To achieve this goal, we first selected all immunogenic antigens of B. melitensis, the main cause of human brucellosis. Then, the final dominant immune epitopes were selected by aligning the sequence of the selected epitopes with all the antigenic epitopes of other human pathogenic brucella species. Evaluating the physicochemical, immunological, and structural quality of the vaccine was performed. In addition, the ability of the vaccine for inducing human immune system was assayed by molecular docking, molecular dynamics (MD) stimulation, and immune simulation. Although some identified epitopes in this study are overlapping with those reported by Saadi et al. [27], the quality of the three-dimensional (3D) vaccine structure improved.
To design an efficient vaccine, nine antigens of B. melitensis were evaluated. Seven antigens from them that had MHC-I, MHC-II, and B cell epitopes were chosen. Overlap T and B epitopes of these antigens were selected and aligned with the other high-risk brucella species in humans. Then, based on the present T and B cells epitopes in these antigens, a multi-epitope vaccine against brucella was predicted. Agonist of Toll-like receptor4 (TLR4), three segments of TTRFC, and the Pan HLA DR-binding epitope (PADRE) sequence were added to the structure. To verify the availability of the vaccine, the secondary and 3D structure, physicochemical properties, antigenicity, and allergenicity of the vaccine construct were analyzed by various bioinformatics software. Then, the interaction effect of the candidate vaccine and TLR4 was analyzed. Gene expression and in silico cloning of the vaccine were evaluated. Finally, the role of the candidate vaccine in activating the human immune response after injection was evaluated. The results indicated that the final protein structure could be used as a candidate multi-epitope peptide vaccine against human brucellosis. It is expected to be capable to induce potent immune responses against brucella antigens.
Materials and Methods
Selection of Antigens and Retrieval of Protein Sequences
The protein antigens of the B. melitensis strain were accessed by electronic databases Google Scholar, PubMed, EMBASE, Scopus, and Web of Science and retrieved from the NCBI database (https://www.ncbi.nlm.nih.gov/) (Table1).
Prediction of T Cell Epitopes and Continuous B Cell Epitopes
The prediction of CTL epitopes was done by an online server CTL Pred (http://crdd.osdd.net/raghava/ctlpred/) with an accuracy of 75.8%, and the IEDB database (www.iedb.org) through IEDB recommended 2.22 method and NetMHC pan 4.0 online server (http://www.cbs.dtu.dk/services/NetMHC/) with an accuracy of 90%. Nine residue-long HTL epitopes were recognized using these servers [30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49]. In addition, the IEDB database (IEDB recommended 2.22 method) (www.iedb.org) and RANKPEP online server (http://imed.med.ucm.es/Tools/rankpep.html) (with an accuracy of 60.0%) were employed to screen helper T cell (HTL) epitopes. These servers predicted eleven amino acid residue-length HTL epitopes [50,51,52,53]. As well as, the prediction of linear B cell epitope was done using methods of Emini surface accessibility and Linear Epitope Prediction 2.0 [54, 55].
Prediction of Interferon-Gamma (IFN-γ) Inducing Epitopes
The overlapped epitopes from previous steps were chosen to investigate whether they can induce IFN-γ immune response by the IFN-gamma epitope server (http://crdd.osdd.net/raghava/ifnepitope/scan.php). The accuracy of this server is estimated at 82.2%. Finally, the epitopes with positive results for the IFN-γ response were selected for the next steps [56].
Evaluating of Antigenicity, Allergenicity, and Toxicity of the Predicted Epitopes
Used epitopes in the designated vaccine must have antigenic properties. Antigenic properties of the predicted epitopes were evaluated by VaxiJen v2.0 (http://www.ddg-pharmfac.net/vaxijen/VaxiJen/VaxiJen.html) server with accuracy of 70% to 89%. Bacteria was selected as the target organism and only probable antigen epitopes which have a threshold value of ≥ 0.4 were selected for the next step [57, 58]. In addition, the allergenic characteristic of each epitope was evaluated by Allergen FP v.1.0 (http://ddg-pharmfac.net/AllergenFP/). The toxic probability of each epitope was assayed by the ToxinPred server (https://webs.iiitd.edu.in/raghava/toxinpred/multi_submit.php) with an accuracy of 94.50% [59,60,61]. Finally, non-allergen and non-toxic epitopes were chosen for the final vaccine construct.
Alignment of Selected Epitopes Between B. melitensis Strain and Other Species of Brucella
To understand the conservation of selected epitopes in high-risk brucella strains, the alignment of the selected epitopes in antigens of B. melitensis and the same antigens in other Brucella species (B. abortus, B. canis, B. suis) was performed. The amino acid sequences of proteins of other species were retrieved from the NCBI (https://www.ncbi.nlm.nih.gov/) database and aligned with the selected epitopes using the CLC sequence viewer [62].
Mapping of Multi-epitope Peptide Vaccine Candidate and Evaluating of the Physiochemical Properties, Solubility Prediction, Antigenicity, and Allergenicity
According to the results of the previous steps, the seven epitopes from each protein of immunogenic 39 kDa, lumazine synthase, superoxide dismutase, ribosomal protein L7/L12, BP26, outer membrane protein (OMP) -31, OMP-16, and OMP-28 were selected to be incorporated in the vaccine construction. The whole construct was designed using joining these epitopes to three adjuvant sequences of Heparin Binding Hemagglutinin (HBHA), PADRE, and the identified epitopes from the Tetanus toxin fragment C (TTFrC). Various segments of the designed vaccine were connected by proper linkers of EAAAK, KK, and GPGPG. After designing the vaccine, its physicochemical properties were computed using Expasy'sProtParam at http://us.expasy.org/tools/protparam.html [63]. The antigenicity of the final vaccine construct was predicted by Vaxijen v2.0 (http://www.ddg-pharmfac.net/vaxijen/VaxiJen/VaxiJen.html) and ANTIGENpro (http://scratch.proteomics.ics.uci.edu/) servers. In addition, the allergenicity of the vaccine construct was evaluated using the AllergenFP v.1.0 (http://ddg-pharmfac.net/AllergenFP/) and AllerTOP v. 2.0 (https://www.ddg-pharmfac.net/AllerTOP/) servers with accuracies of 0.879, and 0.828, respectively. The propensity of protein solubility upon overexpression in E. coli was assayed using SOLpro (http://scratch.proteomics.ics.uci.edu/) with an accuracy of 74.15.
Prediction, and Refinement of the Vaccine Secondary and 3D Structures
The SOMPA online analysis server (https://npsa-prabi.ibcp.fr/cgibin/npsa_automat.pl?) was used for the prediction of the protein secondary structure. This includes the distribution and proportion of three secondary structures of the alpha helix, extended strand, beta turn, and random coil. I-TASSER (https://zhanggroup.org/I-TASSER/) was used for the prediction of the protein 3D structure (https://npsa-prabi.ibcp.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_gor4.html) [64,65,66]. The amino acid sequence of the protein was introduced into this server to construct and analyze the 3D structure of the protein. Then, The 3D structure of the protein was refined by the Galaxy Refine server (http://galaxy.seoklab.org/cgi-bin/submit.cgi?type=REFINE) [67].
Validation of 3D Structure and Prediction of Discontinuous Bcell Epitopes
The 3D structure of the vaccine was validated by ERRAT (https://saves.mbi.ucla.edu/) and Zlab (https://zlab.umassmed.edu/bu/rama/index.pl) [68, 69]. Then discontinuous epitopes of the protein were predicted using the ElliPro server (http://tools.iedb.org/ellipro/). Discontinuous B cell epitopes are the most exposed epitopes that are recognized by antibodies [70].
Evaluating of the Vaccine Interaction with Human Toll-Like Receptor4
Initially, the 3D structure of the human toll-like receptor 4(hTLR4) was obtained from the PDB database (www.rcsb.org) with a code of 3FXI. Next, protein–protein docking of the vaccine structure (as a ligand) and hTLR4 (as a receptor) was performed by HDOCK online server (http://hdock.phys.hust.edu.cn/) [71]. Finally, the complex model had been visualized via Discovery studio 4.5 software [72].The molecular dynamics simulation was performed to evaluate the stability of docking complexes. The iMODS (http://imods.chaconlab.org/) web server calculated the B-factor (a disorder of atoms), structural deformability, and eigenvalue of the vaccine- hTLR4 complex.
Codon Optimization and In Silico Cloning
The efficiency of the multi-epitope vaccine construct in cloning and expression in E. coli as a host is an important step in vaccine designing. For codon optimization, the final amino acid sequence of multi-epitope vaccine was submitted to the Sequence Manipulation Suite (https://www.bioinformatics.org/sms2/rev_trans.html) and its DNA sequence was submitted in the GenScript online server (https://www.genscript.com/tools/rare-codon-analysis). The output of this server provided the codon adaptation index (CAI), GC content, and codon frequency and distribution (CFD) of the optimized sequence. Next, RNA structure server (https://rna.urmc.rochester.edu/RNAstructureWeb/Servers/Predict1/ResultsPages/20220101.152332-6e24fd60/Results.html) predicted the mRNA secondary structure of vaccine thermodynamically and provides minimum free energy (ΔG Kcal/mol) for structure. The restriction sites of ClaI and EcoRV were, respectively, added to the N- and C-terminals of the vaccine DNA sequence using CLC Sequence viewer v8.0 (http://www.cacbio.com) to facilitate the cloning of the vaccine in the E. coli expression system using pBA D24 [73]. Finally, the restriction sites of BmgBI and BamHI were, respectively, added to the N- and C-terminals of the vaccine DNA sequence using CLC Sequence viewer v8.0 (http://www.cacbio.com) to facilitate the expression of the vaccine in the E. coli expression system by the PMT-puro vector.
Immune Simulation
The C-ImmSim server was used to characterize the profile of the stimulated immune response by the multi-epitope peptide vaccine [74]. To produce immune stimulation in humans, an injection was administered with 1000 vaccine molecules per injection for a period of four months. The parameters like a random seed, simulation volume, and simulation step were kept as 12,345, 10 µl, and 1050, respectively.
Results
Selection of Epitopes
Nine antigenic proteins were selected from B. melitensis to predict HLA-I binding epitopes (HLA-A, B, and C) using IEDB database and NetMHC 4 online server. CTL Pred server was used to predict the CTL epitopes of the proteins. RANKEP server and IEDB database predicted HLA-II binding epitopes (DP, DQ, and DR) from these proteins. Linear Bcell epitopes were predicted by the Emini surface accessibility Linear Epitope Prediction 2.0 in the IEDB database. All selected peptides that have the same regions in HLA-I, HLA-II, CTL, and linear B cell epitopes were submitted to the IFN epitope server for evaluating their ability to induce IFN-γ. The antigenicity of epitopes was assayed by VaxiJen v2.0. The allergenic and toxic probability of each epitope were evaluated by AllergenFP and ToxinPred servers. The alignment of the selected epitopes between B. melitensis, B. abortus, B. canis, and B. suis species showed the selected epitopes were conserved between all species. The seven peptides in the vaccine structure were selected based on our selected criteria from proteins of immunogenic 39-kDa, Superoxide dismutase, Ribosomal protein L7/L12, BP26, OMP-31, OMP-16, and OMP-28. The proteins of lumazine synthase and OMP-25 have no efficient peptides. The selected peptides had the highest ability in IFN-γ induction, low allergenicity, and potent antigenic ability and were conserved among the most important pathogenic brucella species (Table 2 and Supplementary Fig. 1).
Vaccine Engineering, Evaluating of Physicochemical Properties, Antigenicity, and Allergenicity
To increase the immunogenicity of the vaccine, adjuvants of HBHA, PADRE, and identified epitopes from TTFrC were joined to the selected peptides from the seven brucella proteins (Fig. 1A). Results showed the designed human brucella vaccine has physicochemical and immunological properties. According to the results of the ProtParam server, the vaccine was stable with an instability index of 29.69 (the stable molecules have an instability index of less than 30). It hada molecular weight of 54.138 KDa and 500 amino acids lenth. In addition, it had an isoelectric point of 7.19, a gravy of − 0.465, and an aliphatic index of 79.52. The vaccine half-life was 30 h (h) in mammalian reticulocytes, 20 h in yeast, and 10 h in E. coli. Based on the prediction of the Solpro server, the vaccine was soluble upon expression in E. coli with a Solpro index of 0.955871. To evaluate the antigenicity of the vaccine by vaxijen server, the “bacteria” option was chosen as a target organism. The threshold for antigenicity was chosen 0.5. The result showed this protein showed appropriate antigenicity (the index equal to or more than 0.5 shows the antigenicity for a protein). In addition, an assessment of antigenicity by Antigen pro showed this vaccine was an antigenic protein with an antigenicity of 0.833277. AllergenFP v.1.0 and AllerTOP v. 2.0 showed the vaccine was non-allergen.
A Schematic representation of the designed multi-epitope peptide vaccine against human Brucellosis. The vaccine consists of twenty-six parts: Epitopes from antigenic proteins of immunogenic 39-kDa, lumazine synthase, superoxide dismutase, ribosomal protein L7/L12, BP26, OMP-16, and OMP-28, adjuvants of HBHA, PADRE, and identified epitopes from TTFrC which join to each other by linkers of KK, EAAAK, and GPGPG. B The secondary structure of the modeled multi-epitope vaccine was predicted by SOPMA server, C The 3D structure of the designed vaccine was predicted via homology modeling by I-Tasser, then the best-predicted model was refined by GalaxyRefine and visualized using Pymol. HBHA is shown in green, TTFrC segments in cyan, predicted epitopes in purple, PADRE in pink, EAAAK linkers in blue, KK linkers in gray, and GPGPG linkers are shown in yellow
Secondary and 3D Structure Modeling, Refinement, and Validation and Prediction of Discontinuous B Cell Epitopes
The secondary structure of the vaccine was predicted using SOPMA online server. From the 500 amino acids of the protein, 229 (45.8%) were alpha helix, 95 (19%) were extended strands, 31 (6.2%) were beta turn, and 145 amino acids (29%) composed the structure of the coils (Fig. 1B). Then I-Tasser predicted the primary 3D model of the human brucellosis vaccine (Supplementary Fig. 2). Next, the selected I-Taser model was refined by Galaxy Refine software (Fig. 1C). The quality of the designed vaccine was evaluated using the Ramachandran plot in the Zlab server and the characteristic atomic interaction in the ERRAT server. The Ramachandran results of the final model showed that the majority of residues (96.487%) are located in the favored region and 2.81% are allowed, and only 0.703% of residues are the outlier. The ERRAT result showed that the refined model had an ERRAT score of 90.890. The results of the Ramachandran plot and ERRAT indicated that the refined 3D structure had the appropriate structure and therefore, could be utilized as a reliable model for further evaluations (Supplementary Fig. 2). The prediction of discontinuous B cell epitopes by Ellipro server revealed the presence of 262 total residues in the vaccine structure that had the potential of discontinuous B cell epitopes. Their scores were varying from 0.513 to 0.788. The sequences of selected epitopes from lumazine synthase, superoxide dismutase, BP26, OMP-16, and OMP-28 were the same in the linear and discontinuous B cells epitopes of the vaccine (Table 3 and Fig. 2).
Discontinuous B cell epitopes predicted by the ElliPro are represented in Table 3. The epitopes are represented by yellow surface, and the bulk of the protein is represented in grey stick
Molecular Docking and MD Simulation of the Vaccine Structure with TLR4 Receptor and Refinement of the Interaction
H-Dock server predicted several models for the interaction complex of vaccine and TLR4. Among predicted models, model 1 was selected as the best-docked complex with the lowest energy score (− 290.25) (Fig. 3). In this complex model, the HBHA segment in the vaccine interacted with TLR4, and the myeloid differentiation factor 2 (MD-2). Molecule stability and physical movements of the vaccine-receptor docked complex were evaluated by the iMODS server. The deformability graph was illustrated in Fig. 4A, hinges were highlighting the deformability regions in the complex. It showed some deflection in the 0 to 1°A range. The B-factor calculates the root mean square value and represents the uncertainty of each atom in the docking complex (Fig. 4B). The eigenvalues of the vaccine and TLR4 docking complex were 2.042744 × 10–5(Fig. 4C). The variance matrix graph of residues displayed in Fig. 4D. The covariance matrix showed a correlation between pairs of residue experience (correlated (red), uncorrelated (white), and anti-correlated (blue)) (Fig. 4E). The elastic network model indicated the connection between atoms and springs. Therefore, stiffer springs were shown in the darker regions. (Fig. 4F). The results of these graphs approved that the amino acids in the complex of vaccine and TLR4 had stable connections.
Docking model (cartoon representation) of human TLR4 in complex with the vaccine obtained using HDOCK server. As it shows HBHA interacted with 4 and MD-2.To more visualized interaction points, some of the interacting residues of the vaccine and TLR4 are magnified. The docked model has been visualized via Discovery studio 4.5 software
In Silico Cloning, Vaccine Optimization, and Prediction of the mRNA Secondary Structure
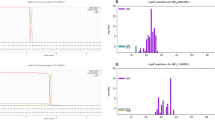
The reverse translation of the protein vaccine into a nucleotide sequence was performed simultaneously using the Sequence Manipulation Suite server to express a high-level protein in E. coli. The gene expression in the E. coli as a host cell was evaluated using the Genscript server. The codon adaptation index (CAI) of the optimized sequences was 1.00, the GC content was 60.01%, and the codon frequency and distribution (CFD) was 0%. These results ensured the potential of the vaccine expression in E. coli. To clone the gene into E. coli, ClaI, and EcoRV restriction sites were added into the N- and C-terminals of the sequence using CLC Sequence viewer v.8. The vaccine gene was inserted into the pBAD24 vector (Fig. 5A). To express the gene into E. coli, BmgBI, and BamHI restriction sites were added into the N- and C-terminals of the sequence using CLC Sequence viewer v.8, and the vaccine gene was inserted into the pMT-puro vector (Fig. 5B). The minimum free energy of the vaccine was determined using RNAstructure server. The optimized mRNA construct showed a ΔG value of − 371.4 kcal/mol. Supplementary Fig. 3 illustrates the predicted secondary structure of vaccine mRNA.
A Cloning the gene into E. coli by ClaI, EcoRV restriction sites, and the pBAD24 vector and using CLC Sequence viewer v.8. The vaccine gene was inserted into the pBAD24 vector. B Expression of the gene into E. coli by BmgBI, and BamH1 restriction sites, pMT-puro vector, and using CLC Sequence viewer v.8
Immune Simulation by the Vaccine Structure
In silico simulation results suggested that the administration of the vaccine had the potential for inducing various immune responses. The humoral response was characterized by an increase in the levels of antibodies and B cell populations. T cell populations (CTL and Th cells) developed increased responses corresponding to the memory cells. The activity of macrophages, dendritic cells, and NK cells were observed consistently throughout the period time of 120 days. A significant increase in the levels of IFN-γ, IL-2, and TGF-β was also found at subsequent exposure. This signifies that the vaccine generated robust immune responses (Fig. 6).
Simulation of the immune responses to vaccination by human brucellosis vaccine. A Levels of immunoglobulins after incitement by a multiepitope vaccine. B Amount of memory B cells (y2) and immunoglobulin isotypes (IgM, IgG1, IgG2) as the immune response. C Amount of active, duplicating, resting, and anergic B cells. D Plasma cell population per state. E Total and memory Th lymphocytes. F Amount of active, duplicating, resting, and anergic population of Th cells. G Amount of different types of Th cells. H Tc cell population per state. I Count of active, duplicating, resting, and anergic Tc cells. J Amount of NK cells. K Amount of DC cells. L Count of macrophage cells. M Count of eosinophil cells. N Produced cytokines
Discussion
Brucellosis is one of the world’s neglected zoonotic diseases. It occurs as a debilitating acute infection that can become chronic and has many complications. Control and vaccination of human brucellosis is the first challenge to eradicating this disease. However, none of the available vaccines are perfect and they may cause brucellosis in humans [75]. Therefore, we should develop a new vaccine to prevent this disease. Identification of immunogenic peptides is an essential step in vaccine design. With helping of bioinformatics approaches, we can recognize immunodominant T- and Bcell epitopes and design potential vaccines for the control of brucellosis. These methods are effective and can reduce the time and cost of experimental methods in the lab [7]. Therefore, in this study, we designed a bioinformatics vaccine against human brucellosis. The vaccine includes 500 amino acid lengths constructed from seven specific epitopes of antigenic proteins of B. melitensis. These epitopes could stimulate both cellular and humoral immunity. In addition, they had not allergenic and toxic properties and are conserved in four main infectious species of humans brucella including B. abortus, B. canis, B. melitensis, and B. suis. The amino acid sequences of three adjuvants of HBHA, PADRE, and three segments of TTFRC were added to the vaccine mapping of human brucella. HBHA is a virulence factor from Mycobacterium tuberculosis that has no systemic toxicity [76, 77]. It possesses a strong immunostimulatory potential such as inducing DC maturation in a TLR 4-dependent manner [77, 78]. PADRE and TTFRC fragments can induce T-helper cell responses and may be useful in the development of vaccines [19, 27, 79]. Therefore, these segments may represent powerful adjuvants for our vaccines. For designing of vaccine with reduced junctional immunogenicity, the linkers of KK and GPGPG were integrated between the epitopes and adjuvants. They increased the stability and quality of the 3D structure of the vaccine and induced neutral theoretical pI. The EAAAK linker was also inserted into the N and C-terminals of HBHA to reduce its interaction of it with other regions and cause more efficient separation [19, 22, 80]. The designed vaccine had high antigenic ability and a lack of allergenic function. An efficient vaccine should possess good physicochemical properties during production, formulation, storage, and consumption. According to the results of bioinformatic predictions, the designed vaccine was stable with a stability index of 29.69. It had a pI of 7.19 and a molecular weight of 54.138 kDa. Analyses of the secondary structure revealed that the vaccine contained mainly alpha helix (45.8%) and random coil (29%). The refinement of the 3D structure of the designed vaccine displayed appropriate characteristics based on the Ramachandran plots and ERRAT results. The result of the Ramachandran plot indicates that 96.487% of the residues are initiated in the favored regions, 2.81% are allowed and only 0.703% of residues are the outlier. In addition, the quality of the 3D structure was 90.890 based on the ERRAT results. The accessibility of a significant number of amino acids in the vaccine as B cell epitopes indicates the high ability of this structure for stimulation of the humoral immune response. For assessment of the ability of the designed vaccine to bind with TLRs on immune cells, the TLR4 was docked with the vaccine. The results showed that the HBHA segment of the vaccine can bind to TLR4 and MD-2. Nezafat et al. demonstrated that the binding of HBHA to TLR4 may be mediated by MD-2 [18]. The interface of the vaccine with TLR4 and MD-2 was signifying that the vaccine may produce both innate and adaptive immune responses. In addition, the iMODS server indicated the interaction was stable. In the production of vaccines, protein must be expressed in a suitable host. The codon optimization, in silico cloning, and evaluating mRNA stability showed that the vaccine can efficiently transcript and translate in E. coli.The immune stimulation showed results consistent with typical immune responses. Following the injection of the vaccine, there was a general increase in the generated immune responses. The development of memory Band Tcells were lasting for several months. Helper T cells were particularly stimulated. In addition, the population of DC, macrophage, and NK cells were increased periodically. Another interesting observation was the rising levels of IFN-γ and IL-2 after the first injection in twenty days period. This showed high levels of TH cells and consequently efficient Ig production. Although, rising in TGF-β in this period may be a little worrying. The next step is planning experimental studies for conforming to the immunogenicity and safety profile of the designed vaccine.
Conclusion
In the current study, using a variety of bioinformatics tools, a multi-epitope peptide vaccine against human brucellosis was designed. The results of this study showed that this vaccine could be possible as a candidate vaccine against all subspecies of brucella which infect humans. It stimulated the profile of immune response, especially humoral and cellular immunity. It had appropriate physicochemical properties, high-quality structure, and the ability for expression in E. coli as a host. However, these results need to be verified by experimental methods.
Data Availability
The data used or analyzed during the current study are available from the corresponding author on reasonable request.
References
Bosilkovski, M., Keramat, F., & Arapović, J. (2021). The current therapeutical strategies in human brucellosis. Infection, 49, 823–832.
Amjadi, O., Rafiei, A., Mardani, M., Zafari, P., & Zarifian, A. (2019). A review of the immunopathogenesis of Brucellosis. Infectious Diseases, 51, 321–333.
Buttigieg, S. C., Savic, S., Cauchi, D., Lautier, E., Canali, M., & Aragrande, M. (2018). Brucellosis control in Malta and Serbia: A one health evaluation. Frontiers in Veterinary Science, 5, 147.
Yagupsky, P. (1999). Detection of Brucellae in blood cultures. Journal of Clinical Microbiology, 37, 3437–3442.
Bossi, P., Tegnell, A., Baka, A., Van Loock, F., Hendriks, J., Werner, A., Maidhof, H., Gouvras, G., Biological, TFO & Chemical Agent THREATS, PHD, European Commission, Luxembourg. (2004). Bichat guidelines for the clinical management of brucellosis and bioterrorism-related brucellosis. Eurosurveillance Weekly, 9, E15–E16.
Kaygusuz, T. O., Kaygusuz, I., Kilic, S., Yalcin, S., & Felek, S. (2005). Investigation of hearing loss in patients with acute brucellosis by standard and high-frequency audiometry. Clinical Microbiology & Infection, 11, 559–563.
Yazdani, Z., Rafiei, A., Yazdani, M., & Valadan, R. (2020). Design an efficient multi-epitope peptide vaccine candidate against SARS-CoV-2: An in silico analysis. Infection and Drug Resistance, 13, 3007.
Bagheri, A., Nezafat, N., Eslami, M., Ghasemi, Y., & Negahdaripour, M. (2021). Designing a therapeutic and prophylactic candidate vaccine against human papillomavirus through vaccinomics approaches. Infection, Genetics and Evolution, 95, 105084.
Yazdani, Z., Rafiei, A., Valadan, R., Ashrafi, H., Pasandi, M., & Kardan, M. (2020). Designing a potent L1 protein-based HPV peptide vaccine: A bioinformatics approach. Computational Biology and Chemistry, 85, 107209.
Dash, R., Das, R., Junaid, M., Akash, M. F. C., Islam, A., & Hosen, S. Z. (2017). In silico-based vaccine design against Ebola virus glycoprotein. Advances and Applications in Bioinformatics and Chemistry, 10, 11.
Khan, M., Hossain, M., Rakib-Uz-Zaman, S., & Morshed, M. (2015). Epitope-based peptide vaccine design and target site depiction against Ebola viruses: An immunoinformatics study. Scandinavian Journal of Immunology, 82, 25–34.
Dey, S., Nandy, A., Basak, S. C., Nandy, P., & Das, S. (2017). A bioinformatics approach to designing a Zika virus vaccine. Computational Biology and Chemistry, 68, 143–152.
Weltman, J. (2016). An immuno-bioinformatic analysis of Zika virus (ZIKV) envelope E protein. Journal of Medical Microbiology & Diagnosis, 5(2161–0703), 1000228.
Dawood, R. M., El-Meguid, M. A., Salum, G. M., El-Wakeel, K., Shemis, M., & El Awady, M. K. (2021). Bioinformatics prediction of B and T cell epitopes within the spike and nucleocapsid proteins of SARS-CoV2. Journal of Infection and Public Health, 14, 169–178.
Noorimotlagh, Z., Karami, C., Mirzaee, S. A., Kaffashian, M., Mami, S., & Azizi, M. (2020). Immune and bioinformatics identification of T cell and B cell epitopes in the protein structure of SARS-CoV-2: A systematic review. International Immunopharmacology., 86, 106738.
Singh, A., Thakur, M., Sharma, L. K., & Chandra, K. (2020). Designing a multi-epitope peptide based vaccine against SARS-CoV-2. Science and Reports, 10, 1–12.
Mohammad, N., Karsabet, M. T., Amani, J., Ardjmand, A., Zadeh, M. R., Gholi, M. K., Saffari, M., & Ghasemi, A. (2016). In silico design of a chimeric protein containing antigenic fragments of Helicobacter pylori; a bioinformatic approach. The Open Microbiology Journal, 10, 97.
Atapour, A., Negahdaripour, M., Ghasemi, Y., Razmjuee, D., Savardashtaki, A., Mousavi, S. M., Hashemi, S. A., Aliabadi, A., & Nezafat, N. (2020). In silico designing a candidate vaccine against breast cancer. International Journal of Peptide Research and Therapeutics, 26, 369–380.
Nezafat, N., Ghasemi, Y., Javadi, G., Khoshnoud, M. J., & Omidinia, E. (2014). A novel multi-epitope peptide vaccine against cancer: An in silico approach. Journal of Theoretical Biology, 349, 121–134.
Safavi, A., Kefayat, A., Abiri, A., Mahdevar, E., Behnia, A. H., & Ghahremani, F. (2019). In silico analysis of transmembrane protein 31 (TMEM31) antigen to design novel multiepitope peptide and DNA cancer vaccines against melanoma. Molecular Immunology, 112, 93–102.
Safavi, A., Kefayat, A., Sotoodehnejadnematalahi, F., Salehi, M., & Modarressi, M. H. (2019). Production, purification, and in vivo evaluation of a novel multiepitope peptide vaccine consisted of immunodominant epitopes of SYCP1 and ACRBP antigens as a prophylactic melanoma vaccine. International Immunopharmacology, 76, 105872.
Yazdani, Z., Rafiei, A., Irannejad, H., Yazdani, M., & Valadan, R. (2020). Designing a novel multiepitope peptide vaccine against melanoma using immunoinformatics approach. Journal of Biomolecular Structure & Dynamics, 40, 1–13.
Chen, Z., Zhu, Y., Sha, T., Li, Z., Li, Y., Zhang, F., & Ding, J. (2021). Design of a new Multi-Epitope vaccine against Brucella based on T and B cell epitopes using bioinformatics methods. Epidemiology and Infection, 149, 1–41.
Li, M., Zhu, Y., Niu, C., Xie, X., Haimiti, G., Guo, W., Yu, M., Chen, Z., Ding, J., & Zhang, F. (2022). Design of a multi-epitope vaccine candidate against Brucella melitensis. Science and Reports, 12, 10146.
Li, Z., Zhang, F., Zhang, C., Wang, C., Lu, P., Zhao, X., Hao, L., & Ding, J. (2019). Immunoinformatics prediction of OMP2b and BCSP31 for designing multi-epitope vaccine against Brucella. Molecular Immunology, 114, 651–660.
Rezaei, M., Rabbani-Khorasgani, M., Zarkesh-Esfahani, S. H., Emamzadeh, R., & Abtahi, H. (2019). Prediction of the Omp16 epitopes for the development of an epitope-based vaccine against Brucellosis. Infectious Disorders - Drug Targets, 19, 36–45.
Saadi, M., Karkhah, A., & Nouri, H. R. (2017). Development of a multi-epitope peptide vaccine inducing robust T cell responses against brucellosis using immunoinformatics based approaches. Infection, Genetics and Evolution, 51, 227–234.
Sha, T., Li, Z., Zhang, C., Zhao, X., Chen, Z., Zhang, F., & Ding, J. (2020). Bioinformatics analysis of candidate proteins Omp2b, P39 and BLS for Brucella multivalent epitope vaccines. Microbial Pathogenesis, 147, 104318.
Yin, D., Li, L., Song, D., Liu, Y., Ju, W., Song, X., Wang, J., Pang, B., Xu, K., & Li, J. (2016). A novel recombinant multi-epitope protein against Brucella melitensis infection. Immunology Letters, 175, 1–7.
Andreatta, M., & Nielsen, M. (2016). Gapped sequence alignment using artificial neural networks: Application to the MHC class I system. Bioinformatics, 32, 511–517.
Bhasin, M., & Raghava, G. P. (2004). Prediction of CTL epitopes using QM, SVM and ANN techniques. Vaccine, 22, 3195–3204.
Chen, C.-Y., Pollack, S., Hunter, D. J., Hirschhorn, J. N., Kraft, P., & Price, A. L. (2013). Improved ancestry inference using weights from external reference panels. Bioinformatics, 29, 1399–1406.
Giguère, S., Drouin, A., Lacoste, A., Marchand, M., Corbeil, J., & Laviolette, F. (2013). MHC-NP: Predicting peptides naturally processed by the MHC. Journal of Immunological Methods, 400, 30–36.
Hoof, I., Peters, B., Sidney, J., Pedersen, L. E., Sette, A., Lund, O., Buus, S., & Nielsen, M. (2009). NetMHCpan, a method for MHC class I binding prediction beyond humans. Immunogenetics, 61, 1–13.
Jurtz, V., Paul, S., Andreatta, M., Marcatili, P., Peters, B., & Nielsen, M. (2017). NetMHCpan-4.0: Improved peptide–MHC class I interaction predictions integrating eluted ligand and peptide binding affinity data. The Journal of Immunology, 199, 3360–3368.
Karosiene, E., Lundegaard, C., Lund, O., & Nielsen, M. (2012). NetMHCcons: A consensus method for the major histocompatibility complex class I predictions. Immunogenetics, 64, 177–186.
Kim, Y., Sidney, J., Pinilla, C., Sette, A., & Peters, B. (2009). Derivation of an amino acid similarity matrix for peptide: MHC binding and its application as a Bayesian prior. BMC Bioinformatics, 10, 1–11.
Lundegaard, C., Lamberth, K., Harndahl, M., Buus, S., Lund, O., & Nielsen, M. (2008). NetMHC-3.0: Accurate web accessible predictions of human, mouse and monkey MHC class I affinities for peptides of length 8–11. Nucleic Acids Research, 36, W509–W512.
Lundegaard, C., Nielsen, M., & Lund, O. (2006). The validity of predicted T-cell epitopes. Trends in Biotechnology, 24, 537–538.
Moutaftsi, M., Peters, B., Pasquetto, V., Tscharke, D. C., Sidney, J., Bui, H.-H., Grey, H., & Sette, A. (2006). A consensus epitope prediction approach identifies the breadth of murine T CD8+-cell responses to vaccinia virus. Nature Biotechnology, 24, 817–819.
Nielsen, M., & Andreatta, M. (2016). NetMHCpan-3.0; improved prediction of binding to MHC class I molecules integrating information from multiple receptor and peptide length datasets. Genome Medicine, 8, 1–9.
Nielsen, M., Lundegaard, C., Blicher, T., Lamberth, K., Harndahl, M., Justesen, S., Røder, G., Peters, B., Sette, A., & Lund, O. (2007). NetMHCpan, a method for quantitative predictions of peptide binding to any HLA-A and-B locus protein of known sequence. PLoS ONE, 2, e796.
Nielsen, M., Lundegaard, C., Worning, P., Lauemøller, S. L., Lamberth, K., Buus, S., Brunak, S., & Lund, O. (2003). Reliable prediction of T-cell epitopes using neural networks with novel sequence representations. Protein Science, 12, 1007–1017.
Peters, B., & Sette, A. (2005). Generating quantitative models describing the sequence specificity of biological processes with the stabilized matrix method. BMC Bioinformatics, 6, 1–9.
Rasmussen, M., Fenoy, E., Harndahl, M., Kristensen, A. B., Nielsen, I. K., Nielsen, M., & Buus, S. (2016). Pan-specific prediction of peptide–MHC class I complex stability, a correlate of T cell immunogenicity. The Journal of Immunology, 197, 1517–1524.
Sidney, J., Assarsson, E., Moore, C., Ngo, S., Pinilla, C., Sette, A., & Peters, B. (2008). Quantitative peptide binding motifs for 19 human and mouse MHC class I molecules derived using positional scanning combinatorial peptide libraries. Immunome Research, 4, 1–14.
Vita, R., Mahajan, S., Overton, J. A., Dhanda, S. K., Martini, S., Cantrell, J. R., Wheeler, D. K., Sette, A., & Peters, B. (2019). The immune epitope database (IEDB): 2018 update. Nucleic Acids Research, 47, D339–D343.
Zhang, H., Lund, O., & Nielsen, M. (2009). The PickPocket method for predicting binding specificities for receptors based on receptor pocket similarities: Application to MHC-peptide binding. Bioinformatics, 25, 1293–1299.
Zhang, Q., Wang, P., Kim, Y., Haste-Andersen, P., Beaver, J., Bourne, P. E., Bui, H.-H., Buus, S., Frankild, S., & Greenbaum, J. (2008). Immune epitope database analysis resource (IEDB-AR). Nucleic Acids Research, 36, W513–W518.
Andreatta, M., Karosiene, E., Rasmussen, M., Stryhn, A., Buus, S., & Nielsen, M. (2015). Accurate pan-specific prediction of peptide-MHC class II binding affinity with improved binding core identification. Immunogenetics, 67, 641–650.
Nielsen, M., & Lund, O. (2009). NN-align. An artificial neural network-based alignment algorithm for MHC class II peptide binding prediction. BMC Bioinform, 10, 1–10.
Nielsen, M., Lundegaard, C., & Lund, O. (2007). Prediction of MHC class II binding affinity using SMM-align, a novel stabilization matrix alignment method. BMC Bioinform, 8, 1–12.
Reche, P. A., Glutting, J.-P., Zhang, H., & Reinherz, E. L. (2004). Enhancement to the RANKPEP resource for the prediction of peptide binding to MHC molecules using profiles. Immunogenetics, 56, 405–419.
Emini, E. A., Hughes, J. V., Perlow, D., & Boger, J. (1985). Induction of hepatitis A virus-neutralizing antibody by a virus-specific synthetic peptide. Journal of Virology, 55, 836–839.
Jespersen, M. C., Peters, B., Nielsen, M., & Marcatili, P. (2017). BepiPred-2.0: Improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Research, 45, W24–W29.
Dhanda, S. K., Vir, P., & Raghava, G. P. (2013). Designing of interferon-gamma inducing MHC class-II binders. Biology Direct, 8, 1–15.
Doytchinova, I. A., & Flower, D. R. (2007). VaxiJen: A server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinformatics, 8, 1–7.
Zaharieva, N., Dimitrov, I., Flower, D., & Doytchinova, I. (2017). Immunogenicity prediction by VaxiJen: A ten year overview. The Journal of Proteomics & Bioinformatics, 10, 298–310.
Buus, S., Lauemøller, S., Worning, P., Kesmir, C., Frimurer, T., Corbet, S., Fomsgaard, A., Hilden, J., Holm, A., & Brunak, S. (2003). Sensitive quantitative predictions of peptide-MHC binding by a ‘Query by Committee’artificial neural network approach. Tissue Antigens, 62, 378–384.
Gupta, S., Kapoor, P., Chaudhary, K., Gautam, A., Kumar, R., Raghava, G. P. S., Consortium OSDD. (2013). In silico approach for predicting toxicity of peptides and proteins. PLoS ONE, 8, e73957.
Gupta, S., Kapoor, P., Chaudhary, K., Gautam, A., Kumar, R., Raghava, G. P., 2015. Peptide toxicity prediction. In Computational peptidology (pp. 143–157). Springer.
Sarwar, M. A., Rehman, A., & Ferzund, J. (2016). Database search, alignment viewer and genomics analysis tools: Big data for bioinformatics. International Journal of Computer Science, Information Technology, & Security, 14, 317.
Gasteiger, E., Hoogland, C., Gattiker, A., Wilkins, M. R., Appel, R. D., & Bairoch, A. (2005). Protein identification and analysis tools on the ExPASy server. In The proteomics protocols handbook (pp. 571–607).
Roy, A., Kucukural, A., & Zhang, Y. (2010). I-TASSER: A unified platform for automated protein structure and function prediction. Nature Protocols, 5, 725–738.
Yang, J., Yan, R., Roy, A., Xu, D., Poisson, J., & Zhang, Y. (2015). The I-TASSER Suite: Protein structure and function prediction. Nature Methods, 12, 7–8.
Yang, J., & Zhang, Y. (2015). I-TASSER server: New development for protein structure and function predictions. Nucleic Acids Research, 43, W174–W181.
Ko, J., Park, H., Heo, L., & Seok, C. (2012). GalaxyWEB server for protein structure prediction and refinement. Nucleic Acids Research, 40, W294–W297.
Anderson, R. J., Weng, Z., Campbell, R. K., & Jiang, X. (2005). Main-chain conformational tendencies of amino acids. Proteins, 60, 679–689.
Colovos, C., & Yeates, T. O. (1993). Verification of protein structures: Patterns of nonbonded atomic interactions. Protein Science, 2, 1511–1519.
Ponomarenko, J., Bui, H.-H., Li, W., Fusseder, N., Bourne, P. E., Sette, A., & Peters, B. (2008). ElliPro: A new structure-based tool for the prediction of antibody epitopes. BMC Bioinformatics, 9, 1–8.
Yan, Y., Tao, H., He, J., & Huang, S.-Y. (2020). The HDOCK server for integrated protein–protein docking. Nature Protocols, 15, 1829–1852.
Biovia, D. S. (2020). BIOVIA workbook, release 2017; BIOVIA pipeline pilot, release 2017. Dassault Systèmes.
Bio-Qiagen, C. (2016). CLC sequence viewer. Aarhus, Denmark.
Rapin, N., Lund, O., Bernaschi, M., & Castiglione, F. (2010). Computational immunology meets bioinformatics: The use of prediction tools for molecular binding in the simulation of the immune system. PLoS ONE, 5, e9862.
O’callaghan, D. (2020). Human brucellosis: Recent advances and future challenges. Infectious Diseases of Poverty, 9, 1–2.
Lei, Y., Shao, J., Ma, F., Lei, C., Chang, H., & Zhang, Y. (2020). Enhanced efficacy of a multi-epitope vaccine for type A and O foot-and-mouth disease virus by fusing multiple epitopes with Mycobacterium tuberculosis heparin-binding hemagglutinin (HBHA), a novel TLR4 agonist. Molecular Immunology, 121, 118–126.
Pethe, K., Alonso, S., Biet, F., Delogu, G., Brennan, M. J., Locht, C., & Menozzi, F. D. (2001). The heparin-binding haemagglutinin of M. tuberculosis is required for extrapulmonary dissemination. Nature, 412, 190–194.
Kumar, S., Sunagar, R., & Gosselin, E. (2019). Bacterial protein toll-like-receptor agonists: A novel perspective on vaccine adjuvants. Frontiers in Immunology, 10, 1144.
Alexander, J., Sidney, J., Southwood, S., Ruppert, J., Oseroff, C., Maewal, A., Snoke, K., Serra, H. M., Kubo, R. T., & Sette, A. (1994). Development of high potency universal DR-restricted helper epitopes by modification of high affinity DR-blocking peptides. Immunity, 1, 751–761.
Arai, R., Ueda, H., Kitayama, A., Kamiya, N., & Nagamune, T. (2001). Design of the linkers which effectively separate domains of a bifunctional fusion protein. Protein Engineering, 14, 529–532.
Funding
This work was supported by the Research and Technology Deputy of Mazandaran University of Medical Sciences [Grant Number 8477].
Author information
Authors and Affiliations
Contributions
AR conceived and designed this study; supervised the project and collected the data. ZY and MG contributed to performing the analysis. ZY wrote the draft. AR and SA edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
The protocol of this study was approved by the Research Ethics committee of Mazandaran University of Medical Sciences (IR.MAZUMS.REC.1399.632).
Consent for Publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yazdani, Z., Rafiei, A., Ghoreyshi, M. et al. In Silico Analysis of a Candidate Multi-epitope Peptide Vaccine Against Human Brucellosis. Mol Biotechnol 66, 769–783 (2024). https://doi.org/10.1007/s12033-023-00698-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-023-00698-y