Abstract
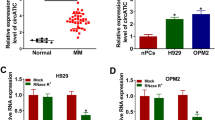
We focused on hsa_circ_0134426 (circ_0134426) to determine its functional effects and targets in multiple myeloma (MM) development. The relative expression of circ_0134426, miR-146b-3p, and neuron-derived neurotrophic factor (NDNF) in the MM samples was confirmed by quantitative real-time PCR (qPCR) or western blotting. The protein levels of epithelial-to-mesenchymal transition (EMT)-linked markers were identified using western blotting. Xenograft models in nude mice were used for in vivo functional analysis of circ_0134426. The binding of miR-146b-3p to circ_0134426 or NDNF was confirmed by dual-luciferase and RIP assays. Poor levels of circ_0134426 and NDNF and high levels of miR-146b-3p were observed in MM bone marrow samples and cell lines. Circ_0134426 overexpression blocked MM cell growth, colony formation, and migration and impeded tumor development in xenograft models. circRNA_0134426 decoys miR-146b-3p to repress its expression. Overexpression of miR-146b-3p restored MM cell activities that were blocked by circ_0134426 overexpression. NDNF is a functional molecule targeted by miR-146b-3p, and miR-146b-3p sequesters NDNF expression. The inhibitory effects of NDNF upregulation on MM cell growth, colony formation, and migration were partially abolished by miR-146b-3p overexpression. Circ_0134426 regulates the miR-146b-3p/NDNF network to restrain the development of MM, to some extent, suggesting that the development of MM therapeutic strategies might focus on circ_0134426 regulatory networks.







Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in this article.
References
Ishiguro, K., Kitajima, H., Niinuma, T., Ishida, T., Maruyama, R., Ikeda, H., et al. (2019). DOT1L inhibition blocks multiple myeloma cell proliferation by suppressing IRF4-MYC signaling. Haematologica, 104(1), 155–165.
Gerecke, C., Fuhrmann, S., Strifler, S., Schmidt-Hieber, M., Einsele, H., & Knop, S. (2016). The diagnosis and treatment of multiple myeloma. Deutsches Arzteblatt international, 113(27–28), 470–476.
Richardson, P. G., Laubach, J., Gandolfi, S., Facon, T., Weisel, K., & O’Gorman, P. (2018). Maintenance and continuous therapy for multiple myeloma. Expert review of anticancer therapy, 18(8), 751–764.
Vo, J. N., Cieslik, M., Zhang, Y., Shukla, S., Xiao, L., Zhang, Y., et al. (2019). The landscape of circular RNA in cancer. Cell, 176(4), 869–81.e13.
Abu, N., & Jamal, R. (2016). Circular RNAs as promising biomarkers: A mini-review. Frontiers in physiology, 7, 355.
Zhao, Z. J., & Shen, J. (2017). Circular RNA participates in the carcinogenesis and the malignant behavior of cancer. RNA biology, 14(5), 514–521.
Cai, H., Li, Y., Niringiyumukiza, J. D., Su, P., & Xiang, W. (2019). Circular RNA involvement in aging: An emerging player with great potential. Mechanisms of ageing and development, 178, 16–24.
Liu, L., Zhang, F., & Li, J. (2021). CircRNA circ_0001821 predicts an unfavorable prognosis and promotes the proliferation of multiple myeloma. Hematology, 26(1), 716–723.
Tang, X., Deng, Z., Ding, P., Qiang, W., Lu, Y., Gao, S., et al. (2022). A novel protein encoded by circHNRNPU promotes multiple myeloma progression by regulating the bone marrow microenvironment and alternative splicing. Journal of Experimental & Clinical Cancer Research, 41(1), 85.
Yu, M., Yu, J., Zhang, Y., Sun, X., Sun, R., Xia, M., et al. (2022). A novel circRNA-miRNA-mRNA network revealed exosomal circ-ATP10A as a biomarker for multiple myeloma angiogenesis. Bioengineered, 13(1), 667–683.
Feng, Y., Zhang, L., Wu, J., Khadka, B., Fang, Z., Gu, J., et al. (2019). CircRNA circ_0000190 inhibits the progression of multiple myeloma through modulating miR-767-5p/MAPK4 pathway. Journal of Experimental & Clinical Cancer Research, 38(1), 54.
Liu, F., Wang, Y. L., Wei, J. M., & Huang, Z. D. (2021). Upregulation of circ_0000142 promotes multiple myeloma progression by adsorbing miR-610 and upregulating AKT3 expression. Journal of Biochemistry, 169(3), 327–336.
Salmena, L., Poliseno, L., Tay, Y., Kats, L., & Pandolfi., P. P. (2011). A ceRNA hypothesis: the rosetta stone of a hidden RNA language? Cell, 146(3), 353–8.
Zhou, M., Gao, Y., Wang, M., Guo, X., Li, X., Zhu, F., et al. (2021). MiR-146b-3p regulates proliferation of pancreatic cancer cells with stem cell-like properties by targeting MAP3K10. Journal of Cancer, 12(12), 3726–3740.
Liu, X., Fu, Q., Bian, X., Fu, Y., Xin, J., Liang, N., et al. (2020). Long non-coding RNA MAPK8IP1P2 inhibits lymphatic metastasis of thyroid cancer by activating hippo signaling via sponging miR-146b-3p. Frontiers in Oncology, 10, 600927.
Kuang, X. L., Zhao, X. M., Xu, H. F., Shi, Y. Y., Deng, J. B., & Sun, G. T. (2010). Spatio-temporal expression of a novel neuron-derived neurotrophic factor (NDNF) in mouse brains during development. BMC Neuroscience, 11, 137.
Joki, Y., Ohashi, K., Yuasa, D., Shibata, R., Kataoka, Y., Kambara, T., et al. (2015). Neuron-derived neurotrophic factor ameliorates adverse cardiac remodeling after experimental myocardial infarction. Circulation. Heart Failure, 8(2), 342–351.
Xia, L., Li, S., Liu, Y., Huang, Y., Ni, B., Wan, L., et al. (2019). NDNF inhibits the migration and invasion of human renal cancer cells through epithelial-mesenchymal transition. Oncology Letters, 17(3), 2969–2975.
Wallington-Beddoe, C. T., & Mynott, R. L. (2021). Prognostic and predictive biomarker developments in multiple myeloma. Journal of hematology & oncology, 14(1), 151.
Zhou, F., Wang, D., Wei, W., Chen, H., Shi, H., Zhou, N., et al. (2020). Comprehensive profiling of circular RNA expressions reveals potential diagnostic and prognostic biomarkers in multiple myeloma. BMC Cancer, 20(1), 40.
Song, Y., Hu, N., Song, X., & Yang, J. (2020). Hsa_Circ_0007841 enhances multiple myeloma chemotherapy resistance through upregulating ABCG2. Technology in Cancer Research & Treatment, 19, 1533033820928371.
Tang, X., Guo, M., Ding, P., Deng, Z., Ke, M., Yuan, Y., et al. (2021). BUB1B and circBUB1B_544aa aggravate multiple myeloma malignancy through evoking chromosomal instability. Signal Transduction and Targeted Therapy, 6(1), 361.
Yu, S., Ai, L., Wei, W., & Pan, J. (2021). circRNA circ-MYBL2 is a novel tumor suppressor and potential biomarker in multiple myeloma. Human Cell, 34(1), 219–228.
Wang, Y., Lin, Q., Song, C., Ma, R., & Li, X. (2020). Depletion of circ_0007841 inhibits multiple myeloma development and BTZ resistance via miR-129-5p/JAG1 axis. Cell cycle (Georgetown, Tex), 19(23), 3289–3302.
Tang, X., Ren, H., Guo, M., Qian, J., Yang, Y., & Gu, C. (2021). Review on circular RNAs and new insights into their roles in cancer. Computational and Structural Biotechnology Journal, 19, 910–928.
Yao, S., Xu, J., Zhao, K., Song, P., Yan, Q., Fan, W., et al. (2018). Down-regulation of HPGD by miR-146b-3p promotes cervical cancer cell proliferation, migration and anchorage-independent growth through activation of STAT3 and AKT pathways. Cell death & disease, 9(11), 1055.
Abulizi, R., Li, B., & Zhang, C. G. (2019). Circ_0071662, a novel tumor biomarker, suppresses bladder cancer cell proliferation and invasion by sponging miR-146b-3p. Oncology research. https://doi.org/10.3727/096504019X15740729375088
Hou, S., Xie, X., Zhao, J., Wu, C., Li, N., Meng, Z., et al. (2020). Downregulation of miR-146b-3p inhibits proliferation and migration and modulates the expression and location of sodium/iodide symporter in dedifferentiated thyroid cancer by potentially targeting MUC20. Frontiers in oncology, 10, 566365.
Zhang, Y., Wu, X., Kai, Y., Lee, C. H., Cheng, F., Li, Y., et al. (2019). Secretome profiling identifies neuron-derived neurotrophic factor as a tumor-suppressive factor in lung cancer. JCI insight, 4, 24.
Cholujova, D., Koklesova, L., LukacovaBujnakova, Z., Dutkova, E., Valuskova, Z., Beblava, P., et al. (2022). In vitro and ex vivo anti-myeloma effects of nanocomposite As(4)S(4)/ZnS/Fe(3)O(4). Science and Reports, 12(1), 17961.
Mahdi, M. A., Yousefi, S. R., Jasim, L. S., & Salavati-Niasari, M. (2022). Green synthesis of DyBa2Fe3O7.988/DyFeO3 nanocomposites using almond extract with dual eco-friendly applications: Photocatalytic and antibacterial activities. International Journal of Hydrogen Energy, 47(31), 14319–30.
Yousefi, S. R., Alshamsi, H. A., Amiri, O., & Salavati-Niasari, M. (2021). Synthesis, characterization and application of Co/Co3O4 nanocomposites as an effective photocatalyst for discoloration of organic dye contaminants in wastewater and antibacterial properties. Journal of Molecular Liquids, 337, 116405.
Yousefi, S. R., Ghanbari, D., Salavati-Niasari, M., & Hassanpour, M. (2016). Photo-degradation of organic dyes: Simple chemical synthesis of Ni(OH)2 nanoparticles, Ni/Ni(OH)2 and Ni/NiO magnetic nanocomposites. Journal of Materials Science: Materials in Electronics, 27(2), 1244–1253.
Yousefi, S. R., Masjedi-Arani, M., Morassaei, M. S., Salavati-Niasari, M., & Moayedi, H. (2019). Hydrothermal synthesis of DyMn2O5/Ba3Mn2O8 nanocomposite as a potential hydrogen storage material. International Journal of Hydrogen Energy, 44(43), 24005–24016.
Yousefi, S. R., Sobhani, A., & Salavati-Niasari, M. (2017). A new nanocomposite superionic system (CdHgI4/HgI2): Synthesis, characterization and experimental investigation. Advanced Powder Technology, 28(4), 1258–1262.
Yousefi, S. R., Ghanbari, D., & Salavati-Niasari, M. (2016). Hydrothermal synthesis of nickel hydroxide nanostructures and flame retardant poly vinyl alcohol and cellulose acetate nanocomposites. Journal of Nanostructures, 6(1), 80–85.
Yousefi, S. R., Sobhani, A., Alshamsi, H. A., & Salavati-Niasari, M. (2021). Green sonochemical synthesis of BaDy(2)NiO(5)/Dy(2)O(3) and BaDy(2)NiO(5)/NiO nanocomposites in the presence of core almond as a capping agent and their application as photocatalysts for the removal of organic dyes in water. RSC Advances, 11(19), 11500–11512.
Yousefi, S. R., Amiri, O., & Salavati-Niasari, M. (2019). Control sonochemical parameter to prepare pure Zn(0.35)Fe(2.65)O(4) nanostructures and study their photocatalytic activity. Ultrason Sonochem., 58, 104619.
Acknowledgements
None.
Funding
This study did not receive any funding in any form.
Author information
Authors and Affiliations
Contributions
MLL and JYL: performed the experiments and data analysis. YYS and MLL: conceived and designed the study. YYS and JYL: made the acquisition of data. YYS and MLL: did the analysis and interpretation of data. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Consent for Publication
Consent for publication was obtained from the participants.
Research involving Human Participants and/or Animals
The present study was approved by the Ethics Committee of Taikang Tongji (Wuhan)Hospital (Wuhan, China). The processing of clinical tissue samples is in strict compliance with the ethical standards of the Declaration of Helsinki. All patients signed written informed consent. This animal experiment was conducted in accordance with the ARRIVE guidelines and was authorized by the Ethics Committee of Taikang Tongji (Wuhan)Hospital.
Informed Consent
All patients signed written informed consent.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, M., Li, J. & Song, Y. Hsa_Circ_0134426 Attenuates the Malignant Biological Behaviors of Multiple Myeloma by Suppressing miR-146b-3p to Upregulate NDNF. Mol Biotechnol 65, 1165–1177 (2023). https://doi.org/10.1007/s12033-022-00618-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-022-00618-6