Abstract
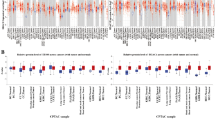
Hepatocellular carcinoma (HCC) is a global health problem with complex etiology and pathogenesis. Microarray data are increasingly being used as a novel and effective method for cancer pathogenesis analysis. An integrative analysis of genes and miRNA for HCC was conducted to unravel the potential prognosis of HCC. Two gene microarray datasets (GSE89377 and GSE101685) and two miRNA expression profiles (GSE112264 and GSE113740) were obtained from Gene Expression Omnibus database. A total of 177 differently expressed genes (DEGs) and 80 differently expressed miRNAs (DEMs) were screened out. Functional enrichment of DEGs was proceeded by Clue GO and these genes were significantly enriched in the chemical carcinogenesis pathway. A protein–protein interaction network was then established on the STRING platform, and ten hub genes (CDC20, TOP2A, ASPM, NCAPG, AURKA, CYP2E1, HMMR, PRC1, TYMS, and CYP4A11) were visualized via Cytoscape software. Then, a miRNA–target network was established to identify the hub dysregulated miRNA. A key miRNA (hsa-miR-124-3p) was filtered. Finally, the miRNA–target–transcription factor network was constructed for hsa-miR-124-3p. The network for hsa-miR-124-3p included two transcription factors (TFs) and five targets. These identified DEGs and DEMs, TFs, targets, and regulatory networks may help advance our understanding of the underlying pathogenesis of HCC.





Similar content being viewed by others
Abbreviations
- HCC:
-
Hepatocellular carcinoma
- PLC:
-
Primary liver cancer
- GEO:
-
Gene Expression Omnibus
- TFs:
-
Transcription factors
- PPI:
-
Protein–protein interaction
- DEGs:
-
Differently expressed genes
- DEMs:
-
Differently expressed miRNAs
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- BP:
-
Biological process
- MF:
-
Molecular function
- CC:
-
Cellular component
References
Dai X, Guo Y, Hu Y, et al. Immunotherapy for targeting cancer stem cells in hepatocellular carcinoma. Theranostics. 2021;11:3489–501.
Bray F, Ferlay J, Soerjomataram I, et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. Ca-Cancer J Clin. 2020;68:394–424.
Chen T, Dai X, Dai J, et al. AFP promotes HCC progression by suppressing the HuR-mediated Fas/FADD apoptotic pathway. Cell Death Dis. 2020;11(10):822.
Wang J, Wang CY. Integrated miRNA and mRNA omics reveal the anti-cancerous mechanism of Licochalcone B on Human Hepatoma Cell HepG2. Food Chem Toxicol. 2021;15:112096.
Zhang J, Song J, Wu D, et al. Hesperetin induces the apoptosis of hepatocellular carcinoma cells via mitochondrial pathway mediated by the increased intracellular reactive oxygen species, ATP and calcium. Med Oncol. 2015;32(4):101.
Chen C, Ge C, Liu Z, et al. ATF3 inhibits the tumorigenesis and progression of hepatocellular carcinoma cells via upregulation of CYR61 expression. J Exp Clin Cancer Res. 2018;37:263.
Yang S, Wang H, Qin C, et al. Up-regulation of CXCL8 expression is associated with a poor prognosis and enhances tumor cell malignant behaviors in liver cancer. 2020. Biosci Rep. https://doi.org/10.1042/BSR20201169.
Tian Z, Jiang H, Liu Y, et al. MicroRNA-133b inhibits hepatocellular carcinoma cell progression by targeting Sirt1. Exp Cell Res. 2016;343:135–47.
Chen H, Bao L, Hu J, et al. ORC6, negatively regulated by miR-1–3p, promotes proliferation, migration, and invasion of hepatocellular carcinoma cells. Front Cell Dev Biol. 2021;9:652292.
Long W, Li Q, Zhang J, et al. Identification of key genes in the tumor microenvironment of lung adenocarcinoma. Med Oncol. 2021;38(7):83.
Gao Y, Zhang S, Zhang Y, et al. Identification of microRNA-target gene-transcription factor regulatory networks in colorectal adenoma using microarray expression data. Front Genet. 2020;11:463.
Janky R, Verfaillie A, Imrichova H, et al. iRegulon: from a gene list to a gene regulatory network using large motif and track collections. PLoS Comput Biol. 2014;10:e1003731.
Llovet JM, Kelley RK, Villanueva A, et al. Hepatocellular carcinoma. Nat Rev Dis Primers. 2021;7:6.
Stults WP, Wei Y. Ambient air emissions of polycyclic aromatic hydrocarbons and female breast cancer incidence in US. Med Oncol. 2018;35:88.
Shih S, Huang YT, Yang HI. A multiple mediator analysis approach to quantify the effects of the ADH1B and ALDH2 genes on hepatocellular carcinoma risk. Genet Epidemiol. 2018;42:394–404.
Wei RR, Zhang MY, Rao HL, et al. Identification of ADH4 as a novel and potential prognostic marker in hepatocellular carcinoma. Med Oncol. 2012;29(2737):2743.
Yu J, Xia X, Dong Y, et al. CYP1A2 suppresses hepatocellular carcinoma through antagonizing HGF/MET signaling. Theranostics. 2021;11:2123–36.
Sugimoto M, Furuta T, Shirai N, et al. Poor metabolizer genotype status of CYP2C19 is a risk factor for developing gastric cancer in Japanese patients with helicobacter pylori infection. Aliment Pharmacol Ther. 2005;22:1033–40.
Kang JS, Wanibuchi H, Morimura K, et al. Role of CYP2E1 in diethylnitrosamine-induced hepatocarcinogenesis in vivo. Cancer Res. 2007;67:11141–6.
Li J, Gao JZ, Du JL, et al. Increased CDC20 expression is associated with development and progression of hepatocellular carcinoma. Int J Oncol. 2014;45:1547–55.
Wong N, Yeo W, Wong WL, et al. TOP2A overexpression in hepatocellular carcinoma correlates with early age onset, shorter patients survival and chemoresistance. Int J Cancer. 2009;124:644–52.
Lin SY, Pan HW, Liu SH, et al. ASPM is a novel marker for vascular invasion, early recurrence, and poor prognosis of hepatocellular carcinoma. Clin Cancer Res. 2008;14:4814–20.
Wang Y, Gao B, Tan PY, et al. Genome-wide CRISPR knockout screens identify NCAPG as an essential oncogene for hepatocellular carcinoma tumor growth. FASEB J. 2019;33:8759–70.
Dauch D, Rudalska R, Cossa G, et al. A MYC-aurora kinase A protein complex represents an actionable drug target in p53-altered liver cancer. Nat Med. 2016;22:744–53.
Gao J, Wang Z, Wang GJ, et al. From hepatofibrosis to hepatocarcinogenesis: higher cytochrome P450 2E1 activity is a potential risk factor. Mol Carcinog. 2018;57:1371–82.
Lei X, Zhang M, Guan B, et al. Identification of hub genes associated with prognosis, diagnosis, immune infiltration and therapeutic drug in liver cancer by integrated analysis. Hum Genomics. 2021;15:39.
Chen J, Rajasekaran M, Xia H, et al. The microtubule-associated protein PRC1 promotes early recurrence of hepatocellular carcinoma in association with the Wnt/beta-catenin signalling pathway. Gut. 2016;65:1522–34.
Wang X, Sun X, Du X, et al. Thymidylate synthase gene polymorphisms as important contributors affecting hepatocellular carcinoma prognosis. Clin Res Hepatol Gastroenterol. 2017;41:319–26.
Eun HS, Cho SY, Lee BS, et al. Cytochrome P450 4A11 expression in tumor cells: a favorable prognostic factor for hepatocellular carcinoma patients. J Gastroenterol Hepatol. 2019;34:224–33.
Gu J, Zhu X, Li Y, et al. miRNA-21 regulates arsenic-induced anti-leukemia activity in myelogenous cell lines. Med Oncol. 2011;28:211–8.
Paydas S, Acikalin A, Ergin M, et al. Micro-RNA (miRNA) profile in Hodgkin lymphoma: association between clinical and pathological variables. Med Oncol. 2016;33(4):34.
Zhang J, Zhang X, Li Z, et al. The miR-124–3p/neuropilin-1 axis contributes to the proliferation and metastasis of triple-negative breast cancer cells and co-activates the TGF-β pathway. Front Oncol. 2021;11:654672.
He D, Zhang X, Tu J. Diagnostic significance and carcinogenic mechanism of pan-cancer gene POU5F1 in liver hepatocellular carcinoma. Cancer Med. 2020;9:8782–800.
Zhang CZ, Chen SL, Wang CH, et al. CBX8 exhibits oncogenic activity via akt/β-catenin activation in hepatocellular carcinoma. Cancer Res. 2018;78:51–63.
Wang CJ, Zhang ZZ, Xu J, et al. SLITRK3 expression correlation to gastrointestinal stromal tumor risk rating and prognosis. World J Gastroenterol. 2015;21:8398–407.
Acknowledgements
We appreciate the GEO database and contributors providing their platforms and their meaningful datasets.
Funding
This work was supported by the Hefei University Scientific Research and Development Fund (20ZR09ZDB) and the Talent Research Fund Project of Hefei University (20RC48).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, J., Wang, C., Yang, L. et al. Identification of the critical genes and miRNAs in hepatocellular carcinoma by integrated bioinformatics analysis. Med Oncol 39, 21 (2022). https://doi.org/10.1007/s12032-021-01622-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-021-01622-7