Abstract
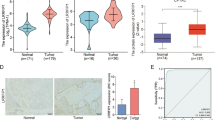
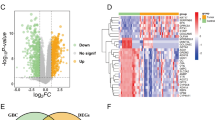
This study is to analyze differentially expressed genes (DEGs) and mutation signatures of pancreatic head cancer and pancreatic body/tail cancer. Pancreatic Adenocarcinoma (PAAD) RNA-seq data, mutation data and clinical data were downloaded and collected from The Cancer Genome Atlas (TCGA), FireHose and CBioPortal. According to the anatomic location, the patients were divided into 146 cases of pancreatic head cancer and 28 cases of pancreatic body/tail cancer. Then survival analysis was performed by Kaplan–Meier and log-rank test. Furthermore, DEGs were screened by R package Deseq2. Gene Ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG) and protein–protein interaction (PPI) were then carried out by DAVID and String. Online tool TIMER was used to analyze the immune cells infiltration. R package maftools and GenVisR were applied to analyze frequently mutated genes and mutant-allele tumor heterogeneity (MATH) of PAAD. Survival of patients with pancreatic body/tail cancer was better than those with pancreatic head cancer (median survival, 24.05 vs 19.45 months, p = 0.048). And 496 significant DEGs (|log2 FoldChange| > 1.5,false discovery rate (FDR) < 0.05) were identified, including 253 downregulated genes and 243 upregulated genes. And there were 13 Go terms (4 biological processes, 6 cellular components and 3 molecular functions) and 3 KEGG pathways (Pancreatic secretion, Fat digestion and absorption, Protein digestion and absorption) (FDR < 0.05). B cells and CD4 + T cells infiltration were more significant in pancreatic head cancer. MATH scores of pancreatic body/tail cancer were higher than pancreatic head cancer, while χ2 test of top 10 frequently mutated genes showed little difference between them. There were prognostic and genetic differences between pancreatic head cancer and pancreatic body/tail cancer. PAAD originated from different location may have different biology natures and should not be treated with same strategy.





Similar content being viewed by others
References
Ilic M, Ilic I. Epidemiology of pancreatic cancer. World J Gastroenterol. 2016;22(44):9694–705. https://doi.org/10.3748/wjg.v22.i44.9694.
Siegel RLMK, Jemal A. Cancer statistics, 2018. CA Cancer J Clin. 2018;68:7–30. https://doi.org/10.3322/caac.21442.
Neoptolemos JPPD, Ghaneh P, et al. Comparison of adjuvant gemcitabine and capecitabine with gemcitabine monotherapy in patients with resected pancreatic cancer (ESPAC-4): a multicentre, open-label, randomised, phase 3 trial. Lancet. 2017;389:1011–24.
Modolell IGL, Malagelada JR. Vagaries of clinical presentation of pancreatic and biliary tract cancer. Ann Oncol. 1999;10(Suppl 4):82–4. https://doi.org/10.1093/annonc/10.
Pitman MB, Lewandrowski K, Shen J, Sahani D, Brugge W, Fernandez-del CC. Pancreatic cysts. 2010;118(1):1–13. https://doi.org/10.1002/cncy.20059.
Artinyan A, Soriano PA, Prendergast C, Low T, Ellenhorn JD, Kim J. The anatomic location of pancreatic cancer is a prognostic factor for survival. HPB (Oxford). 2008;10(5):371–6. https://doi.org/10.1080/13651820802291233.
Watanabe I, Sasaki S, Konishi M, Nakagohri T, Inoue K, Oda T, et al. Onset Symptoms and Tumor Locations as Prognostic Factors of Pancreatic Cancer. 2004;28(2):160–5.
Lau MK, Davila JA, Shaib YH. Incidence and Survival of Pancreatic Head and Body and Tail Cancers: A Population-Based Study in the United States. 2010;39(4):458–62. https://doi.org/10.1097/MPA.0b013e3181bd6489.
Ling Q, Xu X, Zheng S-S, Kalthoff H. The diversity between pancreatic head and body/tail cancers: clinical parameters and in vitro models. Hepatobiliary Pancreatic Dis Int. 2013;12(5):480–7.
Ling Q, Xu X, Ye P, Xie H, Gao F, Hu Q, et al. The prognostic relevance of primary tumor location in patients undergoing resection for pancreatic ductal adenocarcinoma. Oncotarget. 2017;8(9):15159–677. https://doi.org/10.18632/oncotarget.14768.
Bailey P, Chang DK, Nones K, Johns AL, Patch A-M, Gingras M-C, et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature. 2016;531(7592):47–52.
Birnbaum DJ, Bertucci F, Finetti P, Birnbaum D, Mamessier E. Head and body/tail pancreatic carcinomas are not the same tumors. Cancers (Basel). 2019. https://doi.org/10.3390/cancers11040497.
Kamisawa T, Wood LD, Itoi T, Takaori K. Pancreatic cancer. Lancet. 2016;388(10039):73–85. https://doi.org/10.1016/s0140-6736(16)00141-0.
Walter FM, Mills K, Mendonça SC, Abel GA, Basu B, Carroll N, et al. Symptoms and patient factors associated with diagnostic intervals for pancreatic cancer (SYMPTOM pancreatic study): a prospective cohort study. Lancet Gastroenterol Hepatol. 2016;1(4):298–306. https://doi.org/10.1016/S2468-1253(16)30079-6.
Winter JM, Cameron JL, Campbell KA, Arnold MA, Chang DC, Coleman J, et al. 1423 Pancreaticoduodenectomies for pancreatic cancer: a single-institution experience. J. Gastroint. Surg. 2006;10(9):1199–211. https://doi.org/10.1016/j.gassur.2006.08.018.
Nathan H, Wolfgang CL, Edil BH, Choti MA, Herman JM, Schulick RD, et al. Peri-operative mortality and long-term survival after total pancreatectomy for pancreatic adenocarcinoma: a population-based perspective. J Surg Oncol. 2009;99(2):87–92. https://doi.org/10.1002/jso.21189.
Hartwig W, Werner J, Jäger D, Debus J, Büchler MW. Improvement of surgical results for pancreatic cancer. Lancet Oncol. 2013;14(11):e476–e485485. https://doi.org/10.1016/S1470-2045(13)70172-4.
Bilici A. Prognostic factors related with survival in patients with pancreatic adenocarcinoma. World J Gastroenterol. 2014;20(31):10802–12. https://doi.org/10.3748/wjg.v20.i31.10802.
Winer LK, Dhar VK, Wima K, Morris MC, Lee TC, Shah SA, et al. The impact of tumor location on resection and survival for pancreatic ductal adenocarcinoma. J Surg Res. 2019;239:60–6. https://doi.org/10.1016/j.jss.2019.01.061.
Gidekel Friedlander SY, Chu GC, Snyder EL, Girnius N, Dibelius G, Crowley D, et al. Context-dependent transformation of adult pancreatic cells by oncogenic K-Ras. Cancer Cell. 2009;16(5):379–89. https://doi.org/10.1016/j.ccr.2009.09.027.
Hu Y, Zhang Y, Ren J, Wang Y, Wang Z, Zhang J. Statistical Approaches for the Construction and Interpretation of Human Protein-Protein Interaction Network. BioMed Res Int. 2016;2016:5313050. https://doi.org/10.1155/2016/5313050.
Huang Z, Duan H, Li H. Identification of gene expression pattern related to breast cancer survival using integrated tcga datasets and genomic tools. BioMed Res Int. 2015;2015:878546. https://doi.org/10.1155/2015/878546.
Vishnubalaji R, Hamam R, Abdulla MH, Mohammed MAV, Kassem M, Al-Obeed O, et al. Genome-wide mRNA and miRNA expression profiling reveal multiple regulatory networks in colorectal cancer. Cell Death Dis. 2015;6(1):e16–e1414. https://doi.org/10.1038/cddis.2014.556.
Nakata B, Fukunaga S, Noda E, Amano R, Yamada N, Hirakawa K. Chemokine receptor CCR7 expression correlates with lymph node metastasis in pancreatic cancer. Oncology. 2008;74(1–2):69–75. https://doi.org/10.1159/000139126.
Zhang L, Wang D, Li Y, Liu Y, Xie X, Wu Y, et al. CCL21/CCR7 axis contributed to CD133+ pancreatic cancer stem-like cell metastasis via EMT and Erk/NF-kappaB pathway. PLoS ONE. 2016;11(8):e0158529. https://doi.org/10.1371/journal.pone.0158529.
Almendro V, Marusyk A, Polyak K. Cellular Heterogeneity and Molecular Evolution in Cancer. 2013;8(1):277–302. https://doi.org/10.1146/annurev-pathol-020712-163923.
Mroz EA, Rocco JW. MATH, a novel measure of intratumor genetic heterogeneity, is high in poor-outcome classes of head and neck squamous cell carcinoma. Oral Oncol. 2013;49(3):211–5. https://doi.org/10.1016/j.oraloncology.2012.09.007.
Pylayeva-Gupta Y, Das S, Handler JS, Hajdu CH, Coffre M, Koralov SB, et al. IL35-producing B cells promote the development of pancreatic neoplasia. Cancer Discov. 2016;6(3):247–55.
Liu L, Zhao G, Wu W, Rong Y, Jin D, Wang D, et al. Low intratumoral regulatory T cells and high peritumoral CD8+ T cells relate to long-term survival in patients with pancreatic ductal adenocarcinoma after pancreatectomy. Cancer Immunol Immunother. 2016;65(1):73–82.
Tomczak K, Czerwinska P, Wiznerowicz M. The Cancer Genome Atlas (TCGA): an immeasurable source of knowledge. Contemp Oncol. (Pozn). 2015;19(1A):A68–77. https://doi.org/10.5114/wo.2014.47136.
Acknowledgements
This work was supported by the grants from the National Natural Science Foundation of China (81672449, 81871980), The Project of Invigorating Health Care through Science, Technology and Education, Jiangsu Provincial Medical Outstanding Talent (to Yi Miao, JCRCA2016009), Innovation Capability Development Project of Jiangsu Province (No. BM2015004), Jiangsu Key Medical Discipline (General Surgery) (ZDXKA2016005) and Wu Jiepin Medical Foundation (No. 320.2710.1858).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All authors have declared no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yin, L., Xiao, L., Gao, Y. et al. Comparative bioinformatical analysis of pancreatic head cancer and pancreatic body/tail cancer. Med Oncol 37, 46 (2020). https://doi.org/10.1007/s12032-020-01370-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-020-01370-0