Abstract
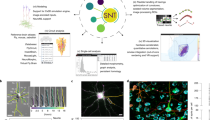
The study of neuron morphology requires robust and comprehensive methods to quantify the differences between neurons of different subtypes and animal species. Several software packages have been developed for the analysis of neuron tracing results stored in the standard SWC format. The packages, however, provide relatively simple quantifications and their non-extendable architecture prohibit their use for advanced data analysis and visualization. We developed nGauge, a Python toolkit to support the parsing and analysis of neuron morphology data. As an application programming interface (API), nGauge can be referenced by other popular open-source software to create custom informatics analysis pipelines and advanced visualizations. nGauge defines an extendable data structure that handles volumetric constructions (e.g. soma), in addition to the SWC linear reconstructions, while remaining lightweight. This greatly extends nGauge’s data compatibility.








Similar content being viewed by others
References
Abdellah, M., Hernando, J., Eilemann, S., Lapere, S., Antille, N., Markram, H., & Schürmann, F. (2018). NeuroMorphoVis: A collaborative framework for analysis and visualization of neuronal morphology skeletons reconstructed from microscopy stacks. Bioinformatics, 34(13), i574–i582.
Ascoli, G. A., Donohue, D. E., & Halavi, M. (2007). NeuroMorpho.Org: a central resource for neuronal morphologies. The Journal of neuroscience: the official journal of the Society for Neuroscience, 27(35), 9247–9251.
Barber, C. B., Dobkin, D. P., & Huhdanpaa, H. (1996). The quickhull algorithm for convex hulls. ACM transactions on mathematical software. Association for Computing Machinery, 22(4), 469–483.
Bates, A. S., Manton, J. D., Jagannathan, S. R., Costa, M., Schlegel, P., Rohlfing, T., & Jefferis, G. S. (2020). The natverse, a versatile toolbox for combining and analysing neuroanatomical data. eLife, 9. https://doi.org/10.7554/eLife.53350
BRAIN Initiative Cell Census Network (BICCN), Adkins, R. S., Aldridge, A. I., Allen, S., Ament, S. A., An, X., et al. (2020, October 21) A Multimodal Cell Census and Atlas of the Mammalian Primary Motor Cortex. Biorxiv. https://doi.org/10.1101/2020.10.19.343129
Claudi, F., Tyson, A. L., & Branco, T. (2020, February 25). Brainrender. A python based software for visualisation of neuroanatomical and morphological data. bioRxiv. https://doi.org/10.1101/2020.02.23.961748
Cuntz, H., Forstner, F., Borst, A., & Häusser, M. (2010). One rule to grow them all: a general theory of neuronal branching and its practical application. PLoS computational biology, 6(8). https://doi.org/10.1371/journal.pcbi.1000877
Dizaji, A. S., Walker, L. A., & Cai, D. (2020). TraceMontage: A method for merging multiple independent neuronal traces. Journal of neuroscience methods, 332, 108560.
Duan, B., Walker, L. A., Roossien, D. H., Shen, F. Y., Cai, D., & Yan, Y. (2020, June 8). Unsupervised Neural Tracing in Densely Labeled Multispectral Brainbow Images. Biorxiv. https://doi.org/10.1101/2020.06.07.138941.
Fukunaga, I., Berning, M., Kollo, M., Schmaltz, A., & Schaefer, A. T. (2012). Two distinct channels of olfactory bulb output. Neuron, 75(2), 320–329.
Gouwens, N. W., Sorensen, S. A., Baftizadeh, F., Budzillo, A., Lee, B. R., Jarsky, T., et al. (2020). Integrated Morphoelectric and Transcriptomic Classification of Cortical GABAergic Cells. Cell, 183(4), 935-953.e19.
Gouwens, N. W., Sorensen, S. A., Berg, J., Lee, C., Jarsky, T., Ting, J., et al. (2019). Classification of electrophysiological and morphological neuron types in the mouse visual cortex. Nature Neuroscience, 22(7), 1182–1195.
Harris, C. R., Millman, K. J., van der Walt, S. J., Gommers, R., Virtanen, P., Cournapeau, D., et al. (2020). Array programming with NumPy. Nature, 585(7825), 357–362.
Hunter, J. D. (2007). Matplotlib: A 2D Graphics Environment. Computing in Science Engineering, 9(3), 90–95.
Jiang, S., Wang, Y., Liu, L., Zhao, S., Chen, M., Zhao, X., et al. (2021, June 10). Petabyte-Scale Multi-Morphometry of Single Neurons for Whole Brains. Biorxiv. https://doi.org/10.1101/2021.01.09.426010.
Kent, B. R. (2014). 3D Scientific Visualization with Blender. Morgan & Claypool Publishers.
Laturnus, S., Kobak, D., & Berens, P. (2020). A Systematic Evaluation of Interneuron Morphology Representations for Cell Type Discrimination. Neuroinformatics. https://doi.org/10.1007/s12021-020-09461-z
Laturnus, S., von Daranyi, A., Huang, Z., & Berens, P. (2020). MorphoPy: A python package for feature extraction of neural morphologies. Journal of Open Source Software, 5(52), 2339.
Li, Y., Walker, L. A., Zhao, Y., Edwards, E. M., Michki, N. S., Cheng, H. P. J., et al. (2020, April 9). Bitbow: a digital format of Brainbow enables highly efficient neuronal lineage tracing and morphology reconstruction in single brains. bioRxiv. https://doi.org/10.1101/2020.04.07.030593
Miyamae, T., Chen, K., Lewis, D. A., & Gonzalez-Burgos, G. (2017). Distinct Physiological Maturation of Parvalbumin-Positive Neuron Subtypes in Mouse Prefrontal Cortex. The Journal of Neuroscience: THe Official Journal of the Society for Neuroscience, 37(19), 4883–4902.
Motta, A., Berning, M., Boergens, K. M., Staffler, B., Beining, M., Loomba, S., et al. (2019). Dense connectomic reconstruction in layer 4 of the somatosensory cortex. Science. https://doi.org/10.1126/science.aay3134
Nanda, S., Chen, H., Das, R., Bhattacharjee, S., Cuntz, H., Torben-Nielsen, B., et al. (2018). Design and implementation of multi-signal and time-varying neural reconstructions. Scientific data, 5, 170207.
Pedregosa, F., Varoquaux, G., Gramfort, A., Michel, V., Thirion, B., Grisel, O., et al. (2011). Scikit-learn: Machine Learning in Python. Journal of machine learning research: JMLR, 12(85), 2825–2830. Accessed 25 Jan 2021
Peng, H., Bria, A., Zhou, Z., Iannello, G., & Long, F. (2014). Extensible visualization and analysis for multidimensional images using Vaa3D. Nature Protocols, 9(1), 193–208.
Phelps, J. S., Hildebrand, D. G. C., Graham, B. J., Kuan, A. T., Thomas, L. A., Nguyen, T. M., et al. (2021). Reconstruction of motor control circuits in adult Drosophila using automated transmission electron microscopy. Cell, 184(3), 759-774.e18.
Quentin, E., Belmer, A., & Maroteaux, L. (2018). Somato-Dendritic Regulation of Raphe Serotonin Neurons; A Key to Antidepressant Action. Frontiers in Neuroscience, 12, 982.
Ramón y Cajal, S. (1892). La rétine des vertébrés. Lierre [etc.]: Van In [etc.].
Roossien, D. H., Sadis, B. V., Yan, Y., Webb, J. M., Min, L. Y., Dizaji, A. S., et al. (2019). Multispectral tracing in densely labeled mouse brain with nTracer. Bioinformatics, 35(18), 3544–3546.
Schindelin, J., Arganda-Carreras, I., Frise, E., Kaynig, V., Longair, M., Pietzsch, T., et al. (2012). Fiji: An open-source platform for biological-image analysis. Nature Methods, 9(7), 676–682.
Scorcioni, R., Polavaram, S., & Ascoli, G. A. (2008). L-Measure: A web-accessible tool for the analysis, comparison and search of digital reconstructions of neuronal morphologies. Nature Protocols, 3(5), 866–876.
Shen, F. Y., Harrington, M. M., Walker, L. A., Cheng, H. P. J., Boyden, E. S., & Cai, D. (2020). Light microscopy based approach for mapping connectivity with molecular specificity. Cold Spring Harbor Laboratory. https://doi.org/10.1101/2020.02.24.963538
Stokes, C. C. A., Teeter, C. M., & Isaacson, J. S. (2014). Single dendrite-targeting interneurons generate branch-specific inhibition. Frontiers in Neural Circuits, 8, 139.
Torben-Nielsen, B. (2014). An efficient and extendable python library to analyze neuronal morphologies. Neuroinformatics, 12(4), 619–622.
Virtanen, P., Gommers, R., Oliphant, T. E., Haberland, M., Reddy, T., Cournapeau, D., et al. (2020). SciPy 1.0: fundamental algorithms for scientific computing in Python. Nature methods, 17(3), 261–272.
Wang, Q., Ding, S.-L., Li, Y., Royall, J., Feng, D., Lesnar, P., et al. (2020). The Allen Mouse Brain Common Coordinate Framework: A 3D Reference Atlas. Cell, 181(4), 936-953.e20.
Yin, W., Brittain, D., Borseth, J., Scott, M. E., Williams, D., Perkins, J., et al. (2020). A petascale automated imaging pipeline for mapping neuronal circuits with high-throughput transmission electron microscopy. Nature Communications, 11(1), 4949.
Acknowledgements
JSW received support from the University of Michigan Women in Science and Engineering Residence Program (WISE-RP) Judith Cram Memorial Fund Research Award. WJL received support from the Magnificent Michigan Fellowship. LAW and DC received support from NSF-1707316 (Neuronex-MINT), NIH-RF1MH123402, and NIH-RF1MH124611. The authors thank Fred Shen for his comments on an early version of the library. LAW thanks Chris Midkiff for his comments on figure design and example code clarity.
Author information
Authors and Affiliations
Contributions
LAW and DC conceptualized the nGauge library, which was then implemented by LAW and JSW. YL and DR provided imaging and neuron reconstruction datasets which were used in library testing. LAW, JSW, YL, WJL, and NM contributed to beta testing of early versions of nGauge and provided comments on the library design. LAW, JSW, and DC wrote the manuscript, which was edited and approved by all authors.
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12021_2022_9573_MOESM1_ESM.eps
Supplementary file 1 (EPS 1169 KB) Supplementary Figure 1 Extended L-Measure Comparison. A collection of 13 functions highlighted in Supplementary Table 1 are compared between nGauge (xaxis) and L-Measure (y-axis) by calculating each metric in both pieces of software. We find that all functions perform as expected.
12021_2022_9573_MOESM2_ESM.xlsx
Supplementary file 2 (XLSX 16 KB) Supplementary Table 1 Implemented nGauge Functions. All functions implemented in nGauge. Along the left-hand side, functions are divided into different modules and function types. For each function, we have identified the closest equivalent function in LnGauge Walker LA, Williams JS, et al. Measure, MorphoPy, and TREEs toolbox, however, not all functions are exact matches due to differencesin API design between all four programs. As a result, in some cases, summations or other summary functions are required to make results match between each program.
Rights and permissions
About this article
Cite this article
Walker, L.A., Williams, J.S., Li, Y. et al. nGauge: Integrated and Extensible Neuron Morphology Analysis in Python. Neuroinform 20, 755–764 (2022). https://doi.org/10.1007/s12021-022-09573-8
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12021-022-09573-8