Abstract
Background
Despite highly effective machinery for the maintenance of genome integrity in human embryonic stem cells (hESCs), the frequency of genetic aberrations during in-vitro culture has been a serious issue for future clinical applications.
Method
By passaging hESCs over a broad range of timepoints (up to 6 years), the isogenic hESC lines with different passage numbers with distinct cellular characteristics, were established.
Result
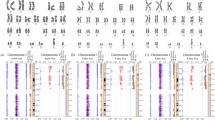
We found that mitotic aberrations, such as the delay of mitosis, multipolar centrosomes, and chromosome mis-segregation, were increased in parallel with polyploidy compared to early-passaged hESCs (EP-hESCs) with normal copy number. Through high-resolution genome-wide approaches and transcriptome analysis, we found that culture adapted-hESCs with a minimal amplicon in chromosome 20q11.21 highly expressed TPX2, a key protein for governing spindle assembly and cancer malignancy. Consistent with these findings, the inducible expression of TPX2 in EP-hESCs reproduced aberrant mitotic events, such as the delay of mitotic progression, spindle stabilization, misaligned chromosomes, and polyploidy.
Conclusion
These studies suggest that the increased transcription of TPX2 in culture adapted hESCs could contribute to an increase in aberrant mitosis due to altered spindle dynamics.
Graphical abstract







Similar content being viewed by others
Data Availability
Source data are available from the corresponding authors upon request. The RNAseq results have been deposited to Gene Expression Omnibus (GEO) under accession number GSE167495 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE167495). The flow cytometry data were deposited in Flowrepository (https://flowrepository.org/id/FR-FCM-Z3W9).
Abbreviations
- hESC:
-
Human embryonic stem cell.
- hPSC:
-
Human pluripotent stem cell.
- hiPSCs:
-
Human induced pluripotent stem cell.
- Chr:
-
Chromosome.
- EP:
-
Early-passage.
- LP:
-
Late-passage.
- DMSO:
-
Dimethyl sulfoxide.
- TPX2:
-
Targeting protein for Xklp2.
- aCGH:
-
Array comparative genomic hybridization.
- CNV:
-
Copy number variation.
- CIN:
-
Chromosome instability.
- 7-AAD:
-
7-Aminoactinomycin D.
- MtOH:
-
Methanol.
- EtOH:
-
Ethanol
- PI:
-
Propidium iodide
References
Weissbein, U., Benvenisty, N., & Ben-David, U. (2014). Quality control: Genome maintenance in pluripotent stem cells. Journal of Cell Biology, 204(2), 153–163.
Maitra, A., Arking, D. E., Shivapurkar, N., Ikeda, M., Stastny, V., Kassauei, K., Sui, G., Cutler, D. J., Liu, Y., Brimble, S. N., et al. (2005). Genomic alterations in cultured human embryonic stem cells. Nature Genetics, 37(10), 1099–1103.
Baker, D. E., Harrison, N. J., Maltby, E., Smith, K., Moore, H. D., Shaw, P. J., Heath, P. R., Holden, H., & Andrews, P. W. (2007). Adaptation to culture of human embryonic stem cells and oncogenesis in vivo. Nature Biotechnology, 25(2), 207–215.
Peterson, S. E., & Loring, J. F. (2014). Genomic instability in pluripotent stem cells: Implications for clinical applications. Journal of Biological Chemistry, 289(8), 4578–4584.
Andrews, P. W., Ben-David, U., Benvenisty, N., Coffey, P., Eggan, K., Knowles, B. B., Nagy, A., Pera, M., Reubinoff, B., Rugg-Gunn, P. J., et al. (2017). Assessing the Safety of Human Pluripotent Stem Cells and Their Derivatives for Clinical Applications. Stem Cell Reports, 9(1), 1–4.
Garber, K. (2015). RIKEN suspends first clinical trial involving induced pluripotent stem cells. Nature Biotechnology, 33(9), 890–891.
Lefort, N., Feyeux, M., Bas, C., Feraud, O., Bennaceur-Griscelli, A., Tachdjian, G., Peschanski, M., & Perrier, A. L. (2008). Human embryonic stem cells reveal recurrent genomic instability at 20q11.21. Nature Biotechnology, 26(12), 1364–1366.
Spits, C., Mateizel, I., Geens, M., Mertzanidou, A., Staessen, C., Vandeskelde, Y., Van der Elst, J., Liebaers, I., & Sermon, K. (2008). Recurrent chromosomal abnormalities in human embryonic stem cells. Nature Biotechnology, 26(12), 1361–1363.
International Stem Cell I, Amps K, Andrews PW, Anyfantis G, Armstrong L, Avery S, Baharvand H, Baker J, Baker D, Munoz MB et al:(2011). Screening ethnically diverse human embryonic stem cells identifies a chromosome 20 minimal amplicon conferring growth advantage. Nature Biotechnology, 29(12):1132–1144.
Biancotti, J. C., & Benvenisty, N. (2011). Aneuploid human embryonic stem cells: Origins and potential for modeling chromosomal disorders. Regenerative Medicine, 6(4), 493–503.
Avery, S., Hirst, A. J., Baker, D., Lim, C. Y., Alagaratnam, S., Skotheim, R. I., Lothe, R. A., Pera, M. F., Colman, A., Robson, P., et al. (2013). BCL-XL mediates the strong selective advantage of a 20q11.21 amplification commonly found in human embryonic stem cell cultures. Stem Cell Reports., 1(5), 379–386.
Cho, S. J., Kim, K. T., Jeong, H. C., Park, J. C., Kwon, O. S., Song, Y. H., Shin, J. G., Kang, S., Kim, W., Shin, H. D., et al. (2018). Selective Elimination of Culture-Adapted Human Embryonic Stem Cells with BH3 Mimetics. Stem Cell Reports, 11(5), 1244–1256.
Markouli, M. C., Couvreu de Deckersberg, E., Regin, M., Nguyen, H. T., Zambelli, F., Keller, A., Dziedzicka, D., de Kock, J., Tilleman, L., van Nieuwerburgh, F., et al. (2019). Gain of 20q1121 in Human Pluripotent Stem Cells Impairs TGF-beta-Dependent Neuroectodermal Commitment. Stem Cell Reports, 13(1), 163–176.
Merkle, F. T., Ghosh, S., Kamitaki, N., Mitchell, J., Avior, Y., Mello, C., Kashin, S., Mekhoubad, S., Ilic, D., Charlton, M., et al. (2017). Human pluripotent stem cells recurrently acquire and expand dominant negative P53 mutations. Nature, 545(7653), 229–233.
Werbowetski-Ogilvie, T. E., Bosse, M., Stewart, M., Schnerch, A., Ramos-Mejia, V., Rouleau, A., Wynder, T., Smith, M. J., Dingwall, S., Carter, T., et al. (2009). Characterization of human embryonic stem cells with features of neoplastic progression. Nature Biotechnology, 27(1), 91–97.
Nori, S., Okada, Y., Nishimura, S., Sasaki, T., Itakura, G., Kobayashi, Y., Renault-Mihara, F., Shimizu, A., Koya, I., Yoshida, R., et al. (2015). Long-term safety issues of iPSC-based cell therapy in a spinal cord injury model: Oncogenic transformation with epithelial-mesenchymal transition. Stem Cell Reports, 4(3), 360–373.
Ben-David, U., Arad, G., Weissbein, U., Mandefro, B., Maimon, A., Golan-Lev, T., Narwani, K., Clark, A. T., Andrews, P. W., Benvenisty, N., et al. (2014). Aneuploidy induces profound changes in gene expression, proliferation and tumorigenicity of human pluripotent stem cells. Nature Communications, 5, 4825.
Jo HY, Lee Y, Ahn H, Han HJ, Kwon A, Kim BY, Ha HY, Kim SC, Kim JH, Kim YO et al: Functional in vivo and in vitro effects of 20q11.21 genetic aberrations on hPSC differentiation. Sci Rep 2020, 10(1):18582.
Martins-Taylor, K., Nisler, B. S., Taapken, S. M., Compton, T., Crandall, L., Montgomery, K. D., Lalande, M., & Xu, R. H. (2011). Recurrent copy number variations in human induced pluripotent stem cells. Nature Biotechnology, 29(6), 488–491.
Narva, E., Autio, R., Rahkonen, N., Kong, L., Harrison, N., Kitsberg, D., Borghese, L., Itskovitz-Eldor, J., Rasool, O., Dvorak, P., et al. (2010). High-resolution DNA analysis of human embryonic stem cell lines reveals culture-induced copy number changes and loss of heterozygosity. Nature Biotechnology, 28(4), 371–377.
Draper, J. S., Smith, K., Gokhale, P., Moore, H. D., Maltby, E., Johnson, J., Meisner, L., Zwaka, T. P., Thomson, J. A., & Andrews, P. W. (2004). Recurrent gain of chromosomes 17q and 12 in cultured human embryonic stem cells. Nature Biotechnology, 22(1), 53–54.
Ben-David, U. (2015). Genomic instability, driver genes and cell selection: Projections from cancer to stem cells. Biochimica et Biophysica Acta, 1849(4), 427–435.
Moon, S. H., Kim, J. S., Park, S. J., Lim, J. J., Lee, H. J., Lee, S. M., & Chung, H. M. (2011). Effect of chromosome instability on the maintenance and differentiation of human embryonic stem cells in vitro and in vivo. Stem Cell Research, 6(1), 50–59.
Lamm, N., Ben-David, U., Golan-Lev, T., Storchova, Z., Benvenisty, N., & Kerem, B. (2016). Genomic Instability in Human Pluripotent Stem Cells Arises from Replicative Stress and Chromosome Condensation Defects. Cell Stem Cell, 18(2), 253–261.
Zhang, J., Hirst, A. J., Duan, F., Qiu, H., Huang, R., Ji, Y., Bai, L., Zhang, F., Robinson, D., Jones, M., et al. (2019). Anti-apoptotic Mutations Desensitize Human Pluripotent Stem Cells to Mitotic Stress and Enable Aneuploid Cell Survival. Stem Cell Reports, 12(3), 557–571.
Jacobs, K., Zambelli, F., Mertzanidou, A., Smolders, I., Geens, M., Nguyen, H. T., Barbe, L., Sermon, K., & Spits, C. (2016). Higher-Density Culture in Human Embryonic Stem Cells Results in DNA Damage and Genome Instability. Stem Cell Reports, 6(3), 330–341.
Liu, W., Ren, Z., Lu, K., Song, C., Cheung, E. C. W., Zhou, Z., & Chen, G. (2018). The Suppression of Medium Acidosis Improves the Maintenance and Differentiation of Human Pluripotent Stem Cells at High Density in Defined Cell Culture Medium. International Journal of Biological Sciences, 14(5), 485–496.
Neumayer, G., Belzil, C., Gruss, O. J., & Nguyen, M. D. (2014). TPX2: Of spindle assembly, DNA damage response, and cancer. Cellular and Molecular Life Sciences, 71(16), 3027–3047.
Asteriti, I. A., Rensen, W. M., Lindon, C., Lavia, P., & Guarguaglini, G. (2010). The Aurora-A/TPX2 complex: A novel oncogenic holoenzyme? Biochimica et Biophysica Acta, 1806(2), 230–239.
Bayliss, R., Sardon, T., Vernos, I., & Conti, E. (2003). Structural basis of Aurora-A activation by TPX2 at the mitotic spindle. Molecular Cell, 12(4), 851–862.
Naso F. D., Sterbini V., Crecca E., Asteriti I. A., Russo A. D., Giubettini M., Cundari E., Lindon C., Rosa A., Guarguaglini G. (2020). Excess TPX2 Interferes with Microtubule Disassembly and Nuclei Reformation at Mitotic Exit. Cells. 9(2).
Aguirre-Portoles, C., Bird, A. W., Hyman, A., & Canamero, M. (2012). Perez de Castro I, Malumbres M: Tpx2 controls spindle integrity, genome stability, and tumor development. Cancer Research, 72(6), 1518–1528.
Heidebrecht, H. J., Adam-Klages, S., Szczepanowski, M., Pollmann, M., Buck, F., Endl, E., Kruse, M. L., Rudolph, P., & Parwaresch, R. (2003). repp86: A human protein associated in the progression of mitosis. Molecular Cancer Research, 1(4), 271–279.
Wang, Y., Wang, H., Yan, Z., Li, G., Hu, G., Zhang, H., Huang, D., Wang, Y., Zhang, X., Yan, Y., et al. (2020). The critical role of dysregulated Hh-FOXM1-TPX2 signaling in human hepatocellular carcinoma cell proliferation. Cell Communication and Signaling: CCS, 18(1), 116.
Carter, S. L., Eklund, A. C., Kohane, I. S., Harris, L. N., & Szallasi, Z. (2006). A signature of chromosomal instability inferred from gene expression profiles predicts clinical outcome in multiple human cancers. Nature Genetics, 38(9), 1043–1048.
Jang, C. Y., Wong, J., Coppinger, J. A., Seki, A., Yates, J. R., 3rd., & Fang, G. (2008). DDA3 recruits microtubule depolymerase Kif2a to spindle poles and controls spindle dynamics and mitotic chromosome movement. The Journal of Cell Biology, 181(2), 255–267.
Schwartz, P. H., Brick, D. J., Nethercott, H. E., & Stover, A. E. (2011). Traditional human embryonic stem cell culture. Methods in Molecular Biology, 767, 107–123.
Barbaric, I., Biga, V., Gokhale, P. J., Jones, M., Stavish, D., Glen, A., Coca, D., & Andrews, P. W. (2014). Time-lapse analysis of human embryonic stem cells reveals multiple bottlenecks restricting colony formation and their relief upon culture adaptation. Stem Cell Reports, 3(1), 142–155.
Kramer, A., Maier, B., & Bartek, J. (2011). Centrosome clustering and chromosomal (in)stability: A matter of life and death. Molecular Oncology, 5(4), 324–335.
Castle, J. C., Loewer, M., Boegel, S., Tadmor, A. D., Boisguerin, V., de Graaf, J., Paret, C., Diken, M., Kreiter, S., Tureci, O., et al. (2014). Mutated tumor alleles are expressed according to their DNA frequency. Science and Reports, 4, 4743.
Witham, F. H., & Hendry, L. B. (1992). Computer modeling of gibberellin-DNA binding. Journal of Theoretical Biology, 155(1), 55–67.
Yao, R., Kondoh, Y., Natsume, Y., Yamanaka, H., Inoue, M., Toki, H., Takagi, R., Shimizu, T., Yamori, T., Osada, H., et al. (2014). A small compound targeting TACC3 revealed its different spatiotemporal contributions for spindle assembly in cancer cells. Oncogene, 33(33), 4242–4252.
Kufer, T. A., Sillje, H. H., Korner, R., Gruss, O. J., Meraldi, P., & Nigg, E. A. (2002). Human TPX2 is required for targeting Aurora-A kinase to the spindle. Journal of Cell Biology, 158(4), 617–623.
Lee, D. F., Su, J., Ang, Y. S., Carvajal-Vergara, X., Mulero-Navarro, S., Pereira, C. F., Gingold, J., Wang, H. L., Zhao, R., Sevilla, A., et al. (2012). Regulation of embryonic and induced pluripotency by aurora kinase-p53 signaling. Cell Stem Cell, 11(2), 179–194.
Deuschle, U., Meyer, W. K., & Thiesen, H. J. (1995). Tetracycline-reversible silencing of eukaryotic promoters. Molecular and Cellular Biology, 15(4), 1907–1914.
Blum, B., & Benvenisty, N. (2009). The tumorigenicity of diploid and aneuploid human pluripotent stem cells. Cell Cycle, 8(23), 3822–3830.
Na, J., Baker, D., Zhang, J., Andrews, P. W., & Barbaric, I. (2014). Aneuploidy in pluripotent stem cells and implications for cancerous transformation. Protein & Cell, 5(8), 569–579.
Nguyen, H. T., Geens, M., Mertzanidou, A., Jacobs, K., Heirman, C., Breckpot, K., & Spits, C. (2014). Gain of 20q11.21 in human embryonic stem cells improves cell survival by increased expression of Bcl-xL. Molecular Human Reproduction, 20(2), 168–177.
Kline-Smith, S. L., & Walczak, C. E. (2004). Mitotic spindle assembly and chromosome segregation: Refocusing on microtubule dynamics. Molecular Cell, 15(3), 317–327.
van Gijn, S. E., Wierenga, E., van den Tempel, N., Kok, Y. P., Heijink, A. M., Spierings, D. C. J., Foijer, F., van Vugt, M., & Fehrmann, R. S. N. (2019). TPX2/Aurora kinase A signaling as a potential therapeutic target in genomically unstable cancer cells. Oncogene, 38(6), 852–867.
Acknowledgements
We thank Dr. Minh Dang Nguyen in University of Calgary for kindly providing TPX2 constructs.
Funding
This work including the design of the study and collection, analysis, and interpretation of data, was supported by a grant from the National Research Foundation of Korea (NRF) (NRF-2020M3A9E4037905 and NRF-2020R1A2C2005914) and Korean Fund for Regenerative Medicine funded by Ministry of Science and ICT, and Ministry of Health and Welfare (Grant number RS-2022–00070316), Republic of Korea.
Author information
Authors and Affiliations
Contributions
HJ.C conceived the overall study design, led the experiments and wrote the manuscript. HC.J and YH.G conducted the experiments, data analysis and wrote the first draft. JG.S and HD.S performed CGH array and exome sequencing and analyzed the data. HS.L analyzed the transcriptome data. YJ.K validated the experiments and generated cell lines. MG.C, D.G, HS.C, JH.L and CY.J performed cell cycle analysis and measured the spindle stability. All authors contributed to manuscript writing and revising and endorsed the final version of this manuscript.
Corresponding authors
Ethics declarations
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file1 (MP4 710 KB)
Supplementary file2 (MP4 900 KB)
Supplementary file3 (MP4 733 KB)
Supplementary file4 (MP4 1054 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jeong, HC., Go, YH., Shin, JG. et al. TPX2 Amplification-Driven Aberrant Mitosis in Culture Adapted Human Embryonic Stem Cells with gain of 20q11.21. Stem Cell Rev and Rep 19, 1466–1481 (2023). https://doi.org/10.1007/s12015-023-10514-4
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-023-10514-4