Abstract
The "bivalent domain" is a unique histone modification region consisting of two histone tri-methylation modifications. Over the years, it has been revealed that the maintenance and dynamic changes of the bivalent domains play a vital regulatory role in the differentiation of various stem cell systems, as well as in other cells, such as immunomodulation. Tri-methylation modifications involved in the formation of the bivalent domains are interrelated and mutually regulated, thus regulating many life processes of cells. Tri-methylation of histone H3 at lysine 4 (H3K4me3), tri-methylation of histone H3 at lysine 9 (H3K9me3) and tri-methylation of histone H3 at lysine 27 (H3K27me3) are the main tri-methylation modifications involved in the formation of bivalent domains. The three form different bivalent domains in pairs. Furthermore, it is equally clear that H3K4me3 is a positive regulator of transcription and that H3K9me3/H3K27me3 are negative regulators. Enzymes related to the regulation of histone methylation play a significant role in the "homeostasis" and "breaking homeostasis" of the bivalent domains. Bivalent domains regulate target genes, upstream transcription, downstream targeting regulation and related cytokines during the establishment and breakdown of homeostasis, and exert the specific regulation of stem cells. Indeed, a unified mechanism to explain the bivalent modification in all stem cells has been difficult to define, and whether the bivalent modification is antagonistic in inducing the differentiation of homologous stem cells is controversial. In this review, we focus on the different bivalent modifications in several key stem cells and explore the main mechanisms and effects of these modifications involved. Finally, we discussed the close relationship between bivalent domains and immune cells, and put forward the prospect of the application of bivalent domains in the field of stem cells.
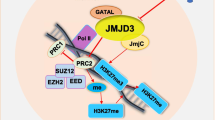
Graphical Abstract




Similar content being viewed by others
Availability of Data and Material
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Yang, X., Tian, D. C., He, W., et al. (2020). Cellular and molecular imaging for stem cell tracking in neurological diseases. Stroke Vasc Neurol. https://doi.org/10.1136/svn-2020-000408
Zamani, A. R. N., Saberianpour, S., Geranmayeh, M. H., et al. (2020). Modulatory effect of photobiomodulation on stem cell epigenetic memory: A highlight on differentiation capacity. Lasers in Medical Science, 35(2), 299–306. https://doi.org/10.1007/s10103-019-02873-7
Takahashi, K., & Yamanaka, S. (2006). Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell, 126(4), 663–676. https://doi.org/10.1016/j.cell.2006.07.024
Wu, Y., & Zhang, W. (2021). The role of E3s in regulating pluripotency of embryonic stem cells and induced pluripotent stem cells. Int J Mol Sci, 22(3), 1168. https://doi.org/10.3390/ijms22031168
Zhang, X. H., & Jin, Z. B. (2021). Patient iPSC-derived retinal organoids: Observable retinal diseases in-a-dish. Histology and Histopathology, 18307. https://doi.org/10.14670/hh-18-307
Introna, M., & Golay, J. (2020). tolerance to bone marrow transplantation: Do mesenchymal stromal cells still have a future for acute or chronic GvHD? Frontiers in Immunology, 11, 609063. https://doi.org/10.3389/fimmu.2020.609063
Danišovič, L., Varga, I., & Polák, S. (2012). Growth factors and chondrogenic differentiation of mesenchymal stem cells. Tissue and Cell, 44(2), 69–73. https://doi.org/10.1016/j.tice.2011.11.005
Murphy, M. B., Moncivais, K., & Caplan, A. I. (2013). Mesenchymal stem cells: Environmentally responsive therapeutics for regenerative medicine. Experimental & Molecular Medicine, 45(11), e54. https://doi.org/10.1038/emm.2013.94
Fukuda, K., & Shinkai, Y. (2020). SETDB1-mediated silencing of retroelements. Viruses, 12(6), 596. https://doi.org/10.3390/v12060596
Kidder, B. L., Hu, G., & Zhao, K. (2014). KDM5B focuses H3K4 methylation near promoters and enhancers during embryonic stem cell self-renewal and differentiation. Genome biology, 15(2), R32. https://doi.org/10.1186/gb-2014-15-2-r32
Wright, H., Aylwin, C. F., Toro, C. A., et al. (2021). Polycomb represses a gene network controlling puberty via modulation of histone demethylase Kdm6b expression. Scientific Reports, 11(1), 1996. https://doi.org/10.1038/s41598-021-81689-4
Matsumura, Y., Nakaki, R., Inagaki, T., et al. (2015). H3K4/H3K9me3 bivalent chromatin domains targeted by lineage-specific DNA methylation pauses adipocyte differentiation. Molecular Cell, 60(4), 584–596. https://doi.org/10.1016/j.molcel.2015.10.025
Wang, L., Jin, Q., Lee, J. E., et al. (2010). Histone H3K27 methyltransferase Ezh2 represses Wnt genes to facilitate adipogenesis. Proceedings of the National Academy of Sciences of the United States of America, 107(16), 7317–7322. https://doi.org/10.1073/pnas.1000031107
Van Nuland, R., Smits, A. H., Pallaki, P., et al. (2013). Quantitative dissection and stoichiometry determination of the human SET1/MLL histone methyltransferase complexes. Molecular and Cellular Biology, 33(10), 2067–2077. https://doi.org/10.1128/mcb.01742-12
Aoki, R., Chiba, T., Miyagi, S., et al. (2010). The polycomb group gene product Ezh2 regulates proliferation and differentiation of murine hepatic stem/progenitor cells. Journal of Hepatology, 52(6), 854–863. https://doi.org/10.1016/j.jhep.2010.01.027
Wei, H., Dong, X., You, Y., et al. (2021). OLIG2 regulates lncRNAs and its own expression during oligodendrocyte lineage formation. BMC Biology, 19(1), 132. https://doi.org/10.1186/s12915-021-01057-6
Choi, H. J., Park, J. H., Park, M., et al. (2015). UTX inhibits EMT-induced breast CSC properties by epigenetic repression of EMT genes in cooperation with LSD1 and HDAC1. EMBO Rep, 16(10), 1288–1298. https://doi.org/10.15252/embr.201540244
Le Boiteux, E., Court, F., Guichet, P. O., et al. (2021). Widespread overexpression from the four DNA hypermethylated HOX clusters in aggressive (IDHwt) glioma is associated with H3K27me3 depletion and alternative promoter usage. Molecular Oncology. https://doi.org/10.1002/1878-0261.12944
Ji, G., Zhou, W., Du, J., et al. (2021). PCGF1 promotes epigenetic activation of stemness markers and colorectal cancer stem cell enrichment. Cell Death & Disease, 12(7), 633. https://doi.org/10.1038/s41419-021-03914-2
Bernstein, B. E., Mikkelsen, T. S., Xie, X., et al. (2006). A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell, 125(2), 315–326. https://doi.org/10.1016/j.cell.2006.02.041
Howe, F. S., Fischl, H., Murray, S. C., et al. (2017). Is H3K4me3 instructive for transcription activation? BioEssays, 39(1), 1–12. https://doi.org/10.1002/bies.201600095
Barski, A., Cuddapah, S., Cui, K., et al. (2007). High-resolution profiling of histone methylations in the human genome. Cell, 129(4), 823–837. https://doi.org/10.1016/j.cell.2007.05.009
Margueron, R., & Reinberg, D. (2011). The Polycomb complex PRC2 and its mark in life. Nature, 469(7330), 343–349. https://doi.org/10.1038/nature09784
Pauler, F. M., Sloane, M. A., Huang, R., et al. (2009). H3K27me3 forms BLOCs over silent genes and intergenic regions and specifies a histone banding pattern on a mouse autosomal chromosome. Genome research, 19(2), 221–233. https://doi.org/10.1101/gr.080861.108
Zhang, J., Matsumura, Y., Kano, Y., et al. (2021). Ubiquitination dependent and independent repression of target genes by SETDB1 reveals a context dependent role for its methyltransferase activity during adipogenesis. Genes to Cells. https://doi.org/10.1111/gtc.12868
Yoo, S., & Bieda, M. C. (2014). Differences among brain tumor stem cell types and fetal neural stem cells in focal regions of histone modifications and DNA methylation, broad regions of modifications, and bivalent promoters. BMC Genomics, 15(1), 724. https://doi.org/10.1186/1471-2164-15-724
Günther, T., & Grundhoff, A. (2010). The epigenetic landscape of latent Kaposi sarcoma-associated herpesvirus genomes. PLoS Pathogens, 6(6), e1000935. https://doi.org/10.1371/journal.ppat.1000935
Cao, Z., Li, Y., Chen, Z., et al. (2015). genome-wide dynamic profiling of histone methylation during nuclear transfer-mediated porcine somatic cell reprogramming. PLoS ONE, 10(12), e0144897. https://doi.org/10.1371/journal.pone.0144897
Roth, S. Y., Denu, J. M., & Allis, C. D. (2001). Histone acetyltransferases. Annual Review of Biochemistry, 70, 81–120. https://doi.org/10.1146/annurev.biochem.70.1.81
Zhao, W., Qiao, L., Yan, S., et al. (2021). Mathematical modeling of histone modifications reveals the formation mechanism and function of bivalent chromatin. iScience, 24(7), 102732. https://doi.org/10.1016/j.isci.2021.102732
Igolkina, A. A., Zinkevich, A., Karandasheva, K. O., et al. (2019). H3K4me3, H3K9ac, H3K27ac, H3K27me3 and H3K9me3 histone tags suggest distinct regulatory evolution of open and condensed chromatin landmarks. Cells, 8(9), 1034. https://doi.org/10.3390/cells8091034
Zhang, T., Cooper, S., & Brockdorff, N. (2015). The interplay of histone modifications - writers that read. EMBO Rep, 16(11), 1467–1481. https://doi.org/10.15252/embr.201540945
Harikumar, A., & Meshorer, E. (2015). Chromatin remodeling and bivalent histone modifications in embryonic stem cells. EMBO Rep, 16(12), 1609–1619. https://doi.org/10.15252/embr.201541011
Li, F., Wan, M., Zhang, B., et al. (2018). Bivalent histone modifications and development. Current Stem Cell Research & Therapy, 13(2), 83–90. https://doi.org/10.2174/1574888x12666170123144743
Saksouk, N., Simboeck, E., & Déjardin, J. (2015). Constitutive heterochromatin formation and transcription in mammals. Epigenetics & Chromatin, 8, 3. https://doi.org/10.1186/1756-8935-8-3
Voigt, P., Tee, W. W., & Reinberg, D. (2013). A double take on bivalent promoters. Genes & Development, 27(12), 1318–1338. https://doi.org/10.1101/gad.219626.113
Piunti, A., & Shilatifard, A. (2016). Epigenetic balance of gene expression by Polycomb and COMPASS families. Science, 352(6290), aad9780. https://doi.org/10.1126/science.aad9780
Mohan, M., Herz, H. M., Smith, E. R., et al. (2011). The COMPASS family of H3K4 methylases in Drosophila. Molecular and Cellular Biology, 31(21), 4310–4318. https://doi.org/10.1128/mcb.06092-11
Bosgana, P., Nikou, S., Dimitrakopoulos, F. I., et al. (2020). H3K4 Methylation status and lysine specific methyltransferase KMT2C expression correlate with prognosis in lung adenocarcinoma. Current Molecular Pharmacology. https://doi.org/10.2174/1874467213999200831130739
Chetverina, D. A., Lomaev, D. V., & Erokhin, M. M. (2020). Polycomb and trithorax group proteins: The long road from mutations in drosophila to use in medicine. Acta Naturae, 12(4), 66–85. https://doi.org/10.32607/actanaturae.11090
Vastenhouw, N. L., & Schier, A. F. (2012). Bivalent histone modifications in early embryogenesis. Current Opinion in Cell Biology, 24(3), 374–386. https://doi.org/10.1016/j.ceb.2012.03.009
Vinogradova, M., Gehling, V. S., Gustafson, A., et al. (2016). An inhibitor of KDM5 demethylases reduces survival of drug-tolerant cancer cells. Nature Chemical Biology, 12(7), 531–538. https://doi.org/10.1038/nchembio.2085
Yu, C., Xiong, C., Tang, J., et al. (2021). Histone demethylase JMJD3 protects against renal fibrosis by suppressing TGFβ and Notch signaling and preserving PTEN expression. Theranostics, 11(6), 2706–2721. https://doi.org/10.7150/thno.48679
Leng, X., Wang, J., An, N., et al. (2020). Histone 3 lysine-27 demethylase KDM6A coordinates with KMT2B to play an oncogenic role in NSCLC by regulating H3K4me3. Oncogene, 39(41), 6468–6479. https://doi.org/10.1038/s41388-020-01449-y
Rea, S., Eisenhaber, F., O’carroll, D., et al. (2000). Regulation of chromatin structure by site-specific histone H3 methyltransferases. Nature, 406(6796), 593–599. https://doi.org/10.1038/35020506
Schultz, D. C., Ayyanathan, K., Negorev, D., et al. (2002). SETDB1: A novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes & Development, 16(8), 919–932. https://doi.org/10.1101/gad.973302
Dodge, J. E., Kang, Y. K., Beppu, H., et al. (2004). Histone H3–K9 methyltransferase ESET is essential for early development. Molecular and Cellular Biology, 24(6), 2478–2486. https://doi.org/10.1128/mcb.24.6.2478-2486.2004
Liu, S., Brind’amour, J., Karimi, M. M., et al. (2014). Setdb1 is required for germline development and silencing of H3K9me3-marked endogenous retroviruses in primordial germ cells. Genes & Development, 28(18), 2041–2055. https://doi.org/10.1101/gad.244848.114
Tan, S. L., Nishi, M., Ohtsuka, T., et al. (2012). Essential roles of the histone methyltransferase ESET in the epigenetic control of neural progenitor cells during development. Development, 139(20), 3806–3816. https://doi.org/10.1242/dev.082198
Koide, S., Oshima, M., Takubo, K., et al. (2016). Setdb1 maintains hematopoietic stem and progenitor cells by restricting the ectopic activation of nonhematopoietic genes. Blood, 128(5), 638–649. https://doi.org/10.1182/blood-2016-01-694810
Takikita, S., Muro, R., Takai, T., et al. (2016). A histone methyltransferase ESET is critical for T cell development. The Journal of Immunology, 197(6), 2269–2279. https://doi.org/10.4049/jimmunol.1502486
Collins, P. L., Kyle, K. E., Egawa, T., et al. (2015). The histone methyltransferase SETDB1 represses endogenous and exogenous retroviruses in B lymphocytes. Proc Natl Acad Sci U S A, 112(27), 8367–8372. https://doi.org/10.1073/pnas.1422187112
Pasquarella, A., Ebert, A., Pereira De Almeida, G., et al. (2016). Retrotransposon derepression leads to activation of the unfolded protein response and apoptosis in pro-B cells. Development, 143(10), 1788–1799. https://doi.org/10.1242/dev.130203
Lawson, K. A., Teteak, C. J., Gao, J., et al. (2013). ESET histone methyltransferase regulates osteoblastic differentiation of mesenchymal stem cells during postnatal bone development. FEBS Letters, 587(24), 3961–3967. https://doi.org/10.1016/j.febslet.2013.10.028
Sachs, M., Onodera, C., Blaschke, K., et al. (2013). Bivalent chromatin marks developmental regulatory genes in the mouse embryonic germline in vivo. Cell Reports, 3(6), 1777–1784. https://doi.org/10.1016/j.celrep.2013.04.032
Khromov, T., Pantakani, D. V., Nolte, J., et al. (2011). Global and gene-specific histone modification profiles of mouse multipotent adult germline stem cells. Molecular Human Reproduction, 17(3), 166–174. https://doi.org/10.1093/molehr/gaq085
Jeon, A. J., & Tucker-Kellogg, G. (2020). Bivalent genes that undergo transcriptional switching identify networks of key regulators of embryonic stem cell differentiation. BMC Genomics, 21(Suppl 10), 614. https://doi.org/10.1186/s12864-020-07009-8
Cui, P., Liu, W., Zhao, Y., et al. (2012). Comparative analyses of H3K4 and H3K27 trimethylations between the mouse cerebrum and testis. Genomics, Proteomics & Bioinformatics, 10(2), 82–93. https://doi.org/10.1016/j.gpb.2012.05.007
Li, B., Howe, L., Anderson, S., et al. (2003). The Set2 histone methyltransferase functions through the phosphorylated carboxyl-terminal domain of RNA polymerase II. Journal of Biological Chemistry, 278(11), 8897–8903. https://doi.org/10.1074/jbc.M212134200
Pray-Grant, M. G., Daniel, J. A., Schieltz, D., et al. (2005). Chd1 chromodomain links histone H3 methylation with SAGA- and SLIK-dependent acetylation. Nature, 433(7024), 434–438. https://doi.org/10.1038/nature03242
Santos-Rosa, H., Schneider, R., Bernstein, B. E., et al. (2003). Methylation of histone H3 K4 mediates association of the Isw1p ATPase with chromatin. Molecular Cell, 12(5), 1325–1332. https://doi.org/10.1016/s1097-2765(03)00438-6
Sims, R. J., 3rd., Chen, C. F., Santos-Rosa, H., et al. (2005). Human but not yeast CHD1 binds directly and selectively to histone H3 methylated at lysine 4 via its tandem chromodomains. Journal of Biological Chemistry, 280(51), 41789–41792. https://doi.org/10.1074/jbc.C500395200
Wysocka, J., Swigut, T., Milne, T. A., et al. (2005). WDR5 associates with histone H3 methylated at K4 and is essential for H3 K4 methylation and vertebrate development. Cell, 121(6), 859–872. https://doi.org/10.1016/j.cell.2005.03.036
Koche, R. P., Smith, Z. D., Adli, M., et al. (2011). Reprogramming factor expression initiates widespread targeted chromatin remodeling. Cell Stem Cell, 8(1), 96–105. https://doi.org/10.1016/j.stem.2010.12.001
Dabiri, Y., Gama-Brambila, R. A., Taškova, K., et al. (2019). Imidazopyridines as potent KDM5 demethylase inhibitors promoting reprogramming efficiency of human iPSCs. iScience, 12, 168–181. https://doi.org/10.1016/j.isci.2019.01.012
Ringrose, L., Ehret, H., & Paro, R. (2004). Distinct contributions of histone H3 lysine 9 and 27 methylation to locus-specific stability of polycomb complexes. Molecular Cell, 16(4), 641–653. https://doi.org/10.1016/j.molcel.2004.10.015
Francis, N. J., Kingston, R. E., & Woodcock, C. L. (2004). Chromatin compaction by a polycomb group protein complex. Science, 306(5701), 1574–1577. https://doi.org/10.1126/science.1100576
Liber, D., Domaschenz, R., Holmqvist, P. H., et al. (2010). Epigenetic priming of a pre-B cell-specific enhancer through binding of Sox2 and Foxd3 at the ESC stage. Cell Stem Cell, 7(1), 114–126. https://doi.org/10.1016/j.stem.2010.05.020
Relaix, F., Rocancourt, D., Mansouri, A., et al. (2005). A Pax3/Pax7-dependent population of skeletal muscle progenitor cells. Nature, 435(7044), 948–953. https://doi.org/10.1038/nature03594
Kidder, B. L., Hu, G., Yu, Z. X., et al. (2013). Extended self-renewal and accelerated reprogramming in the absence of Kdm5b. Molecular and Cellular Biology, 33(24), 4793–4810. https://doi.org/10.1128/mcb.00692-13
Xhabija, B., & Kidder, B. L. (2019). KDM5B is a master regulator of the H3K4-methylome in stem cells, development and cancer. Seminars in Cancer Biology, 57, 79–85. https://doi.org/10.1016/j.semcancer.2018.11.001
Kim, D., Patel, S. R., Xiao, H., et al. (2009). The role of PTIP in maintaining embryonic stem cell pluripotency. Stem Cells, 27(7), 1516–1523. https://doi.org/10.1002/stem.79
Lohmann, F., Loureiro, J., Su, H., et al. (2010). KMT1E mediated H3K9 methylation is required for the maintenance of embryonic stem cells by repressing trophectoderm differentiation. Stem cells (Dayton, Ohio), 28(2), 201–212. https://doi.org/10.1002/stem.278
Hattori, N., Imao, Y., Nishino, K., et al. (2007). Epigenetic regulation of Nanog gene in embryonic stem and trophoblast stem cells. Genes to Cells, 12(3), 387–396. https://doi.org/10.1111/j.1365-2443.2007.01058.x
Wang, H., An, W., Cao, R., et al. (2003). mAM facilitates conversion by ESET of dimethyl to trimethyl lysine 9 of histone H3 to cause transcriptional repression. Molecular Cell, 12(2), 475–487. https://doi.org/10.1016/j.molcel.2003.08.007
Topalovic, V., Schwirtlich, M., Stevanovic, M., et al. (2017). Histone modifications on the promoters of human OCT4 and NANOG genes at the onset of neural differentiation of NT2/D1 cells. Biochemistry (Moscow), 82(6), 715–722. https://doi.org/10.1134/s0006297917060086
Chou, R. H., Yu, Y. L., & Hung, M. C. (2011). The roles of EZH2 in cell lineage commitment. American Journal of Translation Research, 3(3), 243–250.
James, A. W., Pang, S., Askarinam, A., et al. (2012). Additive effects of sonic hedgehog and Nell-1 signaling in osteogenic versus adipogenic differentiation of human adipose-derived stromal cells. Stem Cells and Development, 21(12), 2170–2178. https://doi.org/10.1089/scd.2011.0461
Pei, L., & Tontonoz, P. (2004). Fat’s loss is bone’s gain. The Journal of Clinical Investigation, 113(6), 805–806. https://doi.org/10.1172/jci21311
Diao, S., Yang, D. M., Dong, R., et al. (2015). Enriched trimethylation of lysine 4 of histone H3 of WDR63 enhanced osteogenic differentiation potentials of stem cells from apical papilla. Journal of Endodontia, 41(2), 205–211. https://doi.org/10.1016/j.joen.2014.09.027
He, S., Yang, S., Zhang, Y., et al. (2019). LncRNA ODIR1 inhibits osteogenic differentiation of hUC-MSCs through the FBXO25/H2BK120ub/H3K4me3/OSX axis. Cell Death & Disease, 10(12), 947. https://doi.org/10.1038/s41419-019-2148-2
Rojas, A., Aguilar, R., Henriquez, B., et al. (2015). Epigenetic control of the bone-master runx2 gene during osteoblast-lineage commitment by the histone demethylase JARID1B/KDM5B. Journal of Biological Chemistry, 290(47), 28329–28342. https://doi.org/10.1074/jbc.M115.657825
Wu, H., Gordon, J. A., Whitfield, T. W., et al. (1860). (2017) Chromatin dynamics regulate mesenchymal stem cell lineage specification and differentiation to osteogenesis. Biochimica et Biophysica Acta, Gene Regulatory Mechanisms, 4, 438–449. https://doi.org/10.1016/j.bbagrm.2017.01.003
Zhang, X., Wang, W., Wang, Y., et al. (2020). Extracellular vesicle-encapsulated miR-29b-3p released from bone marrow-derived mesenchymal stem cells underpins osteogenic differentiation. Frontiers in Cell and Development Biology, 8, 581545. https://doi.org/10.3389/fcell.2020.581545
Guo, L., Guo, Y. Y., Li, B. Y., et al. (2019). Histone demethylase KDM5A is transactivated by the transcription factor C/EBPβ and promotes preadipocyte differentiation by inhibiting Wnt/β-catenin signaling. Journal of Biological Chemistry, 294(24), 9642–9654. https://doi.org/10.1074/jbc.RA119.008419
Hemming, S., Cakouros, D., Isenmann, S., et al. (2014). EZH2 and KDM6A act as an epigenetic switch to regulate mesenchymal stem cell lineage specification. Stem Cells, 32(3), 802–815. https://doi.org/10.1002/stem.1573
Isenmann, S., Arthur, A., Zannettino, A. C., et al. (2009). TWIST family of basic helix-loop-helix transcription factors mediate human mesenchymal stem cell growth and commitment. Stem Cells, 27(10), 2457–2468. https://doi.org/10.1002/stem.181
Hemming, S., Cakouros, D., Vandyke, K., et al. (2016). Identification of novel EZH2 targets regulating osteogenic differentiation in mesenchymal stem cells. Stem Cells Dev, 25(12), 909–921. https://doi.org/10.1089/scd.2015.0384
Stachecka, J., Kolodziejski, P. A., Noak, M., et al. (2021). Alteration of active and repressive histone marks during adipogenic differentiation of porcine mesenchymal stem cells. Science and Reports, 11(1), 1325. https://doi.org/10.1038/s41598-020-79384-x
Liu, F., Song, D. Y., Huang, J., et al. (2021). Long non-coding RNA CIR inhibits chondrogenic differentiation of mesenchymal stem cells by epigenetically suppressing ATOH8 via methyltransferase EZH2. Molecular Medicine, 27(1), 12. https://doi.org/10.1186/s10020-021-00272-9
Takada, I., Kouzmenko, A. P., & Kato, S. (2009). Molecular switching of osteoblastogenesis versus adipogenesis: Implications for targeted therapies. Expert Opinion on Therapeutic Targets, 13(5), 593–603. https://doi.org/10.1517/14728220902915310
Ye, L., Fan, Z., Yu, B., et al. (2012). Histone demethylases KDM4B and KDM6B promotes osteogenic differentiation of human MSCs. Cell Stem Cell, 11(1), 50–61. https://doi.org/10.1016/j.stem.2012.04.009
Wróblewski, A., Strycharz, J., Świderska, E., et al. (2021). Chronic and Transient hyperglycemia induces changes in the expression patterns of IL6 and ADIPOQ genes and their associated epigenetic modifications in differentiating human visceral adipocytes. International Journal of Molecular Sciences, 22(13), 6964. https://doi.org/10.3390/ijms22136964
Völker-Albert, M., Bronkhorst, A., Holdenrieder, S., et al. (2020). Histone modifications in stem cell development and their clinical implications. Stem Cell Reports, 15(6), 1196–1205. https://doi.org/10.1016/j.stemcr.2020.11.002
Kampilafkos, P., Melachrinou, M., Kefalopoulou, Z., et al. (2015). Epigenetic modifications in cutaneous malignant melanoma: EZH2, H3K4me2, and H3K27me3 immunohistochemical expression is enhanced at the invasion front of the tumor. American Journal of Dermatopathology, 37(2), 138–144. https://doi.org/10.1097/DAD.0b013e31828a2d54
Lin, B., Lee, H., Yoon, J. G., et al. (2015). Global analysis of H3K4me3 and H3K27me3 profiles in glioblastoma stem cells and identification of SLC17A7 as a bivalent tumor suppressor gene. Oncotarget, 6(7), 5369–5381. https://doi.org/10.18632/oncotarget.3030
Xiu, M., Wang, Y., Li, B., et al. (2021). The role of Notch3 signaling in cancer stemness and chemoresistance: Molecular mechanisms and targeting strategies. Frontiers in Molecular Biosciences, 8, 694141. https://doi.org/10.3389/fmolb.2021.694141
Chen, C., Ma, Z., & Jiang, H. (2021). EMT participates in the regulation of exosomes secretion and function in esophageal cancer cells. Technology in Cancer Research & Treatment, 20, 15330338211033076. https://doi.org/10.1177/15330338211033077
Li, Q., O’malley, M. E., Bartlett, D. L., et al. (2011). Homeobox gene Rhox5 is regulated by epigenetic mechanisms in cancer and stem cells and promotes cancer growth. Molecular Cancer, 10, 63. https://doi.org/10.1186/1476-4598-10-63
Mani, S. A., Guo, W., Liao, M. J., et al. (2008). The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell, 133(4), 704–715. https://doi.org/10.1016/j.cell.2008.03.027
Wang, H., & Unternaehrer, J. J. (2019). Epithelial-mesenchymal transition and cancer stem cells: At the crossroads of differentiation and dedifferentiation. Developmental Dynamics, 248(1), 10–20. https://doi.org/10.1002/dvdy.24678
Van Haaften, G., Dalgliesh, G. L., Davies, H., et al. (2009). Somatic mutations of the histone H3K27 demethylase gene UTX in human cancer. Nature Genetics, 41(5), 521–523. https://doi.org/10.1038/ng.349
Gui, Y., Guo, G., Huang, Y., et al. (2011). Frequent mutations of chromatin remodeling genes in transitional cell carcinoma of the bladder. Nature Genetics, 43(9), 875–878. https://doi.org/10.1038/ng.907
Dalgliesh, G. L., Furge, K., Greenman, C., et al. (2010). Systematic sequencing of renal carcinoma reveals inactivation of histone modifying genes. Nature, 463(7279), 360–363. https://doi.org/10.1038/nature08672
Lee, E., Wang, J., Yumoto, K., et al. (2016). DNMT1 regulates epithelial-mesenchymal transition and cancer stem cells, which promotes prostate cancer metastasis. Neoplasia (New York, N.Y.), 18(9), 553–566. https://doi.org/10.1016/j.neo.2016.07.007
Tan, X., Chen, S., Wu, J., et al. (2017). PI3K/AKT-mediated upregulation of WDR5 promotes colorectal cancer metastasis by directly targeting ZNF407. Cell Death & Disease, 8(3), e2686. https://doi.org/10.1038/cddis.2017.111
Shen, C., Yan, T., Tong, T., et al. (2020). ALKBH4 functions as a suppressor of colorectal cancer metastasis via competitively binding to WDR5. Front Cell Dev Biol, 8, 293. https://doi.org/10.3389/fcell.2020.00293
Wang, F., Zhang, J., Ke, X., et al. (2020). WDR5-Myc axis promotes the progression of glioblastoma and neuroblastoma by transcriptional activating CARM1. Biochemical and Biophysical Research Communications, 523(3), 699–706. https://doi.org/10.1016/j.bbrc.2019.12.101
Zhang, P., Zhang, Y., Mao, L., et al. (2009). Side population in oral squamous cell carcinoma possesses tumor stem cell phenotypes. Cancer Letters, 277(2), 227–234. https://doi.org/10.1016/j.canlet.2008.12.015
Lindner, P., Paul, S., Eckstein, M., et al. (2020). EMT transcription factor ZEB1 alters the epigenetic landscape of colorectal cancer cells. Cell Death & Disease, 11(2), 147. https://doi.org/10.1038/s41419-020-2340-4
Kanayama, K., Chiba, T., Oshima, M., et al. (2019). Genome-wide mapping of bivalent histone modifications in hepatic stem/progenitor cells. Stem Cells International, 2019, 9789240. https://doi.org/10.1155/2019/9789240
Xu, C. R., Cole, P. A., Meyers, D. J., et al. (2011). Chromatin “prepattern” and histone modifiers in a fate choice for liver and pancreas. Science (New York, N.Y.), 332(6032), 963–966. https://doi.org/10.1126/science.1202845
Chiba, T., Seki, A., Aoki, R., et al. (2010). Bmi1 promotes hepatic stem cell expansion and tumorigenicity in both Ink4a/Arf-dependent and -independent manners in mice. Hepatology, 52(3), 1111–1123. https://doi.org/10.1002/hep.23793
Ezhkova, E., Pasolli, H. A., Parker, J. S., et al. (2009). Ezh2 orchestrates gene expression for the stepwise differentiation of tissue-specific stem cells. Cell, 136(6), 1122–1135. https://doi.org/10.1016/j.cell.2008.12.043
Mochizuki-Kashio, M., Mishima, Y., Miyagi, S., et al. (2011). Dependency on the polycomb gene Ezh2 distinguishes fetal from adult hematopoietic stem cells. Blood, 118(25), 6553–6561. https://doi.org/10.1182/blood-2011-03-340554
Murdaugh, R. L., Hoegenauer, K. A., Kitano, A., et al. (2021). The histone H3.3 chaperone HIRA restrains erythroid-biased differentiation of adult hematopoietic stem cells. Stem Cell Reports. https://doi.org/10.1016/j.stemcr.2021.06.009
Lee, G. R. (2018). The balance of Th17 versus Treg cells in autoimmunity. International Journal of Molecular Sciences, 19(3), 730. https://doi.org/10.3390/ijms19030730
Thomas, R. M., Sai, H., & Wells, A. D. (2012). Conserved intergenic elements and DNA methylation cooperate to regulate transcription at the il17 locus. Journal of Biological Chemistry, 287(30), 25049–25059. https://doi.org/10.1074/jbc.M112.351916
Cribbs, A. P., Terlecki-Zaniewicz, S., Philpott, M., et al. (2020). Histone H3K27me3 demethylases regulate human Th17 cell development and effector functions by impacting on metabolism. Proceedings of National Academy of Sciences of the United States of America, 117(11), 6056–6066. https://doi.org/10.1073/pnas.1919893117
Chen, Q., Duan, X., Xu, M., et al. (2020). BMSC-EVs regulate Th17 cell differentiation in UC via H3K27me3. Molecular Immunology, 118, 191–200. https://doi.org/10.1016/j.molimm.2019.12.019
Pereira, R. M., Martinez, G. J., Engel, I., et al. (2014). Jarid2 is induced by TCR signalling and controls iNKT cell maturation. Nature Communications, 5, 4540. https://doi.org/10.1038/ncomms5540
Xiao, X., Shi, X., Fan, Y., et al. (2016). The costimulatory receptor OX40 inhibits interleukin-17 expression through activation of repressive chromatin remodeling pathways. Immunity, 44(6), 1271–1283. https://doi.org/10.1016/j.immuni.2016.05.013
Adoue, V., & Joffre, O. (2020). Endogenous retroviruses: Friend or foe of the immune system? Medecine Sciences : M/S, 36(3), 253–260. https://doi.org/10.1051/medsci/2020022
Kaleviste, E., Saare, M., Leahy, T. R., et al. (2019). Interferon signature in patients with STAT1 gain-of-function mutation is epigenetically determined. European Journal of Immunology, 49(5), 790–800. https://doi.org/10.1002/eji.201847955
Lin, F., Meng, X., Guo, Y., et al. (2019). Epigenetic initiation of the T(H)17 differentiation program is promoted by Cxxc finger protein 1. Sciences Advances, 5(10), eaax1608. https://doi.org/10.1126/sciadv.aax1608
Li, Q., Zou, J., Wang, M., et al. (2014). Critical role of histone demethylase Jmjd3 in the regulation of CD4+ T-cell differentiation. Nature Communications, 5, 5780. https://doi.org/10.1038/ncomms6780
Liu, X., Ren, S., Qu, X., et al. (2015). Mesenchymal stem cells inhibit Th17 cells differentiation via IFN-γ-mediated SOCS3 activation. Immunologic Research, 61(3), 219–229. https://doi.org/10.1007/s12026-014-8612-2
Mi, Q. S., Wang, J., Liu, Q., et al. (2021). microRNA dynamic expression regulates invariant NKT cells. Cellular and Molecular Life Sciences. https://doi.org/10.1007/s00018-021-03895-7
Dobenecker, M. W., Kim, J. K., Marcello, J., et al. (2015). Coupling of T cell receptor specificity to natural killer T cell development by bivalent histone H3 methylation. Journal of Experimental Medicine, 212(3), 297–306. https://doi.org/10.1084/jem.20141499
Acknowledgements
This study was funded by the National Natural Science Foundation of China (Grant No. 31201052). The authors confirm independence from the sponsors; the content of the article has not been influenced by the sponsors.
Funding
This study was funded by the National Natural Science Foundation of China (Grant No. 31201052). The authors confirm independence from the sponsors; the content of the article has not been influenced by the sponsors.
Author information
Authors and Affiliations
Contributions
Han Sun, Feng Ji, Ying Wang and An Wang participated in the writing. Han Sun, Ying Wang and Feng Ji designed and prepared the figures. An Wang and Han Sun designed and prepared the tables. Yin Wang and Xu He were in charge of proofreading manuscripts. Lisha Li, Xu He and Ming Yang designed and polished the paper.
Corresponding authors
Ethics declarations
Ethical Approval
Not applicable.
Open Access
This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licenses, and indicate if changes were made. The images or other third-party material in this article are included in the article’s Creative Commons licenses, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licenses and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licenses, visit http://creativecommons.org/licenses/by/4.0/.
Competing Interests
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sun, H., Wang, Y., Wang, Y. et al. Bivalent Regulation and Related Mechanisms of H3K4/27/9me3 in Stem Cells. Stem Cell Rev and Rep 18, 165–178 (2022). https://doi.org/10.1007/s12015-021-10234-7
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-021-10234-7