Abstract
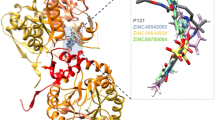
Cryptosporidiosis accounts for a surge in infant (<5 years) mortality and morbidity. To date, several drug discovery efforts have been put in place to develop effective therapeutic options against the causative parasite. Based on a recent report, P131 spares inosine monophosphate dehydrogenase (IMPDH) in a eukaryotic model (mouse IMPDH (mIMPDH)) while binding selectively to the NAD+ site in Cryptosporidium parvum (CpIMPDH). However, no structural detail exists on the underlining mechanisms of P131-CpIMPDH selective targeting till date. To this effect, we investigate the selective inhibitory dynamics of P131 in CpIMPDH relative to mIMPDH via molecular biocomputation methods. Pairwise sequence alignment revealed prominent variations at the NAD+ binding regions of both proteins that accounted for disparate P131 binding activities. The influence of these variations was further revealed by the MM/PBSA energy estimations coupled with per-residue energy decomposition which monitored the systematic binding of the compound. Furthermore, relative high-affinity interactions occurred at the CpIMPDH NAD+ site which were majorly mediated by SER22, VAL24, PRO26, SER354, GLY357, and TYR358 located on chain D. These residues are unique to the parasite IMPDH form and not in the eukaryotic protein, highlighting variations that account for preferential P131 binding. Molecular insights provided herein corroborate previous experimental reports and further underpin the basis of CpIMPDH inhibitor selectivity. Findings from this study could present attractive prospects toward the design of novel anticryptosporidials with improved selectivity and binding affinity against parasitic targets.









Similar content being viewed by others
References
Bouzid, M., Kintz, E., & Hunter, P. R. (2018). Risk factors for Cryptosporidium infection in low and middle income countries: a systematic review and meta-analysis. PLoS Neglected Tropical Diseases, 12(6). https://doi.org/10.1371/journal.pntd.0006553.
Fischer Walker, C. L., Aryee, M. J., Boschi-Pinto, C., & Black, R. E. (2012). Estimating diarrhea mortality among young children in low and middle income countries. PLoS ONE. https://doi.org/10.1371/journal.pone.0029151.
Valentiner-Branth, P., Steinsland, H., Fischer, T. K., Perch, M., Scheutz, F., Dias, F., & Sommerfelt, H. (2003). Cohort study of Guinean children: incidence, pathogenicity, conferred protection, and attributable risk for enteropathogens during the first 2 years of life. Journal of Clinical Microbiology, 41(9), 4238–4245. https://doi.org/10.1128/JCM.41.9.4238-4245.2003.
Liu, L., Johnson, H. L., Cousens, S., Perin, J., Scott, S., Lawn, J. E., & Black, R. E. (2012). Global, regional, and national causes of child mortality: an updated systematic analysis for 2010 with time trends since 2000. The Lancet, 379(9832), 2151–2161. https://doi.org/10.1016/S0140-6736(12)60560-1.
Kotloff, K. L., Nataro, J. P., Blackwelder, W. C., Nasrin, D., Farag, T. H., Panchalingam, S., & Levine, M. M. (2013). Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): a prospective, case-control study. The Lancet, 382(9888), 209–222. https://doi.org/10.1016/S0140-6736(13)60844-2.
Sow, S. O., Muhsen, K., Nasrin, D., Blackwelder, W. C., Wu, Y., & Farag, T. H. et al. (2016). The burden of cryptosporidium diarrheal disease among children <24 months of age in moderate/high mortality regions of sub-Saharan Africa and South Asia, utilizing data from the global enteric multicenter study (GEMS). PLoS Neglected Tropical Diseases, 10(5), e0004729. https://doi.org/10.1371/journal.pntd.0004729.
Odeniran, P. O., & Ademola, I. O. (2019). Epidemiology of Cryptosporidium infection in different hosts in Nigeria: a meta-analysis. Parasitology International, 71(May), 194–206. https://doi.org/10.1016/j.parint.2019.04.007.
Delahoy, M. J., Omore, R., Ayers, T. L., Schilling, K. A., Blackstock, A. J., Ochieng, J. B., … O’Reilly, C. E. (2018). Clinical, environmental, and behavioral characteristics associated with Cryptosporidium infection among children with moderate-to-severe diarrhea in rural western Kenya, 2008–2012: The Global Enteric Multicenter Study (GEMS). PLoS Neglected Tropical Diseases, 12(7). https://doi.org/10.1371/journal.pntd.0006640.
Umejiego, N. N., Li, C., Riera, T., Hedstrom, L., & Striepen, B. (2004). Cryptosporidium parvum IMP dehydrogenase: identification of functional, structural, and dynamic properties that can be exploited for drug design. Journal of Biological Chemistry, 279(39), 40320–40327. https://doi.org/10.1074/jbc.M407121200.
Putignani, L., & Menichella, D. (2010). Global distribution, public health and clinical impact of the protozoan pathogen cryptosporidium. Interdisciplinary Perspectives on Infectious Diseases, 2010. https://doi.org/10.1155/2010/753512.
O’connor, R. M., Shaffie, R., Kang, G., & Ward, H. D. (2011). Cryptosporidiosis in patients with HIV/AIDS. AIDS, 25(5), 549–60. https://doi.org/10.1097/QAD.0b013e3283437e88.
Chalmers, R. M., & Davies, A. P. (2010). Minireview: clinical cryptosporidiosis. Experimental Parasitology, 124(1), 138–46. https://doi.org/10.1016/j.exppara.2009.02.003.
Kang, G., Sarkar, R., & Desai, N. (2012). Cryptosporidiosis: an under-recognized public health problem. Tropical Parasitology, 2(2), 91 https://doi.org/10.4103/2229-5070.105173.
Mor, S., & Tzipori, S. (2008). Cryptosporidiosis in children in sub-Saharan Africa: A lingering challenge. Clinical Infectious Diseases, 47(7), 915–921. https://doi.org/10.1086/591539.
Aldeyarbi, H. M., Abu El-Ezz, N. M. T., & Karanis, P. (2016). Cryptosporidium and cryptosporidiosis: the African perspective. Environmental Science and Pollution Research, 23(14), 13811–13821. https://doi.org/10.1007/s11356-016-6746-6.
Abdool Karim, S. S., Churchyard, G. J., Karim, Q. A., & Lawn, S. D. (2009). HIV infection and tuberculosis in South Africa: an urgent need to escalate the public health response. Lancet, 374(9693), 921–33. https://doi.org/10.1016/S0140-6736(09)60916-8.
Checkley, W., White Jr, A. C., & Jaganath, D. et al. (2015). A review of the global burden, novel diagnostics, therapeutics, and vaccine targets for cryptosporidium. The Lancet Infectious Diseases, 15(1), 85–94. https://doi.org/10.1016/s1473-3099(14)70772-8.
Mead, J. R. (2002). Cryptosporidiosis and the challenges of chemotherapy. Drug Resistance Updates: Reviews and Commentaries in Antimicrobial and Anticancer Chemotherapy, 5(1), 47–57. http://www.ncbi.nlm.nih.gov/pubmed/12127863.
Lee, S., Harwood, M., Girouard, D., Meyers, M. J., Campbell, M. A., Beamer, G., & Tzipori, S. (2017). The therapeutic efficacy of azithromycin and nitazoxanide in the acute pig model of Cryptosporidium hominis. PLoS ONE, 12(10), e0185906. https://doi.org/10.1371/journal.pone.0185906.
Cabada, M. M., & White, A. C. (2010). Treatment of cryptosporidiosis: do we know what we think we know? Current Opinion in Infectious Diseases, 23(5), 494–9. https://doi.org/10.1097/QCO.0b013e32833de052.
Sparks, H., Nair, G., Castellanos-Gonzalez, A., & White, A. C. (2015). Treatment of Cryptosporidium: what we know, gaps, and the way forward. Current Tropical Medicine Reports, 2(3), 181–187. https://doi.org/10.1007/s40475-015-0056-9.
Fan, X., Upadhyaya, B., Wu, L., Koh, C., Santín-Durán, M., Pittaluga, S., & Jain, A. (2012). CD40 agonist antibody mediated improvement of chronic Cryptosporidium infection in patients with X-linked hyper IgM syndrome. Clinical Immunology, 143(2), 152–161. https://doi.org/10.1016/j.clim.2012.01.014.
Hewitt, R. G., Yiannoutsos, C. T., Higgs, E. S., Carey, J. T., Geiseler, P. J., Soave, R., & Bender, J. F. (2000). Paromomycin: no more effective than placebo for treatment of cryptosporidiosis in patients with advanced human immunodeficiency virus infection. AIDS Clinical Trial Group. Clinical Infectious Diseases, 31(4), 1084–92. https://doi.org/10.1086/318155.
Hussien, S. M. M., Abdella, O. H., Abu-Hashim, A. H., Aboshiesha, G. A., Taha, M. A. A., El-Shemy, A. S., & El-Bader, M. M. (2013). Comparative study between the effect of nitazoxanide and paromomycine in treatment of cryptosporidiosis in hospitalized children. Journal of the Egyptian Society of Parasitology, 43(2), 463–470. http://www.ncbi.nlm.nih.gov/pubmed/24260825.
Allam, A. F., & Shehab, A. Y. (2002). Efficacy of azithromycin, praziquantel and mirazid in treatment of cryptosporidiosis in school children. Journal of the Egyptian Society of Parasitology, 32(3), 969–978. http://www.ncbi.nlm.nih.gov/pubmed/12512828.
Raja, K., Abbas, Z., Hassan, S. M., Luck, N. H., Aziz, T., & Mubarak, M. (2014). Prevalence of cryptosporidiosis in renal transplant recipients presenting with acute diarrhea at a single center in Pakistan. Journal of Nephropathology, 3(4), 127–131. https://doi.org/10.12860/jnp.2014.25.
Holmberg, S. D., Moorman, A. C., Von Bargen, J. C., Palella, F. J., Loveless, M. O., Ward, D. J., & Navin, T. R. (1998). Possible effectiveness of clarithromycin and rifabutin for cryptosporidiosis chemoprophylaxis in HIV disease. HIV Outpatient Study (HOPS) Investigators. JAMA, 279(5), 384–386. https://doi.org/10.1001/jama.279.5.384.
Amenta, M., Dalle Nogare, E. R., Colomba, C., Prestileo, T. S., Di Lorenzo, F., Fundaro, S., & Ferrieri, A. (1999). Intestinal protozoa in HIV-infected patients: effect of rifaximin in Cryptosporidium parvum and Blastocystis hominis infections. Journal of Chemotherapy, 11(5), 391–395. https://doi.org/10.1179/joc.1999.11.5.391.
Gathe, J. C., Mayberry, C., Clemmons, J., & Nemecek, J. (2008). Resolution of severe cryptosporidial diarrhea with rifaximin in patients with AIDS. Journal of Acquired Immune Deficiency Syndromes, 48(3), 363–364. https://doi.org/10.1097/QAI.0b013e31817beb78.
Legrand, F., Grenouillet, F., Larosa, F., Dalle, F., Saas, P., Millon, L., & Rohrlich, P. S. (2011). Diagnosis and treatment of digestive cryptosporidiosis in allogeneic haematopoietic stem cell transplant recipients: a prospective single centre study. Bone Marrow Transplantation, 46(6), 858–862. https://doi.org/10.1038/bmt.2010.200.
Bonatti, H., Barroso, L. F., Sawyer, R. G., Kotton, C. N., & Sifri, C. D. (2012). Cryptosporidium enteritis in solid organ transplant recipients: Multicenter retrospective evaluation of 10 cases reveals an association with elevated tacrolimus concentrations. Transplant Infectious Disease, 14(6), 635–648. https://doi.org/10.1111/j.1399-3062.2012.00719.x.
Krause, I., Amir, J., Cleper, R., Dagan, A., Behor, J., Samra, Z., & Davidovits, M. (2012). Cryptosporidiosis in children following solid organ transplantation. Pediatric Infectious Disease Journal, 31(11), 1135–1138. https://doi.org/10.1097/INF.0b013e31826780f7.
Chen, L., Wilson, D. J., Xu, Y., Aldrich, C. C., Felczak, K., Sham, Y. Y., & Pankiewicz, K. W. (2010). Triazole-linked inhibitors of inosine monophosphate dehydrogenase from human and mycobacterium tuberculosis. Journal of Medicinal Chemistry, 53(12), 4768–4778. https://doi.org/10.1021/jm100424m.
Gollapalli, D. R., MacPherson, I. S., Liechti, G., Gorla, S. K., Goldberg, J. B., & Hedstrom, L. (2010). Structural determinants of inhibitor selectivity in prokaryotic IMP dehydrogenases. Chemistry and Biology, 17(10), 1084–1091. https://doi.org/10.1016/j.chembiol.2010.07.014.
Hedstrom, L., Liechti, G., Goldberg, J. B., & Gollapalli, D. R. (2011). The antibiotic potential of prokaryotic IMP dehydrogenase inhibitors. Current Medicinal Chemistry, 18(13), 1909–18. https://doi.org/10.2174/092986711795590129.
Usha, V., Hobrath, J. V., Gurcha, S. S., Reynolds, R. C., & Besra, G. S. (2012). Identification of novel Mt-Guab2 inhibitor series active against M. tuberculosis. PLoS ONE, 7(3). https://doi.org/10.1371/journal.pone.0033886.
Wang, W., & Hedstrom, L. (1997). Kinetic mechanism of human inosine 5’-monophosphate dehydrogenase type II: random addition of substrates and ordered release of products. Biochemistry, 36(28), 8479–8483. https://doi.org/10.1021/bi970226n.
Zimmermann, A., Gu, J. J., Spychala, J., & Mitchell, B. S. (1996). Inosine monophosphate dehydrogenase expression: transcriptional regulation of the type I and type II genes. In Advances in Enzyme Regulation (Vol. 36, pp. 75–84). Elsevier Ltd. https://doi.org/10.1016/0065-2571(95)00012-7.
Striepen, B., Pruijssers, A. J. P., Huang, J., Li, C., Gubbels, M. J., Umejiego, N. N., & Kissinger, J. C. (2004). Gene transfer in the evolution of parasite nucleotide biosynthesis. Proceedings of the National Academy of Sciences of the United States of America, 101(9), 3154–3159. https://doi.org/10.1073/pnas.0304686101.
Zhang, R. G., Evans, G., Rotella, F. J., Westbrook, E. M., Beno, D., Huberman, E., & Collart, F. R. (1999). Characteristics and crystal structure of bacterial inosine-5’-monophosphate dehydrogenase. Biochemistry, 38(15), 4691–4700. https://doi.org/10.1021/bi982858v.
Kim, Y., Makowska-Grzyska, M., Gorla, S. K., Gollapalli, D. R., Cuny, G. D., Joachimiak, A., & Hedstrom, L. (2015). Structure of Cryptosporidium IMP dehydrogenase bound to an inhibitor with in vivo antiparasitic activity. Acta Crystallographica Section F: Structural Biology Communications, 71, 531–538. https://doi.org/10.1107/S2053230X15000187.
Makowska-Grzyska, M., Kim, Y., Maltseva, N., Osipiuk, J., Gu, M., Zhang, M., & Joachimiak, A. (2015). A novel cofactor-binding mode in bacterial IMP dehydrogenases explains inhibitor selectivity. Journal of Biological Chemistry, 290(9), 5893–5911. https://doi.org/10.1074/jbc.M114.619767.
Hedstrom, L. (2009). IMP dehydrogenase: structure, mechanism, and inhibition. Chemical Reviews, 109(7), 2903–2928. https://doi.org/10.1021/cr900021w.
Felczak, K., Chen, L., Wilson, D., Williams, J., Vince, R., Petrelli, R., & Pankiewicz, K. W. (2011). Cofactor-type inhibitors of inosine monophosphate dehydrogenase via modular approach: Targeting the pyrophosphate binding sub-domain. Bioorganic and Medicinal Chemistry, 19(5), 1594–1605. https://doi.org/10.1016/j.bmc.2011.01.042.
Colby, T. D., Vanderveen, K., Strickler, M. D., Markham, G. D., & Goldstein, B. M. (1999). Crystal structure of human type II inosine monophosphate dehydrogenase: implications for ligand binding and drug design. Proceedings of the National Academy of Sciences of the United States of America, 96(7), 3531–3536. https://doi.org/10.1073/pnas.96.7.3531.
Gorla, S. K., Kavitha, M., Zhang, M., Liu, X., Sharling, L., Gollapalli, D. R., & Cuny, G. D. (2012). Selective and potent urea inhibitors of Cryptosporidium parvum inosine 5′-monophosphate dehydrogenase. Journal of Medicinal Chemistry, 55(17), 7759–7771. https://doi.org/10.1021/jm3007917.
Macpherson, I. S., Kirubakaran, S., Gorla, S. K., Riera, T. V., D’Aquino, J. A., Zhang, M., & Hedstrom, L. (2010). The structural basis of Cryptosporidium-specific IMP dehydrogenase inhibitor selectivity. Journal of the American Chemical Society, 132(4), 1230–1. https://doi.org/10.1021/ja909947a.
Gorla, S. K., McNair, N. N., Yang, G., Gao, S., Hu, M., Jala, V. R., & Hedstrom, L. (2014). Validation of IMP dehydrogenase inhibitors in a mouse model of cryptosporidiosis. Antimicrobial Agents and Chemotherapy, 58(3), 1603–1614. https://doi.org/10.1128/AAC.02075-13.
Johnson, C. R., Gorla, S. K., Kavitha, M., Zhang, M., Liu, X., Striepen, B., & Hedstrom, L. (2013). Phthalazinone inhibitors of inosine-5’-monophosphate dehydrogenase from Cryptosporidium parvum. Bioorganic & Medicinal Chemistry Letters, 23(4), 1004–1007. https://doi.org/10.1016/j.bmcl.2012.12.037.
Kirubakaran, S., Gorla, S. K., Sharling, L., Zhang, M., Liu, X., Ray, S. S., & Cuny, G. D. (2012). Structure-activity relationship study of selective benzimidazole-based inhibitors of Cryptosporidium parvum IMPDH. Bioorganic & Medicinal Chemistry Letters, 22(5), 1985–1988. https://doi.org/10.1016/j.bmcl.2012.01.029.
Maurya, S. K., Gollapalli, D. R., Kirubakaran, S., Zhang, M., Johnson, C. R., Benjamin, N. N., & Cuny, G. D. (2009). Triazole inhibitors of Cryptosporidium parvum inosine 5’-monophosphate dehydrogenase. Journal of Medicinal Chemistry, 52(15), 4623–4630. https://doi.org/10.1021/jm900410u.
Sharling, L., Liu, X., Gollapalli, D. R., Maurya, S. K., Hedstrom, L., & Striepen, B. (2010). A screening pipeline for antiparasitic agents targeting Cryptosporidium inosine monophosphate dehydrogenase. PLoS Neglected Tropical Diseases, 4(8), e794. https://doi.org/10.1371/journal.pntd.0000794.
Umejiego, N. N., Gollapalli, D., Sharling, L., Volftsun, A., Lu, J., Benjamin, N. N., & Hedstrom, L. (2008). Targeting a prokaryotic protein in a eukaryotic pathogen: identification of lead compounds against cryptosporidiosis. Chemistry & Biology, 15(1), 70–77. https://doi.org/10.1016/j.chembiol.2007.12.010.
Sun, Z., Khan, J., Makowska-Grzyska, M., Zhang, M., Cho, J. H., Suebsuwong, C., & Cuny, G. D. (2014). Synthesis, in vitro evaluation and cocrystal structure of 4-oxo-[1]benzopyrano[4,3-c]pyrazole Cryptosporidium parvum inosine 5’-monophosphate dehydrogenase (CpIMPDH) inhibitors. Journal of Medicinal Chemistry, 57(24), 10544–10550. https://doi.org/10.1021/jm501527z.
Gorla, S. K., Kavitha, M., Zhang, M., Chin, J. E. W., Liu, X., Striepen, B., & Cuny, G. D. (2013). Optimization of benzoxazole-based inhibitors of Cryptosporidium parvum inosine 5’-monophosphate dehydrogenase. Journal of Medicinal Chemistry, 56(10), 4028–4043. https://doi.org/10.1021/jm400241j.
Sievers, F., Wilm, A., Dineen, D., Gibson, T. J., Karplus, K., Li, W., … Higgins, D. G. (2011). Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Molecular Systems Biology, 7. https://doi.org/10.1038/msb.2011.75.
Abrahamsen, M. S., Templeton, T. J., Enomoto, S., Abrahante, J. E., Zhu, G., Lancto, C. A., & Kapur, V. (2004). Complete genome sequence of the apicomplexan, Cryptosporidium parvum. Science, 304(5669), 441–445. https://doi.org/10.1126/science.1094786.
Tiedeman, A. A., & Smith, J. M. (1991). Isolation and sequence of a cDNA encoding mouse IMP dehydrogenase. Gene, 97(2), 289–293. https://doi.org/10.1016/0378-1119(91)90065-j.
Waterhouse, A., Bertoni, M., Bienert, S., Studer, G., Tauriello, G., Gumienny, R., & Schwede, T. (2018). SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Research, 46(W1), W296–W303. https://doi.org/10.1093/nar/gky427.
Sintchak, M. D., Fleming, M. A., Futer, O., Raybuck, S. A., Chambers, S. P., Caron, P. R., & Wilson, K. P. (1996). Structure and mechanism of inosine monophosphate dehydrogenase in complex with the immunosuppressant mycophenolic acid. Cell, 85(6), 921–930. https://doi.org/10.1016/S0092-8674(00)81275-1.
Wiederstein, M., & Sippl, M. J. (2007). ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Research, 35(Web Server), W407–W410. https://doi.org/10.1093/nar/gkm290.
Eisenberg, D., Lüthy, R., & Bowie, J. U. (1997). VERIFY3D: assessment of protein models with three-dimensional profiles. Methods in Enzymology, 277, 396–404.
Gopalakrishnan, K., Sowmiya, G., Sheik, S. S., & Sekar, K. (2007). Ramachandran plot on the web (2.0). Protein & Peptide Letters, 14(7), 669–671. https://doi.org/10.2174/092986607781483912.
Pettersen, E. F., Goddard, T. D., Huang, C. C., Couch, G. S., Greenblatt, D. M., Meng, E. C., & Ferrin, T. E. (2004). UCSF Chimera—a visualization system for exploratory research and analysis. Journal of Computational Chemistry, 25(13), 1605–1612. https://doi.org/10.1002/jcc.20084.
Eswar, N., Webb, B., Marti-Renom, M. A., Madhusudhan, M. S., Eramian, D., Shen, M.-Y., & Sali, A. (2006). Comparative protein structure modeling using Modeller. Current Protocols in Bioinformatics, 5, Unit-5.6 https://doi.org/10.1002/0471250953.bi0506s15.
Trott, O., & Olson, A. J. (2010). Software news and update AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. Journal of Computational Chemistry, 31(2), 455–461. https://doi.org/10.1002/jcc.21334.
Wang, Z., Sun, H., Yao, X., Li, D., Xu, L., Li, Y., & Hou, T. (2016). Comprehensive evaluation of ten docking programs on a diverse set of protein-ligand complexes: the prediction accuracy of sampling power and scoring power. Physical Chemistry Chemical Physics, 18(18), 12964–12975. https://doi.org/10.1039/c6cp01555g.
Machaba, K. E., Mhlongo, N. N., & Soliman, M. E. S. (2018). Induced mutation proves a potential target for TB therapy: a molecular dynamics study on LprG. Cell Biochemistry and Biophysics, 76(3), 345–356. https://doi.org/10.1007/s12013-018-0852-7.
Oguntade, S., Ramharack, P., & Soliman, M. E. (2017). Characterizing the ligand-binding landscape of Zika NS3 helicase-promising lead compounds as potential inhibitors. Future Virology, 12(6), 261–273. https://doi.org/10.2217/fvl-2017-0014.
Olotu, F. A., & Soliman, M. E. S. (2018). From mutational inactivation to aberrant gain-of-function: Unraveling the structural basis of mutant p53 oncogenic transition. Journal of Cellular Biochemistry, 119(3), 2646–2652. https://doi.org/10.1002/jcb.26430.
Agoni, C., Munsamy, G., Ramhrack, P., & Soliman, M. (2020). Human rhinovirus inhibition through capsid “Canyon” perturbation: structural insights into the role of a novel benzothiophene derivative. Cell Biochemistry and Biophysics, 78, 3–13.
Agoni, C., Salifu, E. Y., Munsamy, G., Olotu, F. A., & Soliman, M. (2019). CF3‐pyridinyl substitution on anti‐malarial therapeutics: probing differential ligand binding and dynamical inhibitory effects of a novel triazolopyrimidine‐based inhibitor on Plasmodium falciparum Dihydroorotate dehydrogenase. Chemistry & Biodiversity. https://doi.org/10.1002/cbdv.201900365.
Olotu, F. A., & Soliman, M. E. S. (2019). Dynamic perspectives into the mechanisms of mutation-induced p53-DNA binding loss and inactivation using active perturbation theory: structural and molecular insights toward the design of potent reactivators in cancer therapy. Journal of Cellular Biochemistry, 120(1), 951–966. https://doi.org/10.1002/jcb.27458.
Nair, P. C., & Miners, J. O. (2014). Molecular dynamics simulations: from structure function relationships to drug discovery. In Silico Pharmacology, 2(1). https://doi.org/10.1186/s40203-014-0004-8.
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A., & Case, D. A. (2004). Development and testing of a general amber force field. Journal of Computational Chemistry, 25(9), 1157–1174. https://doi.org/10.1002/jcc.20035.
Grest, G. S., & Kremer, K. (1986). Molecular dynamics simulation for polymers in the presence of a heat bath. Physical Review A, 33(5), 3628–3631. https://doi.org/10.1103/PhysRevA.33.3628.
Berendsen, H. J. C., Postma, J. P. M., Van Gunsteren, W. F., Dinola, A., & Haak, J. R. (1984). Molecular dynamics with coupling to an external bath. The Journal of Chemical Physics, 81(8), 3684–3690. https://doi.org/10.1063/1.448118.
Ryckaert, J.-P., Ciccotti, G., & Berendsen, H. J. C. (1977). Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes.Journal of Computational Physics, 23(3), 327–341.
Roe, D. R., & Cheatham, T. E. (2013). PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. Journal of Chemical Theory and Computation, 9(7), 3084–3095. https://doi.org/10.1021/ct400341p.
Seifert, E. (2014). OriginPro 9.1: scientific data analysis and graphing software-software review. Journal of Chemical Information and Modeling, 54(5), 1552. https://doi.org/10.1021/ci500161d.
Genheden, S., & Ryde, U. (2015). The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opinion on Drug Discovery, 10(5), 449–461. https://doi.org/10.1517/17460441.2015.1032936.
Hou, T., Wang, J., Li, Y., & Wang, W. (2011). Assessing the performance of the MM/PBSA and MM/GBSA methods. 1. The accuracy of binding free energy calculations based on molecular dynamics simulations. Journal of Chemical Information and Modeling, 51(1), 69–82. https://doi.org/10.1021/ci100275a.
Chaudhary, N., & Aparoy, P. (2017). Deciphering the mechanism behind the varied binding activities of COXIBs through Molecular Dynamic Simulations, MM-PBSA binding energy calculations and per-residue energy decomposition studies. Journal of Biomolecular Structure & Dynamics, 35(4), 868–882. https://doi.org/10.1080/07391102.2016.1165736.
Gupta, A., Chaudhary, N., & Aparoy, P. (2018). MM-PBSA and per-residue decomposition energy studies on 7-Phenyl-imidazoquinolin-4(5H)-one derivatives: identification of crucial site points at microsomal prostaglandin E synthase-1 (mPGES-1) active site. International Journal of Biological Macromolecules, 119, 352–359. https://doi.org/10.1016/j.ijbiomac.2018.07.050.
Beg, A., Khan, F., Lobb, K., Islam, A., Ahmad, F., & Hassan, M. (2019). High throughput screening, docking, and molecular dynamics studies to identify potential inhibitors of human calcium/calmodulin-dependent protein kinase IV. Journal of Biomolecular Structure & Dynamics, 37, 2179–2192.
Dahiya, R., Mohammad, T., Roy, S., Anwar, S., Gupta, P., Haque, A., & Ahmad, F. (2019). Investigation of inhibitory potential of quercetin to the pyruvate dehydrogenase kinase 3: towards implications in anticancer therapy. International Journal of Biological Macromolecules, 136, 1076–1085.
Fatima, S., Mohammad, T., Jairajpuri, D., Rehman, M., Hussain, A., Samim, M., & … Hassan, M. (2019). Identification and evaluation of glutathione conjugate gamma-l-glutamyl-l-cysteine for improved drug delivery to the brain. Journal of Biomolecular Structure & Dynamics, 38(12), 3610–3620.
Gulzar, M., Ali, S., Khan, F., Khan, P., Taneja, P., & Hassan, M. (2019). Binding mechanism of ca eic acid and simvastatin to the integrin linked kinase for therapeutic implications: a comparative docking and MD simulation studies. Journal of Biomolecular Structure & Dynamics, 37, 4327–4337.
Kuzmanic, A., & Zagrovic, B. (2010). Determination of ensemble-average pairwise root mean square deviation from experimental B-factors. Biophysical Journal, 98, 861–871.
Naz, F., Shahbaaz, M., Bisetty, K., Islam, A., Ahmad, F., & Hassan, M. (2015). Designing new kinase inhibitor derivatives as therapeutics against common complex diseases: structural basis of microtubule affinity-regulating kinase 4 (MARK4) inhibition. OMICS, 19, 700–711.
Naz, F., Shahbaaz, M., Khan, S., Bisetty, K., Islam, A., Ahmad, F., & Hassan, M. (2015). PKR-inhibitor binds efficiently with human microtubule affinity-regulating kinase 4. Journal of Molecular Graphics and Modelling, 62, 245–252.
Naz, H., Shahbaaz, M., Haque, M., Bisetty, K., Islam, A., Ahmad, F., & Hassan, M. (2017). Urea-induced denaturation of human calcium/calmodulin-dependent protein kinase IV: a combined spectroscopic and MD simulation studies. Journal of Biomolecular Structure & Dynamics, 35, 463–475.
Menendez, C., Mazola, Y., Guirola, O., Palomares, S., Chinea, G., Hernandez, L., & Musacchio, A. (2015). A comparative molecular dynamics study of thermophilic and mesophilic beta-fructosidase enzymes. Journal of Molecular Modeling, 21, 2772.
Ali, S., Hassan, M., Islam, A., & Ahmad, F. (2014). A review of methods available to estimate solvent-accessible surface areas of soluble proteins in the folded and unfolded states. Current Protein & Peptide Science, 15, 456–476.
Durham, E., Dorr, B., Woetzel, N., Staritzbichler, R., & Meiler, J. (2009). Solvent accessible surface area approximations for rapid and accurate protein structure prediction. Journal of Molecular Modeling, 15(9), 1093–1108. https://doi.org/10.1007/s00894-009-0454-9.
Ali, S., Khan, F., Mohammad, T., Lan, D., Hassan, M., & Wang, Y. (2019). Identification and evaluation of inhibitors of lipase from Malassezia restricta using virtual high-throughput screening and molecular dynamics studies. International Journal of Molecular Sciences, 20, 884.
Khan, F., Shahbaaz, M., Bisetty, K., Waheed, A., Sly, W., Ahmad, F., & Hassan, M. (2016). Large scale analysis of the mutational landscape in beta-glucuronidase: a major player of mucopolysaccharidosis type VII. Gene, 576, 36–44.
Bhagavan, N. V., & Ha, C.-E. (2015). Nucleotide metabolism. In Essentials of medical biochemistry (pp. 465–487). Elsevier. https://doi.org/10.1016/b978-0-12-416687-5.00025-7.
Acknowledgements
Appreciation goes to Center for High Performance Computing, Cape Town, South Africa for providing computational resources.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Omolabi, K.F., Agoni, C., Olotu, F.A. et al. Molecular Basis of P131 Cryptosporidial-IMPDH Selectivity—A Structural, Dynamical and Mechanistic Stance. Cell Biochem Biophys 79, 11–24 (2021). https://doi.org/10.1007/s12013-020-00950-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-020-00950-1