Abstract
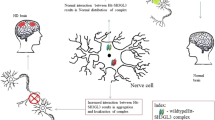
Huntington’s disease (HD) is a neurodegenerative disorder that is caused by an abnormal elongation of the polyglutamine (polyQ) chain in the Huntington (Htt) protein. At present, the normal function of Htt of neurons as well as the mechanism by which selective neurodegeneration is caused by the expanded polyQ chain in Htt remains ambiguous. A gain of function as a result of the elongated polyQ chain can lead to abnormal interaction of the Htt protein with its interacting partners, thereby resulting in the neuropathological changes seen in the Huntington’s disease. Recent research indicates protein kinase C and casein kinase substrate in neurons protein 1 (PACSIN1) as one of the interacting partners of Htt protein. It has proven experimentally that the mutant Htt and PACSIN1 formed aggregates in the cytoplasm. This aggregation is believed to be a cause for Huntington’s disease. In our study, we performed in silico investigations to predict the biomolecular mechanism of Htt/PACSIN1 interaction that could be one of the major triggers of the disease. Biomolecular interaction and molecular dynamics simulation analysis were performed to understand the dynamic behavior of native and mutant structures at the atomic level. Mutant Htt showed more interaction with its biological partner than the native Htt due to its expansion of interaction surface and flexible nature of binding residues. Our investigation of native and mutant Htt clearly shows that the structural and functional consequences of the polyQ elongation cause HD. Because of the central role of the Htt-PACSIN1 complex in maintaining connections between neurons, these differences likely contribute to the mechanism responsible for HD progression.






Similar content being viewed by others
References
HDCTG. (1993). A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington’s disease chromosomes. Cell, 72, 971–983.
Trottier, Y., et al. (1995). Polyglutamine expansion as a pathological epitope in Huntington’s disease and four dominant cerebellar ataxias. Nature, 378, 403–406.
DiFiglia, M., et al. (1995). Huntington is a cytoplasmic protein associated with vesicles in human and rat brain neurons. Neuron, 14, 1075–1081.
Chai, Y., et al. (2001). The role of protein composition in specifying nuclear inclusion formation in polyglutamine disease. Journal of Biological Chemistry, 276, 44889–44897.
Faber, P. W., et al. (1998). Huntington interacts with a family of WW domain proteins. Human Molecular Genetics, 7, 1463–1474.
Steffan, J. S., et al. (2000). The Huntington’s disease protein interactswith p53 and CREB-binding protein and represses transcription. Proceedings of the National Academy of Sciences USA, 97, 6763–6768.
Zuccato, C., et al. (2001). Loss of Huntington-mediated BDNF gene transcription in Huntington’s disease. Science, 293, 493–498.
Cattaneo, E., et al. (2001). Loss of normal huntington function: New developments in Huntington’s disease research. Trends in Neurosciences, 24, 182–188.
Boutell, J. M., et al. (1999). Aberrant interactions of transcriptional repressor proteins with the Huntington’s disease gene product, Huntington. Human Molecular Genetics, 8, 1647–1655.
Dunah, A. W., et al. (2002). Sp1 and TAFII130 transcriptional activity disrupted in early Huntington’s disease. Science, 296, 2238–2243.
Slepnev, V. I., & De Camilli, P. (2000). Accessory factors in clathrin dependent synaptic vesicle endocytosis. Nature Reviews Neuroscience, 1, 161–172.
Sun, Y., et al. (2001). Polyglutamine-expanded Huntington promotes sensitization of N-methyl-D-aspartate receptors via post-synaptic density 95. Journal of Biological Chemistry, 276, 24713–24718.
Bates, G., et al. (2002). Huntington’s disease (3rd ed.). New York: Oxford University Press.
Ontoya, A., Price, B. H., Menear, M., & Lepage, M. (2006). Brain imaging and cognitive dysfunctions in Huntington’s disease. The Journal of Psychiatry & Neuroscience, 31(1), 21–29.
Vonsattel, J. P., et al. (1985). Neuropathological classification of Huntington’s disease. Journal of Neuropathology and Experimental Neurology, 44, 559–577.
Gunawardena, S., et al. (2003). Disruption of axonal transport by loss of Huntington or expression of pathogenic polyQ proteins in Drosophila. Neuron, 40, 25–40.
Szebenyi, G., et al. (2003). Neuropathogenic forms of Huntington and androgen receptor inhibit fast axonal transport. Neuron, 40, 41–52.
Van der Burg, J. M., Björkqvist, M., & Brundin, P. (2009). Beyond the brain: Widespread pathology in Huntington’s disease. The Lancet Neurology, 8(8), 765–774. doi:10.1016/S1474-4422(09)70178-4.
Scherzinger, E., et al. (1997). Huntington-encoded polyglutamine expansions form amyloid-like protein aggregates in vitro and in vivo. Cell, 90, 549–558.
Wanker, E. E. (2000). Protein aggregation and pathogenesis of Huntington’s disease: mechanisms and correlations. The Journal of Biological Chemistry, 381, 937–942.
Poirier, M. A., Jiang, H., & Ross, C. A. (2005). A structure-based analysis of huntingtin mutant polyglutamine aggregation and toxicity: evidence for a compact beta-sheet structure. Human Molecular Genetics, 14, 765–774.
Mangiarini, L., Sathasivam, K., Seller, M., Cozens, B., Harper, A., Hetherington, C., et al. (1996). Exon 1 of the HD gene with an expanded CAG repeat is sufficient to cause a progressive neurological phenotype in transgenic mice. Cell, 87, 493–506.
Nucifora, L. G., Burke, K. A., Feng, X., Arbez, N., Zhu, S., Miller, J., et al. (2012). Identification of novel potentially toxic oligomers formed in vitro from mammalian-derived expanded huntingtin exon-1 protein. Journal of Biological Chemistry, 287, 16017–16028.
Zhang, Q. C., Yeh, T.-L., Leyva, A., Frank, L. G., Miller, J., Kim, Y. E., et al. (2011). A compact beta model of huntingtin toxicity. Journal of Biological Chemistry, 286, 8188–8196.
Zhang, Z., Norris, J., Schwartz, C., & Alexov, E. (2011). In silico and in vitro investigations of the mutability of disease-causing missense mutation sites in spermine synthase. PLoS One, 6(5), e20373.
DiFiglia, M., et al. (1997). Aggregation of Huntington in neuronal intranuclear inclusions and dystrophic neurites in brain. Science, 277, 1990–1993.
Sharp, A. H., et al. (1995). Widespread expression of Huntington’s disease gene (IT15) protein product. Neuron, 14, 1065–1074.
Sittler, A., Wälter, S., Wedemeyer, N., Hasenbank, R., Scherzinger, E., Eickhoff, H., et al. (1998). SH3GL3 associates with the huntington exon 1 protein and promotes the formation of polygln-containing protein aggregates. Molecular Cell, 2, 427–436.
Sathasivam, et al. (1997). Aberrant processing of the Fugu HD (FrHD) mRNA in mouse cells and in transgenic mice. Human Molecular Genetics, 6(12), 2141–2149.
Modregger, J., et al. (2002). PACSIN 1 interacts with Huntington and is absent from synaptic varicosities in presymptomatic Huntington’s disease brains. Human Molecular Genetics, 11, 2547–2558.
Nicniocaill, B., Haraldsson, B., Hansson, O., O’Connor, W. T., & Brundin, P. (2001). Altered striatal amino acid neurotransmitter release monitored using microdialysis in R6/1 Huntington transgenic mice. European Journal of Neuroscience, 13, 206–210.
Purohit, R., Rajendran, V., & Sethumadhavan, R. (2011). Relationship between mutation of serine residue at 315th position in M. tuberculosis catalase-peroxidase enzyme and isoniazid susceptibility: An in silico analysis. Journal of Molecular Modeling, 17(4), 869–877.
Kumar, A., & Purohit, R. (2012). Computational investigation of pathogenic nsSNPs in CEP63 protein. Gene, 503(1), 75–82.
Rajendran, V., Purohit, R., & Sethumadhavan, R. (2012). In silico investigation of molecular mechanism of laminopathy caused by a point mutation (R482W) in lamin A/C protein. Amino Acids, 43(2), 603–615.
Rajendran, V., & Sethumadhavan, R. (2014). Drug resistance mechanism of PncA in Mycobacterium tuberculosis. Journal of Biomolecular Structure & Dynamics, 32(2), 209–221.
Purohit, R. (2014). Role of ELA region in auto-activation of mutant KIT receptor: a molecular dynamics simulation insight. Journal of Biomolecular Structure & Dynamics, 32(7), 1033–1046.
Gopalakrishnan, C., Kalsi, N., Jethi, S., & Purohit, R. (2015). Computational investigation of molecular mechanism and neuropathological implications in Huntington disease. Molecular and Cellular Biochemistry, 409(1–2), 1–11.
Kalsi, N., Gopalakrishnan, C., Rajendran, V., & Purohit, R. (2016). Biophysical aspect of phosphatidylinositol 3-kinase and role of oncogenic mutants (E542K & E545K). Journal of Biomolecular Structure & Dynamics,. doi:10.1080/07391102.2015.1127774.
Zhang, Y. (2008). I-TASSER server for protein 3D structure prediction. BMC Bioinformatics, 9, 40.
Dominguez, C., Boelens, R., & Bonvin, A. M. (2003). HADDOCK: A protein–protein docking approach based on biochemical or biophysical information. Journal of the American Chemical Society, 125, 1731–1737.
Dominguez, C., Boelens, R., & Bonvin, A. M. (2003). The HADDOCK web server for data-driven biomolecular docking. Nature Protocols, 5, 883–897.
Nilges, M. (1995). Calculation of protein structures with ambiguous distance restraints. Automated assignment of ambiguous NOE crosspeaks and disulphide connectivities. Journal of Molecular Biology, 245, 645–660.
Nilges, M., Macias, M. J., O’Donoghue, S. I., & Oschkinat, H. (1997). Automated NOESY interpretation with ambiguous distance restraints: the refined NMR solution structure of the pleckstrin homology domain from beta-spectrin. Journal of Molecular Biology, 269, 408–422.
Brunger, A. T., Adams, P. D., Clore, G. M., DeLano, W. L., Gros, P., Grosse- Kunstleve, R. W., et al. (1998). Crystallography and NMR system: A new software suite for macromolecular structure determination. Acta Crystallographica. Section D, Biological Crystallography, 54, 905–921.
Porollo, A., & Meller, J. (2007). Prediction-based fingerprints of protein-protein interactions. Proteins: Structure, Function, and Bioinformatics, 66, 630–645.
Porollo, A., & Meller, J. (2012). Computational methods for prediction of protein-protein interaction sites. In W. Cai & H. Hong (Eds.), Protein-protein interactions—computational and experimental tools, InTech (Vol. 472, pp. 3–26).
Hess, B., Kutzner, C., Spoel, D., & Lindahl, E. (2008). GROMACS 4: Algorithms for highly efficient, load-balanced, and scalable molecular simulation. Journal of Chemical Theory and Computation, 4, 435–447.
Spoel D., et al. (2005). GROMACS: Fast, flexible, and free. Journal of Computational Chemistry, 26, 1701–1718.
Berendsen, H. J. C., Postma, J. P. M., Gunsteren, W. F. V., & Hermans, J. (1981). Interaction models for water in relation to protein hydration. In B. Pullman (Ed.), Intermolecular forces (pp. 331–342). Dordrecht: D. Reidel Publishing Company.
Berendsen, H. J. C., Postma, J. P. M., DiNola, A., & Hakk, J. R. (1984). Molecular dynamics with coupling to an external bath. The Journal of Chemical Physics, 81, 3684–3690.
Essmann, U., Perera, L., Berkowitz, M. L., Darden, T., Lee, H., & Pedersen, L. G. (1995). A smooth particle mesh Ewald method. The Journal of Chemical Physics, 103, 8577–8593.
Halperin, I., Ma, B., Wolfson, H., & Nussinov, R. (2002). Principles of docking: An overview of search algorithms and a guide to scoring functions. Proteins: Structure, Function, and Genetics, 47, 409–443.
Janin, J., Henrick, K., Moult, J., Eyck, L. T., Sternberg, M. J., Vajda, S., et al. (2003). CAPRI: A critical assessment of predicted interactions. Proteins: Structure, Function, and Genetics, 52, 2–9.
van Dijk, M., van Dijk, A. D. J., Hsu, V., Boelens, R., & Bonvin, A. M. J. J. (2006). Information-driven protein–DNA docking using HADDOCK: It is a matter of flexibility. Nucleic Acids Research., 34(11), 3317–3325. doi:10.1093/nar/gkl412.
Kaltenbach, L. S., et al. (2007). Huntingtin interacting proteins are genetic modifiers of Neurodegeneration. PLoS Genetics, 3(5), e82.
Acknowledgments
We gratefully acknowledge the CSIR-Institute of Himalayan Bioresource Technology, Palampur, Vellore Institute of Technology University and Bioinformatics Resources and Applications Facility (BRAF), C-DAC, Pune for providing the facilities to carry out this work. This manuscript represents CSIR-IHBT communication number 4022.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Additional information
Chandrasekhar Gopalakrishnan and Shraddha Jethi have contributed equally to this manuscript.
Rights and permissions
About this article
Cite this article
Gopalakrishnan, C., Jethi, S., Kalsi, N. et al. Biophysical Aspect of Huntingtin Protein During polyQ: An In Silico Insight. Cell Biochem Biophys 74, 129–139 (2016). https://doi.org/10.1007/s12013-016-0728-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-016-0728-7