Abstract
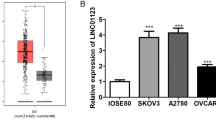
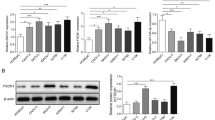
Previous reports indicate that long intergenic non-coding RNA LINC00665 naturally occurred vital effects in various cancers. Herein, the role of LINC00665 in ovarian cancer progress was explored. We found that LINC00665 was upregulated in ovarian cancer cell lines. Besides, a series of assays including flow cytometry, wound-healing, transwell, cell counting Kit-8 (CCK-8), and EdU assay confirmed that the knockdown of LINC00665 could reduce the viability, proliferation, and migration of SKOV-3 and OVCAR-3 cells. Accumulating evidence indicates that many lncRNAs can function as endogenous miRNA sponges by competitively binding common miRNAs. In this study, the bioinformatics analysis suggests that LNC00665 specifically binds to miR-181a-5p. LINC00665 downregulated the miR-181a-5p in SKOV-3 and OVCAR-3 cells. The knockdown of miR-181a-5p evidently reverses the inhibitory effect of sh-LINC00662. Besides, FH2 domain containing 1 (FHDC1) has been proved to deed as an effective target of miR-181a-5p. The results reveal the knockdown of LINC00665 facilitates ovarian cancer via development by sponging miR-181a-5p and up-regulating FHDC1 expression. These may contribute to ovarian cancer therapy.





Similar content being viewed by others
Data Availability
All data generated or analyzed during this study are included in this published article.
References
Bray, F., Ferlay, J., Soerjomataram, I., Siegel, R. L., Torre, L. A., & Jemal, A. (2018). Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians, 68, 394–424.
Torre, L. A., Trabert, B., DeSantis, C. E., Miller, K. D., Samimi, G., Runowicz, C. D., Gaudet, M. M., Jemal, A., & Siegel, R. L. (2018). Ovarian cancer statistics, 2018. CA: A Cancer Journal for Clinicians, 68, 284–296.
Zhang, M. L., Peng, P., Wu, C. X., Gong, Y. M., Zhang, S. W., Chen, W. Q., & Bao, P. P. (2019). Report of breast cancer incidence and mortality in China registry regions, 2008–2012. Zhonghua Zhong Liu Za Zhi, 41, 315–320.
Marth, C., Reimer, D., & Zeimet, A. G. (2017). Front-line therapy of advanced epithelial ovarian cancer: standard treatment. Ann Oncol, 28(Suppl 8), viii36–viii39.
Qiu, M. T., Hu, J. W., Yin, R., & Xu, L. (2013). Long noncoding RNA: An emerging paradigm of cancer research. Tumour Biology, 34, 613–620.
Lalevee, S., & Feil, R. (2015). Long noncoding RNAs in human disease: Emerging mechanisms and therapeutic strategies. Epigenomics, 7, 877–879.
DiStefano, J. K. (2018). The Emerging Role of Long Noncoding RNAs in Human Disease. Methods in Molecular Biology, 1706, 91–110.
Xue, M., Zhuo, Y., & Shan, B. (2017). MicroRNAs, Long Noncoding RNAs, and their functions in human disease. Methods in Molecular Biology, 1617, 1–25.
Lv, P., Qiu, X., Gu, Y., Yang, X., Xu, X., & Yang, Y. (2019). Long non-coding RNA SNHG6 enhances cell proliferation, migration and invasion by regulating miR-26a-5p/MAPK6 in breast cancer. Biomedicine & Pharmacotherapy, 110, 294–301.
Sun, X., Huang, T., Zhang, C., Zhang, S., Wang, Y., Zhang, Q., & Liu, Z. (2019). Long non-coding RNA LINC00968 reduces cell proliferation and migration and angiogenesis in breast cancer through up-regulation of PROX1 by reducing hsa-miR-423-5p. Cell Cycle, 18, 1908–1924.
Jia, H., Wang, X., & Sun, Z. (2018). Exploring the molecular pathogenesis and biomarkers of high risk oral premalignant lesions on the basis of long noncoding RNA expression profiling by serial analysis of gene expression. European Journal of Cancer Prevention, 27, 370–378.
Wen, D. Y., Lin, P., Pang, Y. Y., Chen, G., He, Y., Dang, Y. W., & Yang, H. (2018). Expression of the long intergenic non-protein coding RNA 665 (LINC00665) gene and the cell cycle in hepatocellular carcinoma using the cancer genome atlas, the gene expression omnibus, and quantitative real-time polymerase chain reaction. Medical Science Monitor, 24, 2786–2808.
Cong, Z., Diao, Y., Xu, Y., Li, X., Jiang, Z., Shao, C., Ji, S., Shen, Y., De, W., & Qiang, Y. (2019). Long non-coding RNA linc00665 promotes lung adenocarcinoma progression and functions as ceRNA to regulate AKR1B10-ERK signaling by sponging miR-98. Cell Death & Disease, 10, 84.
Adami, G. R., Tangney, C. C., Tang, J. L., Zhou, Y., Ghaffari, S., Naqib, A., Sinha, S., Green, S. J., & Schwartz, J. L. (2018). Effects of green tea on miRNA and microbiome of oral epithelium. Science and Reports, 8, 5873.
Rodriguez, L. G., Wu, X., & Guan, J. L. (2005). Wound-healing assay. Methods in Molecular Biology, 294, 23–29.
Marshall, J. (2011). Transwell((R)) invasion assays. Methods in Molecular Biology, 769, 97–110.
Li, Q., Zhang, J., Zhou, J., Yang, B., Liu, P., Cao, L., Jing, L., & Liu, H. (2018). lncRNAs are novel biomarkers for differentiating between cisplatin-resistant and cisplatin-sensitive ovarian cancer. Oncology Letters, 15, 8363–8370.
Shen, W., Xie, X., Liu, M., & Wang, L. (2020). Diagnostic Value of lncRNA ROR in Differentiating Ovarian Cancer Patients. Clinical laboratory, 66.
Shi, Y., Gao, S., Zheng, Y., Yao, M., & Ruan, F. (2019). LncRNA CASC15 Functions As An Unfavorable Predictor Of Ovarian Cancer Prognosis And Inhibits Tumor Progression Through Regulation Of miR-221/ARID1A Axis. Oncotargets and Therapy, 12, 8725–8736.
Li, Y., Kuscu, C., Banach, A., Zhang, Q., Pulkoski-Gross, A., Kim, D., Liu, J., Roth, E., Li, E., Shroyer, K. R., Denoya, P. I., Zhu, X., Chen, L., & Cao, J. (2015). miR-181a-5p inhibits cancer cell migration and angiogenesis via downregulation of matrix metalloproteinase-14. Cancer Research, 75, 2674–2685.
Qi, H., Xiao, Z., & Wang, Y. (2019). Long non-coding RNA LINC00665 gastric cancer tumorigenesis by regulation miR-149-3p/RNF2 axis. Oncotargets and Therapy, 12, 6981–6990.
Chen, W., Yu, Z., Huang, W., Yang, Y., Wang, F., & Huang, H. (2020). LncRNA LINC00665 promotes prostate cancer progression via miR-1224-5p/SND1 Axis. Oncotargets and Therapy, 13, 2527–2535.
Petrillo, M., Zannoni, G. F., Beltrame, L., Martinelli, E., DiFeo, A., Paracchini, L., Craparotta, I., Mannarino, L., Vizzielli, G., Scambia, G., D’Incalci, M., Romualdi, C., & Marchini, S. (2016). Identification of high-grade serous ovarian cancer miRNA species associated with survival and drug response in patients receiving neoadjuvant chemotherapy: A retrospective longitudinal analysis using matched tumor biopsies. Annals of Oncology, 27, 625–634.
Copeland, S. J., Thurston, S. F., & Copeland, J. W. (2016). Actin- and microtubule-dependent regulation of Golgi morphology by FHDC1. Molecular Biology of the Cell, 27, 260–276.
Chen, H. Q., Zhao, J., Li, Y., He, L. X., Huang, Y. J., Shu, W. Q., Cao, J., Liu, W. B., & Liu, J. Y. (2018). Gene expression network regulated by DNA methylation and microRNA during microcystin-leucine arginine induced malignant transformation in human hepatocyte L02 cells. Toxicology Letters, 289, 42–53.
Author information
Authors and Affiliations
Contributions
S.W conducted most of the experiments. J.W. interpreted and analyzed the data. Y.W wrote the first draft of the article. J.L. finalized the manuscript. J.W. conceived the study and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics Approval and Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, S., Wang, Y., Lu, J. et al. LncRNA LINC00665 Promotes Ovarian Cancer Cell Proliferation and Inhibits Apoptosis via Targeting miR-181a-5p/FHDC. Appl Biochem Biotechnol 194, 3819–3832 (2022). https://doi.org/10.1007/s12010-022-03943-3
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-022-03943-3