Abstract
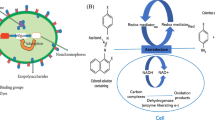
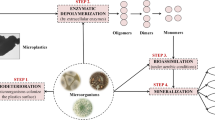
Biofouling leads to unfavorable problems in underwater structures like ships’ hulls, aquaculture cages, fishnets, petroleum pipelines, sensors, and other equipment. The main target of antifouling applications is to prevent the early steps of fouling, namely microfouling (bacterial biofilm). In recent years, investigations into the development of environmentally friendly, nontoxic products in antifouling coating technology are becoming more widespread. The main purpose of this study was to investigate the antibiofilm activity of commercial hydrolytic enzymes (α-amylase, xylanase, glucanase, protease, pectinase, lipase, viscozyme, and lysing complex) against marine biofilm bacteria (Pseudoalteromonas agarivorans FJ040188, Vibrio lentus FJ200649, and Exiguobacterium homiense FJ200653). The experimental method involved a prevention and detachment test based on bacterial adhesion in a microplate and biomass quantification of the total biofilm using DAPI fluorescent dye after a 24-hour incubation period. Results from the screening of the biofilm percentage reduction in the enzymes, a lysing complex enzyme, were seen to be the most effective in both the prevention of bacterial adhesion and the detachment of adhered bacteria. Epifluorescence microscopy analysis of DAPI-stained adhered bacteria using lysing complex confirmed these results. It was seen that the mixtures of enzymes selected for present study have the potential to be employed as environmentally friendly antifouling additives in marine antifouling coatings.





Similar content being viewed by others
References
Kristensen, JB, Meyer, RL, Laursen, BS, Shipovskov, S, Besenbacher, F, Poulsen, CH, “Antifouling enzymes and the biochemistry of marine settlement.” Biotechnol. Adv., 26 471–481 (2008)
Holmström, C, Rittschof, D, Kjelleberg, S, “Inhibition of settlement of larvae of Balanus amphitrite and Ciona intestinalis by a surface-colonising marine bacterium.” Appl. Environ. Microbiol., 58 2111–2115 (1992)
Wieczorek, SK, Todd, CD, “Biofilms cues and larval settlement.” Biofouling., 12 81–118 (1998)
Qian, PY, Thiyagarajan, V, Lau, SCK, Cheung, SCK, “Relationship between community profile in biofilm and attachment of the acorn barnacle Balanus Amphitrite.” Aquat. Microb. Ecol., 33 225–237 (2003)
Leroy, C, Delbarre, C, Ghillebaert, F, Compere, C, Combes, D, “Effects of commercial enzymes on the adhesion of a marine biofilm-forming bacterium.” Biofouling, 24 (1) 11–22 (2008)
Sutherland, IW, “The biofilm matrix—an immobilized but dynamic microbial environment.” Trends Microbiol., 9 222–227 (2001)
Allison, DG, “Molecular architecture of the biofilm matrix.” In: Lens, P (ed.) Biofilms in Medicine, Industry, and Environment Technology, pp. 81–90. IWA Publishing, London (2003)
Branda, SS, Vik, A, Friedman, L, Kolter, R, “Biofilms: the matrix revisited.” Trends Microbiol., 13 20–26 (2005)
Lejars, M, Margaillan, A, Bressy, C, “Fouling Release Coatings: A Nontoxic Alternative to Biocidal Antifouling Coatings.” Chem. Rev., 112 4347–4390 (2012)
Zanaroli, G, Negroni, A, Calisti, C, Ruzzi, M, Fava, F, “Selection of commercial hydrolytic enzymes with potential antifouling activity in marine environments.” Enzym. Microb. Technol., 49 574–579 (2011)
Kristensen, JB, Meyer, RL, Poulsen, CH, Kragh, KM, Besenbacher, F, Laursen, BS, “Biomimetic silica encapsulation of enzymes for replacement of biocides in antifouling coatings.” Green Chem., 12 387–394 (2010)
Flemming, HC, Wingender, J, “The biofilm matrix.” Nat. Rev. Microbiol., 8 623–633 (2010)
Rawlings, N, Morton, F, Barrett, A, “MEROPS: the peptidase database.” Nucleic Acids Res., 34 270–272 (2006)
Callow, JA, Callow, ME, The Ulva spore adhesive system. In Biological adhesives, Springer, Berlin Heidelberg, Germany (2006)
Olsen, SM, Pedersen, LT, Laursen, MH, Kiil, S, Dam-Johansen, K, “Enzyme-based antifouling coatings: a review.” Biofouling., 23 (5) 369–383 (2007)
Aehle, W, Enzymes in industry: products and applications, 2nd ed. Wiley, Germany (2003)
Kacar, A, Kocyigit, A, Ozdemir, G, Cihangir, B, “The development of biofilm bacteria on panels coated by different antifouling paints in the marinas.” Fresen. Environ. Bull., 18 (11) 2004–2012 (2009)
Somogyi, M, “Modifications of two methods for the assay of amylase.” Clin. Chem., 6 23–35 (1960)
Tomarelli, RM, Charney, J, Harding, ML, “The use of azoalbumin as a substrate in the colorimetric determination of peptic and tryptic activity.” J. Lab. Clin. Med., 34 428–433 (1949)
Kilcawley, KN, Wilkinson, MG, Fox, PF, “Determination of key enzyme activities in commercial peptidase and lipase preparations from microbial or animal sources.” Enzyme Microb. Technol., 31 310–320 (2002)
Leroy, C, “Lutte contre les salissures marines: approche par procédés enzymatiques.” Ph.D. thesis, University of Toulouse, France (2006)
Larsen, P, Nielsen, JL, Dueholm, MS, Wetzel, R, Otzen, D, Nielsen, PH, “Amyloid adhesions are abundant in natural biofilms.” Environ. Microbiol., 9 (12) 3077–3090 (2007)
Suzuki, Y, Tsujimoto, Y, Matsui, H, Watanabe, K, “Decomposition of extremely hard-to-degrade animal proteins by thermophilic bacteria.” J. Biosci. Bioeng., 102 (2) 73–81 (2006)
Pettitt, M, Henry, S, Callow, J, Clare, A, “Activity of commercial enzymes on settlement and adhesion of cypris larvae of the barnacle Balanus amphitrite, spores of the green alga Ulva linza, and the diatom Navicula perminuta.” Biofouling, 20 299–311 (2004)
Allermann, K, Schneider, I, Walstrøm, E, Andersen, BH, Andersen, TT, Højenvang, J, “Substitution af biocider i bundmaling til skibe med enzyme.” Danish Environmental Protection Agency., (2004)
Callow, JA, Stanley, MS, Wetherbee, R, Callow, ME, “Cellular and molecular approaches to understanding primary adhesion in Enteromorpha: an overview.” Biofouling, 16 (2–4) 141–150 (2000)
Aldridge, IY, Chmurny, AB, Durham, DR, Roberts, RL, “Proteases to inhibit and remove biofilm.” EP Patent 0,590,746B, 1998
Hollis, GC, Terry, JP, Jaquess, PA, “Composition and methods for removing or preventing biofilm.” WO Patent 92/13807, 1992
Johansen, C, Falholt, P, Gram, L, “Enzymatic removal and disinfection of bacterial biofilms.” Appl. Environ. Microbiol., 63 (9) 3724–3728 (1997)
Hamade, KR, Yamammori, TN, Yoshio, KO, “Glucoxide derivatives for enzyme modification, lipid-coated enzymes, method of producing such enzymes and antifouling paint composition.” US Patent 5,770,188, 1996
Huijs, FM, Klijnstra, JW, van Zanten, J, “Antifouling coating comprising a polymer with functional groups bonded to an enzyme.” European patent application EP1,661,955A1, 2004
Kato, N, “Enzyme-containing antifouling emulsion coating compositions.” Japanese patent JP63202677, 1987
Iwamura, G, Kuwamura, S, Shoji, A, Kito, K, “Antifouling coating materials containing immobilized enzymes.” Japanese patent JP02227471, 1989
Kuwamura, S, Iwamura, G, Shoji, A, Kito, K, “Antifouling coatings containing enzymes.” Japanese patent JP02227465, 1989
Bonaventura, C, Bonaventura, J, Hooper, IR, “Anti-fouling methods using enzyme coatings.” US patent number 5998200, 1991
Allermann, K, Schneider, I, “Antifouling paint composition comprising rosin and enzyme.” US patent Application Publication US2002/0166237A1, 2000
Bonaventura, C, Bonaventura, J, Hooper, IR, “Anti-fouling methods using enzyme coatings.” US patent number US 5,998,200, 1999
Huijs, FM, Klijnstra, JW, “Antifouling coating comprising a polymer with functional groups bonded to an enzyme.” European patent number EP 1,661,955, 2006
Acknowledgments
This study was supported by Scientific Research Projects Coordination Unit of Dokuz Eylul University (Project Number: 2015.KB.FEN.015).
Conflict of interest
The authors have declared no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Aykin, E., Omuzbuken, B. & Kacar, A. Microfouling bacteria and the use of enzymes in eco-friendly antifouling technology. J Coat Technol Res 16, 847–856 (2019). https://doi.org/10.1007/s11998-018-00161-7
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11998-018-00161-7