Abstract
Purpose of Review
Cardiac cell-based therapy represents a promising approach for cardiac repair. However, one of the main challenges is cardiac arrhythmias associated with stem cell transplantation. The current review summarizes the recent progress in model systems for addressing mechanisms of arrhythmogenesis in cardiac repair.
Recent Findings
Animal models have been extensively developed for mechanistic studies of cardiac arrhythmogenesis. Advances in human induced pluripotent stem cells (hiPSCs), patient-specific disease models, tissue engineering, and gene editing have greatly enhanced our ability to probe the mechanistic bases of cardiac arrhythmias. Additionally, recent development in multiscale computational studies and machine learning provides yet another powerful tool to quantitatively decipher the mechanisms of cardiac arrhythmias.
Summary
Advancing efforts towards the integrations of experimental and computational studies are critical to gain insights into novel mitigation strategies for cardiac arrhythmias in cell-based therapy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cardiovascular disease is the leading cause of morbidity and mortality worldwide and causes more deaths than all cancers combined [1]. Despite significant advances in therapy and management, heart failure (HF) remains a life-threatening disease with a 5-year mortality rate of 45–60% [1]. Therefore, there is a compelling need to seek new options for patients suffering from HF. Since adult cardiac myocytes are unable to proliferate sufficiently to replace the damaged tissue, stem cell therapy represents a promising approach for the treatment of end-stage HF, since it aims at generating new functional myocardium and inducing neoangiogenesis. However, therapeutic strategies using cell-based therapy have not produced full restorative functions [2, 3]. A high rate of transplanted stem cell loss (90% within the first few days) has been observed [4, 5]. Moreover, stem cell transplantation has been shown to be associated with occurences of cardiac arrhythmias [6•, 7••, 8, 9], which represents one of the main challenges in the field of cardiac cell-based therapy.
Animal models have been extensively developed for mechanistic studies of cardiac arrhythmogenesis [10•, 11,12,13], enabling genetic modification using gain- and loss-of-function strategies. However, limitations exist including significant electrophysiological differences between human and animal hearts, costs, as well as ethical considerations. To circumvent some of these shortcomings, cellular and multicellular models for arrhythmogenesis have been widely used, which are further enhanced by advances in human-induced pluripotent stem cells (hiPSCs), tissue engineering [14,15,16,17,18,19], and gene editing. The hiPSC-derived cardiomyocytes (hiPSC-CMs) can be obtained from healthy or diseased individuals, and provide the inexhaustible source for human disease modeling. Finally, recent development in multiscale computational studies provides yet another powerful tool to quantitatively test the mechanisms of cardiac arrhythmias in cardiac repair [20••].
The current review will summarize the recent progress in model systems for addressing mechanisms of cardiac arrhythmogenesis, and serve as a discussion platform to gain insights into mechanistic underpinnings and novel mitigation strategies for arrhythmogenesis in cardiac repair.
Animal Models for Cardiac Arrhythmias
Numerous species have been utilized to study the underlying mechanisms of arrhythmogenesis, ranging from small animals such as zebrafish to large animal models such as pigs and nonhuman primates (Fig. 1). Although no animal models perfectly replicate arrhythmogenesis seen in humans, the multitudes of animal models have substantially advanced our understanding of different aspects of arrhythmogenesis. Small animal models are generally used for mechanistic discoveries, drug screening, and testing, while larger animals are reserved for validation of these key findings and to further establish drug safety profiles.
Cardiac Hypertrophy
Electrical and structural remodeling in pathological cardiac hypertrophy has been shown to increase patient’s susceptibility to ventricular arrhythmias (VAs) and sudden cardiac death (SCD). Electrical remodeling in cardiac hypertrophy is well documented and includes action potential (AP) prolongation [21] and conduction delay [22]. One of the most commonly used animal models of cardiac hypertrophy is surgical aortic constriction, which elevates afterload and consequently induces adverse structural and electrical remodeling. Aortic constriction is commonly used in mouse models due to the ease of genetic manipulation; however, it has been adapted to rats [21], guinea pigs [23], rabbits [24], and pigs [25]. Additionally, animal models with volume overload can induce pathological remodeling of the heart that result in cardiac hypertrophy, such as arterio-venous shunt formation and aortic regurgitation [26].
Myocardial Ischemia and Infarction
Coronary artery disease (CAD), leading to myocardial ischemia and infarction, is a frequent cause of cardiac arrhythmia. Cardiac ischemia and infarction result in complex electrophysiological remodeling [27•]. In murine models, myocardial infarction is typically induced by ligation of the left anterior descending (LAD) coronary artery. In larger animal models, ligation of other coronary arteries such as the left circumflex or right coronary artery has also been used [28].
Atrial and Ventricular Tachyarrhythmia
Atrial fibrillation (AF) is the most prevalent cardiac arrhythmia seen clinically [29]. A commonly used model of AF is atrial tachypacing. Although it is employed in dogs [30], sheep [31], pigs [32], and rabbits [33], its usage has not been extended over to smaller animals due to their significantly higher heart rates [34]. Tachypacing may also be used on the ventricles to study HF-induced arrhythmogenesis [35].
Heart Failure
Heart failure (HF) occurs when the cardiac output is no longer able to meet the metabolic demands of the body, and is frequently associated with cardiac arrhythmias [36]. Animal models of cardiac hypertrophy, myocardial ischemia/infarction, and atrial and ventricular tachypacing [35] can produce HF when the intervention is prolonged, combined, and/or severe. Although not necessary in smaller animals, larger animals may require more than one intervention to induce HF. For instance, a commonly used rabbit model of HF involves aortic valve cusp perforation, followed by aortic constriction, to create a volume and subsequent pressure overload [26]. These multiple interventions are required since HF is uncommon with pressure overload in rabbits, but they may induce an increase in mortality.
Genetic Models
Gene-targeted animal models predominantly in mice have greatly expanded our mechanistic understanding of long QT syndrome (LQTS), short QT syndrome (SQTS), Brugada syndrome, catecholaminergic polymorphic ventricular tachyarrhythmia (CPVT), sick sinus syndrome, cardiac conduction disease, and familial AF [37]. Genetic manipulation in larger animals have also been described including LQTS 1 and 2 models in rabbits, overexpression of TGFB1 in goats, and mutated SCN5A in pigs [10•]. Indeed, emerging technologies for genome editing, including clustered regularly interspaced short palindromic repeats (CRISPR)-Cas9 mediated gene editing, have enabled the generation of genetic models for arrhythmias in larger animals.
Animal models have significantly expanded our understanding of the mechanistic underpinning of arrhythmogenesis, but there exist advantages and limitations that should be considered. Besides the obvious issues with costs and the length of time required for model generation, there are inherent species-dependent differences at the molecular and cellular levels that impact pathophysiology. Indeed, these differences must be evaluated and understood in order to translate therapeutic findings to clinical practice. Advantages of using small animal models are their relatively inexpensive costs, short gestational period and large litter size, and relative ease of genetic manipulation. However, small animals possess cardiovascular anatomy and physiology that are substantially different from humans [34]. For instance, AP profiles, electrocardiograms, and heart rate are significantly different between humans and rodents [10•]. These differences make larger animal models more suitable for cardiac electrophysiology and arrhythmia studies. Nonetheless, the usage of small animal models in arrhythmia research provides valuable insights, guides the design of studies in larger animals, and eventual translation to humans.
Ex Vivo Models for Cardiac Arrhythmias
Studies in cardiac arrhythmogenesis have also taken advantage of the ex vivo Langendorff-perfused heart (Fig. 1). Isolated hearts from rabbits, guinea pigs, sheep, and mice have been extensively used to investigate cardiac repolarization [38, 39], AF [40, 41], ventricular fibrillation (VF) [42,43,44], torsades de pointes (TdP) [45], AV node conduction [46, 47], and hypokalemia-induced arrhythmia [38, 48, 49]. Explanted human hearts from patients who undergo heart transplantation have been invaluable in cardiac arrhythmia studies [44]. Additionally, ex vivo Langendorff-perfused hearts are employed for drug testing [46, 50,51,52,53,54]. The ex vivo studies enable regional-specific interrogations including cardiac conduction systems, sinoatrial and atrioventricular nodes, regional heterogeneity, and cell-cell interactions. Finally, advanced imaging techniques such as optical or electrical mapping, two-photon microscopy, and optogenetics have greatly enhanced the impact of these ex vivo models [55,56,57,58,59,60]. One main pitfall of the model is the absence of cardiac innervation, which is a critical factor for arrhythmogenesis.
Cellular and Tissue Models for Arrhythmogenesis
For deciphering the mechanisms of arrhythmogenesis, in vitro models are useful tools that can simplify experiments by restricting the factors of interest to be tested independently. The models vary from murine to human cardiomyocytes and from single cells to multi-cellular preparations (Fig. 1). Each of these models is associated with distinct advantages and disadvantages.
Cardiomyocytes From Animal Models
Primary cardiomyocytes can be harvested from animal models and are critical for our understanding of mechanisms of arrhythmias at the cellular and subcellular levels. The source of focal arrhythmic activity—spontaneously generated or triggered AP—either due to ion channelopathies, altered ion channel expression, subcellular localization, or post-translational modifications, and drug-modulated channel function, can be readily deciphered using patch-clamp and imaging techniques. Myocardial infarction models exhibiting arrhythmias in vivo that have been attributed to reduced Ca2+ or K+ currents were determined in single cardiomyocytes [61, 62]. Underlying cellular mechanisms of arrhythmias in atrial and ventricular tachycardia models were similarly determined in isolated single cells [63]. Studies using single cardiomyocytes enable focused experiments on the cellular and subcellular mechanisms in the absence of other contributing pro-arrhythmic factors, such as fibrosis.
Cell Lines of Cardiomyocytes
A readily available source of cardiomyocytes is highly desirable for in vitro studies; however, the lack of regenerative capability of adult cardiomyocytes requires that primary culture is used as the main source. To circumvent limited proliferative potential of cardiomyocytes, cell lines have been generated by restoring the proliferative ability. This can be achieved by infecting cardiomyocytes with simian virus (SV) 40, a type of DNA tumor virus, that expresses large and small T antigens, which cooperatively inhibit tumor suppressors and transform the cells to escape senescence [64]. HL-1 cells are commercially available, immortalized murine atrial cardiomyocytes that originated from a subcutaneously engrafted atrial tumor, AT-1 expressing SV40 large T-antigen under the atrial natriuretic factor promoter, in C57BL/6J mice [65]. The ability of this cell line to expand indefinitely in vitro and to allow cryo-storage, while maintaining the characteristics and function of cardiomyocytes, make this cell line an attractive alternative to primary cardiomyocytes. Since its derivation, this line has been used in a large number of studies, including those investigating reentry spiral waves [66]. A human cardiomyocyte line, AC16, has been similarly generated through fusion of primary ventricular cardiomyocytes with a SV40-immortalized human fibroblasts [67]. Although this cell line expresses cardiomyocyte-specific markers and exhibits outward currents, it is unable to generate APs due to its deficiency in inward currents. This caveat in electrophysiological characteristic makes it unsuitable for arrhythmia studies.
HiPSC-Derived Cardiomyocytes
Considerable insights in cardiac arrhythmias have been derived from various species. However, the mechanisms responsible for arrhythmogenesis in the context of human electrophysiology may be significantly different from animal models with differing electrophysiology. Therefore, human cardiomyocytes remain the most reliable model for investigating the mechanisms of human arrhythmias. Primary human cardiomyocytes are not easily obtainable. Their laboratory usage is further limited by the difficulty of maintenance in culture. HiPSCs that can differentiate into all somatic cell types, including cardiomyocytes, have garnered great interest in recent years as a readily available option. These cells not only open the possibility of having an unlimited cell source as well as genetic modification or imposed environmental conditions but hiPSCs can also be generated from patients suffering from cardiac arrhythmias to generate patient-specific in vitro models. To date, numerous LQTS-, SQTS-, Brugada syndrome-, and CPVT-hiPSC lines have been generated to study the channelopathies [68,69,70,71,72,73]. Cardiomyocytes differentiated from these diseased lines faithfully exhibit abnormal APs and arrhythmias as expected. The predictive value of hiPSC-derived cardiomyocytes for drug testing and safety screening has also been demonstrated [74,75,76]. Additionally, this model has been utilized to iteratively improve the moiety of a target compound, mexiletine, for therapeutic potency and safety [77].
HiPSC-derived cardiomyocytes are not, however, without disadvantages. There are two key issues associated with using these cells for studying arrhythmias: (1) the heterogeneity of cardiomyocyte subtypes in the differentiated population and (2) immaturities of hiPSC-derived cardiomyocytes. Widely used cardiomyogenesis protocols to differentiate hiPSCs yield a heterogeneous population of pacemaker-, atrial-, and ventricular-like subtypes, as classified by the AP profiles [78]. The heterogeneity in electrophysiology among the subtypes can mask dysfunctional effects due to differential expression of ionic currents. Indeed, only the atrial and ventricular subtypes differentiated from LQT1-patient hiPSCs exhibit significantly prolonged AP duration, while the electrophysiology of the pacemaking subtype appears normal [79]. The results highlight the importance of using the appropriate subtype for studying the arrhythmia in question.
A few strategies have been devised to minimize subtype impurities. Enrichment of desired cardiomyocyte subtype requires a surface marker with high specificity for the subtype. Ventricular cardiomyocyte enrichment by selecting CORIN-positive or CD77-positive/CD200-negative cardiomyocytes has been reported, but the specificity could still be improved [80, 81]. An alternative strategy to reduce undesired cardiomyocyte subtype is direct differentiation to a specific subtype. Directed differentiation protocols that promote a higher fraction of ventricular, atrial, or pacemaking subtype have been reported [82,83,84,85,86,87,88]; however, none of the protocols could attain 100% yield of the desired subtype.
Immaturity of hiPSC-derived cardiomyocytes compared to adult cells, from excitation to contraction, is another issue that may confound experimental results. For instance, the pacemaking, hyperpolarization-activated cyclic nucleotide-gated (HCN4) channels that are absent in the adult working cardiomyocytes are present in all hiPSC-derived cardiomyocytes [89]. Conversely, inwardly rectifying Kir2.1 channels that are present to stabilize the membrane potential in adult working cardiomyocytes are largely absent in hiPSC-derived cardiomyocytes [90]. Consequently, all hiPSC-derived cardiomyocytes exhibit automaticity or phase-4 depolarization, both of which can be a trigger for arrhythmia. Therefore, only a preparation with stable baseline response should be used for the experiments. Whenever possible, the baseline response should also be recorded for comparison against induced dysfunction. The lack of Kir2.1 also means that cardiomyocytes differentiated from a LQT7 patient-specific hiPSC line are unlikely to be good candidates to study this particular channelopathy until these cells further mature.
To remedy this developmental deficiency, maturation strategies have been devised, which range from electrical, mechanical, to metabolic conditioning [90,91,92,93,94,95]. Significant improvements in all functional aspects have been observed with maturation conditioning, but none can attain the characteristics matching the adult cells. Despite immaturities, hiPSC-CMs have been used to study numerous channelopathies and for drug screening with expected presentation of dysfunction and accuracy [68,69,70,71,72,73,74,75,76]. Of note, immature hiPSC-derived cardiomyocytes have been reported to exhibit greater sensitivity to pro-arrhythmogenic drugs than those that have undergone the maturation process. Increased rate of false-positive arrhythmic events needs to be taken into account when using these cells for drug screening.
Tissue Models
One arrhythmogenic mechanism that fails to be captured at the single cell level is the impulse propagation in arrhythmia formation. Ectopic beats triggered by automaticity, early or delayed after depolarizations (EADs or DADs) are focal events that only lead to arrhythmia if these are propagated to the neighboring cells. Consequently, arrhythmias can only be observed in multi-cellular tissue models. Besides cellular contribution of abnormal electrophysiology, cell-cell electrical coupling affecting the conduction velocity is another factor that dictates the wave length of the AP wave front. Additionally, electrophysiological heterogeneities, including dispersion of repolarization or source-sink mismatch, are triggers for arrhythmias that only become apparent at the tissue levels.
Conduction velocity and wave front reentry can be observed in either 2D or 3D tissue models composed of a syncytium of cardiomyocytes via optical mapping. Since primary adult cardiomyocytes from animals do not form monolayers in culture, most in vitro tissue models have relied on primary neonatal or hiPSC-derived cardiomyocytes that retain morphological plasticity to reorganize and allow electrical coupling with neighboring cells. For the 2D model, cardiomyocytes plated in typical cell cultureware exhibit random cellular orientation unlike the anisotropic alignment of the myocardium. Conduction pattern is affected by the cardiomyocyte alignment, which is reflected in the increased incidence of arrhythmic events even in healthy monolayer of hiPSC-derived cardiomyocytes, making this model less accurate for testing pro-arrhythmic triggers. With advances in microfabrication technology, patterned surface to induce aligned monolayers of hiPSC-derived cardiomyocytes has been shown to reduce the incidence of arrhythmia, creating a more stable baseline tissue model for assessing pro-arrhythmic factors [96, 97].
While 2D hiPSC-cardiomyocyte-based tissue models can display arrhythmic events, there are limitations in the organization and maturity of cardiomyocytes. Specifically, 3D tissues in a patch configuration have been reported to be necessary in modeling arrhythmic phenomenon such as torsade de pointes with fluctuating excitation intervals that can only be recreated in the presence of a meandering wave origin [98]. This is not possible in a 2D configuration with a stationary wave center. Other 3D hiPSC-derived cardiomyocyte-based tissue configurations for testing arrhythmic events include linear or circular strips, small microtissues, and organoid chambers [16, 99,100,101,102,103]. These tissue models present a more mature cardiomyocyte phenotype than the 2D counterpart. However, most 3D cardiac tissues in the literature have added fibroblasts to facilitate tissue compaction. Depending on the fibroblast type, the electrical coupling with the cardiomyocytes may differ, which may result in different source-sink ratio that can affect the conduction pattern and intrinsic arrhythmogenicity [104]. Therefore, there are many factors to consider when choosing a tissue model for in vitro arrhythmia studies.
Mathematical Models for Addressing the Mechanisms of Arrhythmogenesis
Since cardiac excitability is sculpted by the beat-to-beat feedback among three nonlinear dynamic processes including Ca2+ dynamic, APs, and mechanical contractions, multiscale computational modeling offers unparalleled advantages in deciphering cardiac arrhythmia mechanisms (Fig. 1). The fundamental question is how to fully integrate experimental and clinical information and mathematical modeling to accurately simulate cardiac structural and electrophysiological properties at subcellular, cellular, tissue, and organ levels. This multiscale modeling is a challenge considering the complexity and heterogeneity of cardiac tissues. With the recent development of computational techniques, research in cardiac computational modeling has greatly expanded with the support from experimental and clinical investigators [105,106,107,108,109]. Computational models of different cell types including sinoatrial node (SAN) cells, atrial myocytes, atrioventricular (AV) node cells, Purkinje fibers, and ventricular myocytes have been developed. For understanding human cardiac diseases, many computational models across species have been developed using experimental data from animal studies for validation and comparison. Here, we will focus on the human cardiac models.
Atrial and AV Node Models
The early mathematical models of the AP from adult human atrial cells were developed by Nygren et al. and Courtemanche et al. in 1998 [110, 111]. With accumulation of human atrial experimental data, several human atrial models have been refined including the AV node [112,113,114,115,116,117,118,119,120]. The significance of these models was demonstrated by simulating the AP alternans, AF, conduction block, and AF-associated electrical remodeling in single cells, one-, two-, and three-dimensional tissues [112, 116, 121,122,123,124]. Computational models are powerful tools to study the mechanism of AF, and have the potential to guide clinical treatment of AF using catheter ablation [125, 126]. In the past decade, 3D computational modeling has been used to guide and optimize AF ablation and therapy to minimize the ablation lesions and improve clinical outcomes [107].
Ventricle Models
The first mathematical model of the AP from human ventricular myocytes was reported by Priebe and Beuckelmann in 1998 [106, 127]. The models for ventricular myocytes and tissues were further refined using larger human dataset [106, 128,129,130,131,132,133]. The models have been used to explore and predict the mechanism of initiation, maintenance, and termination of ventricular arrhythmias, including simulations of triggered activities, AP and conduction velocity restitutions, inherited arrhythmias, prediction of AP alternans, the vulnerable window, and efficacy of antiarrhythmic drugs [106]. More recently, computational models have been developed and applied to characterize ventricular arrhythmias under different clinical settings and diseased states including heart failure, cardiac ischemia, and maintenance of torsades de pointes [107, 134,135,136,137,138]. Multiscale modeling has been successfully used to demonstrate the origins of ECG morphology and abnormal ECG features resulted from diseases [139,140,141,142,143]. With the aid of the advanced computational techniques, modeling of patient-specific cardiac diseases becomes feasible with successful applications in predicting patient’s risk for ventricular arrhythmias and SCD [144, 145]. Treatment of ventricular tachycardia and defibrillator implantation have also been facilitated by using computational modeling to predict the optimal ablation targets and defibrillation locations [146, 147].
Purkinje Fiber Models
Purkinje fibers play critical roles as initiating sites for ventricular tachycardia and fibrillation. An AP model of human Purkinje fibers was first developed by Tusscher and Panfilov [148]. Based on detailed biophysical kinetic analysis of the ion channels in human Purkinje fibers, a new computational model was proposed and applied to simulate LQTS [149, 150]. The 3D Purkinje fiber network model was established based on imaging data, which integrates the Purkinje fibers with the ventricle to simulate the electrical activation sequences in the ventricle [151]. Recently, ionic currents of human Purkinje-related electrophysiology, pacemaker activity, and arrhythmogenicity were further revealed by incorporating Purkinje-specific ionic currents and Ca2+ handling [152]. The structural and functional integration of ventricular and Purkinje fiber models will significantly improve our understandings of the mechanisms of ventricular arrhythmias. Additionally, the structural and functional contribution of cardiac fibroblasts also needs to be considered [153].
SAN Models
Because of the very limited experimental data from human SAN, the development of mathematical model for human SAN lags behind the model development for human atrial and ventricular myocytes. Seemann et al. published the first human SAN AP model as part of a 3D human atria model [154]. This is followed by an AP model based on the mRNA expression of ion channels in healthy adult human SAN tissues [155]. A recent study integrates the membrane and Ca2+ clock and constructs a comprehensive mathematical model for SAN cells [156].
Virtual Heart Models
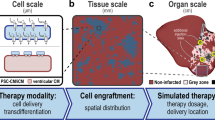
One major advantage of computational modeling is the expansion of personalized medicine by integrating patient information and clinical data into the model to develop personalized treatment strategies [157]. A recent virtual heart technology for guiding the ablation of ventricular tachycardia demonstrates the highly promising future for treatment of cardiac arrhythmias [158]. The incorporation of machine learning and artificial intelligence in the diagnosis and treatment of cardiac arrhythmias will provide powerful tool sets for the prevention, prediction, and treatment of cardiac arrhythmias (Fig. 1) [159]. Indeed, multiscale 3D whole-heart modeling has recently been developed to determine how varying parameters of cell delivery and transdifferentiation could result in focal ectopy, heart block, and reentry [20••].
Finally, the innervation, spatial, and temporal dynamic feedback at tissue and organ levels are required to be integrated into the models for understanding the beat-to-beat electrical excitability in the heart. Therefore, joint efforts of computational biologists, experimental biologists, and physicians are necessary to integrate the multi-discipline knowledge in computational modeling. Additional validation with experimental and clinical data followed by reiterative refinement of the computational models will help to pave the way for deeper understandings of the complex mechanisms of cardiac arrhythmogenesis, assist the diagnosis, and guide the treatment.
Conclusions and Perspectives
The mechanistic understanding of cardiac arrhythmogenesis is critical for the development of cardiac cell-based therapy. Animal models from multiple species provide experimental platforms for mechanistic hypothesis testing. Ex vivo and in vitro models enable the application of several cutting-edge techniques including the development of patient-specific hiPSC-CMs and CRISPR-Cas9 gene editing. Recent advances in multiscale computational models for arrhythmogenesis represent the joint efforts by computational biologists, biophysicists, experimental biologists, and physicians, and provide quantitative and powerful tools for the mechanistic understanding of arrhythmogenesis in cardiac cell-based therapy. Further integrations of experimental, computational, and the virtual heart model will provide critical insights into novel mitigation strategies for cardiac arrhythmias in cell-based therapy.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Mozaffarian D, Benjamin EJ, Go AS, Arnett DK, Blaha MJ, Cushman M, et al. Heart disease and stroke statistics—2016 update: a report from the American Heart Association. Circulation. 2016;133:e38–360.
Dimmeler S, Zeiher AM. Cell therapy of acute myocardial infarction: open questions. Cardiology. 2009;113:155–60.
Dimmeler S, Losordo D. Stem cells review series: an introduction. Circ Res. 2011;109:907–9.
Zhang WY, Ebert AD, Narula J, Wu JC. Imaging cardiac stem cell therapy: translations to human clinical studies. J Cardiovasc Transl Res. 2011;4:514–22.
Sirish P, Thai PN, Lee JH, Yang J, Zhang XD, Ren L, et al. Suppression of inflammation and fibrosis using soluble epoxide hydrolase inhibitors enhances cardiac stem cell-based therapy. Stem Cells Transl Med. 2020;9:1570–84.
• Lalit PA, Hei DJ, Raval AN, Kamp TJ. Induced pluripotent stem cells for post-myocardial infarction repair: remarkable opportunities and challenges. Circ Res. 2014;114:1328–45 The study provides an overview of the opportunities and challenges of using induced pluripotent stem cells for cardiac cell-based therapy.
•• Chong JJ, Murry CE. Cardiac regeneration using pluripotent stem cells—progression to large animal models. Stem Cell Res. 2014;13:654–65 The review article discussed cardiac cell-based therapy in non-human primate models including large-scale remuscularization, electromechanical coupling and short-term arrhythmias.
Liu YW, Chen B, Yang X, Fugate JA, Kalucki FA, Futakuchi-Tsuchida A, et al. Human embryonic stem cell-derived cardiomyocytes restore function in infarcted hearts of non-human primates. Nat Biotechnol. 2018;36:597–605.
Almeida SO, Skelton RJ, Adigopula S, Ardehali R. Arrhythmia in stem cell transplantation. Card Electrophysiol Clin. 2015;7:357–70.
• Clauss S, Bleyer C, Schuttler D, Tomsits P, Renner S, Klymiuk N, et al. Animal models of arrhythmia: classic electrophysiology to genetically modified large animals. Nat Rev Cardiol. 2019;16:457–75 This recent review article provides an overview of different models for cardiac arrhythmia research as well as advantages and disadvantages for each model.
Choy L, Yeo JM, Tse V, Chan SP, Tse G. Cardiac disease and arrhythmogenesis: mechanistic insights from mouse models. Int J Cardiol Heart Vasc. 2016;12:1–10.
Dobrev D, Wehrens XHT. Mouse models of cardiac arrhythmias. Circ Res. 2018;123:332–4.
Huang CL. Murine electrophysiological models of cardiac arrhythmogenesis. Physiol Rev. 2017;97:283–409.
Sommariva E, Stadiotti I, Perrucci GL, Tondo C, Pompilio G. Cell models of arrhythmogenic cardiomyopathy: advances and opportunities. Dis Model Mech. 2017;10:823–35.
Pourrier M, Fedida D. The emergence of human induced pluripotent stem cell-derived cardiomyocytes (hiPSC-CMs) as a platform to model arrhythmogenic diseases. Int J Mol Sci. 2020;21:657.
Goldfracht I, Protze S, Shiti A, Setter N, Gruber A, Shaheen N, et al. Generating ring-shaped engineered heart tissues from ventricular and atrial human pluripotent stem cell-derived cardiomyocytes. Nat Commun. 2020;11:75.
Casini S, Verkerk AO, Remme CA. Human iPSC-derived cardiomyocytes for investigation of disease mechanisms and therapeutic strategies in inherited arrhythmia syndromes: strengths and limitations. Cardiovasc Drugs Ther. 2017;31:325–44.
Sinnecker D, Goedel A, Laugwitz KL, Moretti A. Induced pluripotent stem cell-derived cardiomyocytes: a versatile tool for arrhythmia research. Circ Res. 2013;112:961–8.
Shinnawi R, Gepstein L. iPCS cell modeling of inherited cardiac arrhythmias. Curr Treat Options Cardiovasc Med. 2014;16:331.
•• Yu JK, Franceschi W, Huang Q, Pashakhanloo F, Boyle PM, Trayanova NA. A comprehensive, multiscale framework for evaluation of arrhythmias arising from cell therapy in the whole post-myocardial infarcted heart. Sci Rep. 2019;9:9238 The study provides computational framework to explore how varying parameters of cell delivery and transdifferentiation may contribute to cardiac arrhythmias, focal ectopy, heart block, and reentry, in cell-based therapy.
Jin H, Chemaly ER, Lee A, Kho C, Hadri L, Hajjar RJ, et al. Mechanoelectrical remodeling and arrhythmias during progression of hypertrophy. FASEB J. 2010;24:451–63.
Winterton SJ, Turner MA, O'Gorman DJ, Flores NA, Sheridan DJ. Hypertrophy causes delayed conduction in human and guinea pig myocardium: accentuation during ischaemic perfusion. Cardiovasc Res. 1994;28:47–54.
Ahmmed GU, Dong PH, Song G, Ball NA, Xu Y, Walsh RA, et al. Changes in Ca2+ cycling proteins underlie cardiac action potential prolongation in a pressure-overloaded guinea pig model with cardiac hypertrophy and failure. Circ Res. 2000;86:558–70.
Pogwizd Steven M, Qi M, Yuan W, Samarel Allen M, Bers Donald M. Upregulation of Na+/Ca2+ exchanger expression and function in an arrhythmogenic rabbit model of heart failure. Circ Res. 1999;85:1009–19.
Ishikawa K, Aguero J, Oh JG, Hammoudi N, Fish LA, Leonardson L, et al. Increased stiffness is the major early abnormality in a pig model of severe aortic stenosis and predisposes to congestive heart failure in the absence of systolic dysfunction. J Am Heart Assoc. 2015;4:e001925.
Pogwizd SM, Bers DM. Rabbit models of heart disease. Drug Discov Today Dis Model. 2008;5:185–93.
• Hegyi B, Bossuyt J, Griffiths LG, Shimkunas R, Coulibaly Z, Jian Z, et al. Complex electrophysiological remodeling in postinfarction ischemic heart failure. Proc Natl Acad Sci U S A. 2018;115:E3036–44 The study provides mechanistic understanding for the contributions of multiple ionic currents and their remodeling in heart disease, and highlight the need to consider the integration of multiple ionic currents in designing therapeutic strategies for treating arrhythmias in heart failure.
Corr PB, Pearle DL, Hinton JR, Roberts WC, Gillis RA. Site of myocardial infarction. A determinant of the cardiovascular changes induced in the cat by coronary occlusion. Circ Res. 1976;39:840–7.
Wijffels MC, Kirchhof CJ, Dorland R, Allessie MA. Atrial fibrillation begets atrial fibrillation. A study in awake chronically instrumented goats. Circulation. 1995;92:1954–68.
Shi Y, Ducharme A, Li D, Gaspo R, Nattel S, Tardif JC. Remodeling of atrial dimensions and emptying function in canine models of atrial fibrillation. Cardiovasc Res. 2001;52:217–25.
Filgueiras-Rama D, Price NF, Martins RP, Yamazaki M, Avula UMR, Kaur K, et al. Long-term frequency gradients during persistent atrial fibrillation in sheep are associated with stable sources in the left atrium. Circ Arrhythm Electrophysiol. 2012;5:1160–7.
Citerni C, Kirchhoff J, Olsen LH, Sattler SM, Gentilini F, Forni M, et al. Characterization of atrial and ventricular structural remodeling in a porcine model of atrial fibrillation induced by atrial tachypacing. Front Vet Sci. 2020;7:179.
Jia X, Zheng S, Xie X, Zhang Y, Wang W, Wang Z, et al. MicroRNA-1 accelerates the shortening of atrial effective refractory period by regulating KCNE1 and KCNB2 expression: an atrial tachypacing rabbit model. PLoS One. 2013;8:e85639.
Milani-Nejad N, Janssen PML. Small and large animal models in cardiac contraction research: advantages and disadvantages. Pharmacol Ther. 2014;141:235–49.
Kaab S, Nuss HB, Chiamvimonvat N, O'Rourke B, Pak PH, Kass DA, et al. Ionic mechanism of action potential prolongation in ventricular myocytes from dogs with pacing-induced heart failure. Circ Res. 1996;78:262–73.
Ling LH, Kistler PM, Kalman JM, Schilling RJ, Hunter RJ. Comorbidity of atrial fibrillation and heart failure. Nat Rev Cardiol. 2016;13:131–47.
Wilde AAM, Bezzina CR. Genetics of cardiac arrhythmias. Heart. 2005;91:1352–8.
Sabir IN, Fraser JA, Killeen MJ, Grace AA, Huang CL. The contribution of refractoriness to arrhythmic substrate in hypokalemic Langendorff-perfused murine hearts. Pflugers Arch. 2007;454:209–22.
Wiegerinck RF, van Veen TA, Belterman CN, Schumacher CA, Noorman M, de Bakker JM, et al. Transmural dispersion of refractoriness and conduction velocity is associated with heterogeneously reduced connexin43 in a rabbit model of heart failure. Heart Rhythm. 2008;5:1178–85.
Ravelli F, Allessie M. Effects of atrial dilatation on refractory period and vulnerability to atrial fibrillation in the isolated Langendorff-perfused rabbit heart. Circulation. 1997;96:1686–95.
Yamazaki M, Morgenstern S, Klos M, Campbell K, Buerkel D, Kalifa J. Left atrial coronary perfusion territories in isolated sheep hearts: implications for atrial fibrillation maintenance. Heart Rhythm. 2010;7:1501–8.
Sabir IN, Ma N, Jones VJ, Goddard CA, Zhang Y, Kalin A, et al. Alternans in genetically modified langendorff-perfused murine hearts modeling catecholaminergic polymorphic ventricular tachycardia. Front Physiol. 2010;1:126.
Tse G, Hothi SS, Grace AA, Huang CL. Ventricular arrhythmogenesis following slowed conduction in heptanol-treated, Langendorff-perfused mouse hearts. J Physiol Sci. 2012;62:79–92.
de Bakker JM, Coronel R, Tasseron S, Wilde AA, Opthof T, Janse MJ, et al. Ventricular tachycardia in the infarcted, Langendorff-perfused human heart: role of the arrangement of surviving cardiac fibers. J Am Coll Cardiol. 1990;15:1594–607.
Gerhardy A, Scholtysik G, Schaad A, Haltiner R, Hess T. Generating and influencing torsades de pointes-like polymorphic ventricular tachycardia in isolated guinea pig hearts. Basic Res Cardiol. 1998;93:285–94.
Lehmann HI, Richter D, Prokesch H, Graeff C, Prall M, Simoniello P, et al. Atrioventricular node ablation in Langendorff-perfused porcine hearts using carbon ion particle therapy: methods and an in vivo feasibility investigation for catheter-free ablation of cardiac arrhythmias. Circ Arrhythm Electrophysiol. 2015;8:429–38.
Janse MJ, van Capelle FJ, Freud GE, Durrer D. Circus movement within the AV node as a basis for supraventricular tachycardia as shown by multiple microelectrode recording in the isolated rabbit heart. Circ Res. 1971;28:403–14.
Killeen MJ, Thomas G, Gurung IS, Goddard CA, Fraser JA, Mahaut-Smith MP, et al. Arrhythmogenic mechanisms in the isolated perfused hypokalaemic murine heart. Acta Physiol (Oxford). 2007;189:33–46.
Maruyama M, Ai T, Chua SK, Park HW, Lee YS, Shen MJ, et al. Hypokalemia promotes late phase 3 early afterdepolarization and recurrent ventricular fibrillation during isoproterenol infusion in Langendorff perfused rabbit ventricles. Heart Rhythm. 2014;11:697–706.
Ghais NS, Zhang Y, Mistry B, Grace AA, Huang CL. Anti-arrhythmic effects of cyclopiazonic acid in Langendorff-perfused murine hearts. Prog Biophys Mol Biol. 2008;98:281–8.
Tse G, Sun B, Wong ST, Tse V, Yeo JM. Anti-arrhythmic effects of hypercalcemia in hyperkalemic, Langendorff-perfused mouse hearts. Biomed Rep. 2016;5:301–10.
Tse G, Tse V, Yeo JM, Sun B. Atrial anti-arrhythmic effects of heptanol in Langendorff-perfused mouse hearts. PLoS One. 2016;11:e0148858.
Cao ZZ, Tian YJ, Hao J, Zhang PH, Liu ZP, Jiang WZ, et al. Barbaloin inhibits ventricular arrhythmias in rabbits by modulating voltage-gated ion channels. Acta Pharmacol Sin. 2018;39:357–70.
Tse G, Tse V, Yeo JM. Ventricular anti-arrhythmic effects of heptanol in hypokalaemic, Langendorff-perfused mouse hearts. Biomed Rep. 2016;4:313–24.
Olejnickova V, Novakova M, Provaznik I. Isolated heart models: cardiovascular system studies and technological advances. Med Biol Eng Comput. 2015;53:669–78.
Motayagheni N. Modified Langendorff technique for mouse heart cannulation: improved heart quality and decreased risk of ischemia. MethodsX. 2017;4:508–12.
Swift LM, Jaimes R 3rd, McCullough D, Burke M, Reilly M, Maeda T, et al. Optocardiography and electrophysiology studies of ex vivo Langendorff-perfused hearts. J Vis Exp. 2019;153:e60472.
Sill B, Hammer PE, Cowan DB. Optical mapping of Langendorff-perfused rat hearts. J Vis Exp. 2009;30:e1138.
Nygren A, Kondo C, Clark RB, Giles WR. Voltage-sensitive dye mapping in Langendorff-perfused rat hearts. Am J Physiol Heart Circ Physiol. 2003;284:H892–902.
Boyle PM, Karathanos TV, Trayanova NA. Cardiac optogenetics: 2018. JACC Clin Electrophysiol. 2018;4:155–67.
Aggarwal R, Boyden PA. Diminished Ca2+ and Ba2+ currents in myocytes surviving in the epicardial border zone of the 5-day infarcted canine heart. Circ Res. 1995;77:1180–91.
Jiang M, Cabo C, Yao J, Boyden PA, Tseng G. Delayed rectifier K currents have reduced amplitudes and altered kinetics in myocytes from infarcted canine ventricle. Cardiovasc Res. 2000;48:34–43.
O'Rourke B, Kass DA, Tomaselli GF, Kaab S, Tunin R, Marban E. Mechanisms of altered excitation-contraction coupling in canine tachycardia-induced heart failure, I: experimental studies. Circ Res. 1999;84:562–70.
Ahuja D, Saenz-Robles MT, Pipas JM. SV40 large T antigen targets multiple cellular pathways to elicit cellular transformation. Oncogene. 2005;24:7729–45.
Claycomb WC, Lanson NA Jr, Stallworth BS, Egeland DB, Delcarpio JB, Bahinski A, et al. HL-1 cells: a cardiac muscle cell line that contracts and retains phenotypic characteristics of the adult cardiomyocyte. Proc Natl Acad Sci U S A. 1998;95:2979–84.
Hong JH, Choi JH, Kim TY, Lee KJ. Spiral reentry waves in confluent layer of HL-1 cardiomyocyte cell lines. Biochem Biophys Res Commun. 2008;377:1269–73.
Davidson MM, Nesti C, Palenzuela L, Walker WF, Hernandez E, Protas L, et al. Novel cell lines derived from adult human ventricular cardiomyocytes. J Mol Cell Cardiol. 2005;39:133–47.
Itzhaki I, Maizels L, Huber I, Zwi-Dantsis L, Caspi O, Winterstern A, et al. Modelling the long QT syndrome with induced pluripotent stem cells. Nature. 2011;471:225–9.
Lahti AL, Kujala VJ, Chapman H, Koivisto AP, Pekkanen-Mattila M, Kerkela E, et al. Model for long QT syndrome type 2 using human iPS cells demonstrates arrhythmogenic characteristics in cell culture. Dis Model Mech. 2012;5:220–30.
Carvajal-Vergara X, Sevilla A, D'Souza SL, Ang YS, Schaniel C, Lee DF, et al. Patient-specific induced pluripotent stem-cell-derived models of LEOPARD syndrome. Nature. 2010;465:808–12.
Veerman CC, Mengarelli I, Guan K, Stauske M, Barc J, Tan HL, et al. hiPSC-derived cardiomyocytes from Brugada syndrome patients without identified mutations do not exhibit clear cellular electrophysiological abnormalities. Sci Rep. 2016;6:30967.
Sasaki K, Makiyama T, Yoshida Y, Wuriyanghai Y, Kamakura T, Nishiuchi S, et al. Patient-specific human induced pluripotent stem cell model assessed with electrical pacing validates S107 as a potential therapeutic agent for catecholaminergic polymorphic ventricular tachycardia. PLoS One. 2016;11:e0164795.
Shinnawi R, Shaheen N, Huber I, Shiti A, Arbel G, Gepstein A, et al. Modeling reentry in the short QT syndrome with human-induced pluripotent stem cell-derived cardiac cell sheets. J Am Coll Cardiol. 2019;73:2310–24.
Lee EK, Tran DD, Keung W, Chan P, Wong G, Chan CW, et al. Machine learning of human pluripotent stem cell-derived engineered cardiac tissue contractility for automated drug classification. Stem Cell Rep. 2017;9:1560–72.
Saleem U, van Meer BJ, Katili PA, Mohd Yusof NAN, Mannhardt I, Garcia AK, et al. Blinded, multicenter evaluation of drug-induced changes in contractility using human-induced pluripotent stem cell-derived cardiomyocytes. Toxicol Sci. 2020;176:103–23.
Feric NT, Pallotta I, Singh R, Bogdanowicz DR, Gustilo M, Chaudhary K, et al. Engineered cardiac tissues generated in the Biowire II: a platform for human-based drug discovery. Toxicol Sci. 2019;172:89–97.
McKeithan WL, Feyen DAM, Bruyneel AAN, Okolotowicz KJ, Ryan DA, Sampson KJ, et al. Reengineering an antiarrhythmic drug using patient hiPSC cardiomyocytes to improve therapeutic potential and reduce toxicity. Cell Stem Cell. 2020;27:813–21 e816.
Yechikov S, Copaciu R, Gluck JM, Deng W, Chiamvimonvat N, Chan JW, et al. Same-single-cell analysis of pacemaker-specific markers in human induced pluripotent stem cell-derived cardiomyocyte subtypes classified by electrophysiology. Stem Cells. 2016;34:2670–80.
Moretti A, Bellin M, Welling A, Jung CB, Lam JT, Bott-Flugel L, et al. Patient-specific induced pluripotent stem-cell models for long-QT syndrome. N Engl J Med. 2010;363:1397–409.
Veevers J, Farah EN, Corselli M, Witty AD, Palomares K, Vidal JG, et al. Cell-surface marker signature for enrichment of ventricular cardiomyocytes derived from human embryonic stem cells. Stem Cell Rep. 2018;11:828–41.
Zhang JZ, Termglinchan V, Shao NY, Itzhaki I, Liu C, Ma N, et al. A human iPSC double-reporter system enables purification of cardiac lineage subpopulations with distinct function and drug response profiles. Cell Stem Cell. 2019;24:802–11 e805.
Karakikes I, Senyei GD, Hansen J, Kong CW, Azeloglu EU, Stillitano F, et al. Small molecule-mediated directed differentiation of human embryonic stem cells toward ventricular cardiomyocytes. Stem Cells Transl Med. 2014;3:18–31.
Weng Z, Kong CW, Ren L, Karakikes I, Geng L, He J, et al. A simple, cost-effective but highly efficient system for deriving ventricular cardiomyocytes from human pluripotent stem cells. Stem Cells Dev. 2014;23:1704–16.
Lee JH, Protze SI, Laksman Z, Backx PH, Keller GM. Human pluripotent stem cell-derived atrial and ventricular cardiomyocytes develop from distinct mesoderm populations. Cell Stem Cell. 2017;21:179–94 e174.
Protze SI, Liu J, Nussinovitch U, Ohana L, Backx PH, Gepstein L, et al. Sinoatrial node cardiomyocytes derived from human pluripotent cells function as a biological pacemaker. Nat Biotechnol. 2017;35:56–68.
Yechikov S, Kao HKJ, Chang CW, Pretto D, Zhang XD, Sun YH, et al. NODAL inhibition promotes differentiation of pacemaker-like cardiomyocytes from human induced pluripotent stem cells. Stem Cell Res. 2020;49:102043.
Ren J, Han P, Ma X, Farah EN, Bloomekatz J, Zeng XI, et al. Canonical Wnt5b signaling directs outlying Nkx2.5+ mesoderm into pacemaker cardiomyocytes. Dev Cell. 2019;50:729–43 e725.
Liang W, Han P, Kim EH, Mak J, Zhang R, Torrente AG, et al. Canonical Wnt signaling promotes pacemaker cell specification of cardiac mesodermal cells derived from mouse and human embryonic stem cells. Stem Cells. 2020;38:352–68.
Sartiani L, Bettiol E, Stillitano F, Mugelli A, Cerbai E, Jaconi ME. Developmental changes in cardiomyocytes differentiated from human embryonic stem cells: a molecular and electrophysiological approach. Stem Cells. 2007;25:1136–44.
Lieu DK, Fu JD, Chiamvimonvat N, Tung KC, McNerney GP, Huser T, et al. Mechanism-based facilitated maturation of human pluripotent stem cell-derived cardiomyocytes. Circ Arrhythm Electrophysiol. 2013;6:191–201.
Eng G, Lee BW, Protas L, Gagliardi M, Brown K, Kass RS, et al. Autonomous beating rate adaptation in human stem cell-derived cardiomyocytes. Nat Commun. 2016;7:10312.
Ruan JL, Tulloch NL, Saiget M, Paige SL, Razumova MV, Regnier M, et al. Mechanical stress promotes maturation of human myocardium from pluripotent stem cell-derived progenitors. Stem Cells. 2015;33:2148–57.
Tulloch NL, Muskheli V, Razumova MV, Korte FS, Regnier M, Hauch KD, et al. Growth of engineered human myocardium with mechanical loading and vascular coculture. Circ Res. 2011;109:47–59.
Feyen DAM, McKeithan WL, Bruyneel AAN, Spiering S, Hormann L, Ulmer B, et al. Metabolic maturation media improve physiological function of human iPSC-derived cardiomyocytes. Cell Rep. 2020;32:107925.
Nunes SS, Miklas JW, Liu J, Aschar-Sobbi R, Xiao Y, Zhang B, et al. Biowire: a platform for maturation of human pluripotent stem cell-derived cardiomyocytes. Nat Methods. 2013;10:781–7.
Wang J, Chen A, Lieu DK, Karakikes I, Chen G, Keung W, et al. Effect of engineered anisotropy on the susceptibility of human pluripotent stem cell-derived ventricular cardiomyocytes to arrhythmias. Biomaterials. 2013;34:8878–86.
Shum AM, Che H, Wong AO, Zhang C, Wu H, Chan CW, et al. A micropatterned human pluripotent stem cell-based ventricular cardiac anisotropic sheet for visualizing drug-induced arrhythmogenicity. Adv Mater. 2017;29:1602448.
Kawatou M, Masumoto H, Fukushima H, Morinaga G, Sakata R, Ashihara T, et al. Modelling torsade de pointes arrhythmias in vitro in 3D human iPS cell-engineered heart tissue. Nat Commun. 2017;8:1078.
Zhao Y, Rafatian N, Feric NT, Cox BJ, Aschar-Sobbi R, Wang EY, et al. A platform for generation of chamber-specific cardiac tissues and disease modeling. Cell. 2019;176:913–27 e918.
Schaaf S, Shibamiya A, Mewe M, Eder A, Stohr A, Hirt MN, et al. Human engineered heart tissue as a versatile tool in basic research and preclinical toxicology. PLoS One. 2011;6:e26397.
Li RA, Keung W, Cashman TJ, Backeris PC, Johnson BV, Bardot ES, et al. Bioengineering an electro-mechanically functional miniature ventricular heart chamber from human pluripotent stem cells. Biomaterials. 2018;163:116–27.
Giacomelli E, Bellin M, Sala L, van Meer BJ, Tertoolen LG, Orlova VV, et al. Three-dimensional cardiac microtissues composed of cardiomyocytes and endothelial cells co-differentiated from human pluripotent stem cells. Development. 2017;144:1008–17.
Giacomelli E, Meraviglia V, Campostrini G, Cochrane A, Cao X, van Helden RWJ, et al. Human-iPSC-derived cardiac stromal cells enhance maturation in 3D cardiac microtissues and reveal non-cardiomyocyte contributions to heart disease. Cell Stem Cell. 2020;26:862–79 e811.
Zhang J, Tao R, Campbell KF, Carvalho JL, Ruiz EC, Kim GC, et al. Functional cardiac fibroblasts derived from human pluripotent stem cells via second heart field progenitors. Nat Commun. 2019;10:2238.
Mayourian J, Sobie EA, Costa KD. An introduction to computational modeling of cardiac electrophysiology and arrhythmogenicity. Methods Mol Biol. 2018;1816:17–35.
Moreno JD, Clancy CE. Using computational modeling to predict arrhythmogenesis and antiarrhythmic therapy. Drug Discov Today Dis Model. 2009;6:71–84.
Niederer SA, Lumens J, Trayanova NA. Computational models in cardiology. Nat Rev Cardiol. 2019;16:100–11.
Roberts BN, Yang PC, Behrens SB, Moreno JD, Clancy CE. Computational approaches to understand cardiac electrophysiology and arrhythmias. Am J Physiol Heart Circ Physiol. 2012;303:H766–83.
Deng D, Jiao P, Ye X, Xia L. An image-based model of the whole human heart with detailed anatomical structure and fiber orientation. Comput Math Methods Med. 2012;2012:891070.
Nygren A, Fiset C, Firek L, Clark JW, Lindblad DS, Clark RB, et al. Mathematical model of an adult human atrial cell: the role of K+ currents in repolarization. Circ Res. 1998;82:63–81.
Courtemanche M, Ramirez RJ, Nattel S. Ionic mechanisms underlying human atrial action potential properties: insights from a mathematical model. Am J Phys. 1998;275:H301–21.
Wilhelms M, Hettmann H, Maleckar MM, Koivumaki JT, Dossel O, Seemann G. Benchmarking electrophysiological models of human atrial myocytes. Front Physiol. 2012;3:487.
Grandi E, Pandit SV, Voigt N, Workman AJ, Dobrev D, Jalife J, et al. Human atrial action potential and Ca2+ model: sinus rhythm and chronic atrial fibrillation. Circ Res. 2011;109:1055–66.
Maleckar MM, Greenstein JL, Trayanova NA, Giles WR. Mathematical simulations of ligand-gated and cell-type specific effects on the action potential of human atrium. Prog Biophys Mol Biol. 2008;98:161–70.
Podziemski P, Zebrowski JJ. A simple model of the right atrium of the human heart with the sinoatrial and atrioventricular nodes included. J Clin Monit Comput. 2013;27:481–98.
Jackowska-Zduniak B, Forys U. Mathematical model of the atrioventricular nodal double response tachycardia and double-fire pathology. Math Biosci Eng. 2016;13:1143–58.
Inada S, Hancox JC, Zhang H, Boyett MR. One-dimensional mathematical model of the atrioventricular node including atrio-nodal, nodal, and nodal-his cells. Biophys J. 2009;97:2117–27.
Climent AM, Guillem MS, Zhang Y, Millet J, Mazgalev TN. Functional mathematical model of dual pathway AV nodal conduction. Am J Physiol Heart Circ Physiol. 2011;300:H1393–401.
orgensen P, Schafer C, Guerra PG, Talajic M, Nattel S, Glass L. A mathematical model of human atrioventricular nodal function incorporating concealed conduction. Bull Math Biol. 2002;64:1083–99.
Nayebpour M, Talajic M, Nattel S. Quantitation of dynamic AV nodal properties and application to predict rate-dependent AV conduction. Am J Phys. 1991;261:H292–300.
Lim B, Kim J, Hwang M, Song JS, Lee JK, Yu HT, et al. In situ procedure for high-efficiency computational modeling of atrial fibrillation reflecting personal anatomy, fiber orientation, fibrosis, and electrophysiology. Sci Rep. 2020;10:2417.
Trayanova NA. Mathematical approaches to understanding and imaging atrial fibrillation: significance for mechanisms and management. Circ Res. 2014;114:1516–31.
Aronis KN, Ali RL, Liang JA, Zhou S, Trayanova NA. Understanding AF mechanisms through computational modelling and simulations. Arrhythmia Electrophysiol Rev. 2019;8:210–9.
Talajic M, Papadatos D, Villemaire C, Glass L, Nattel S. A unified model of atrioventricular nodal conduction predicts dynamic changes in Wenckebach periodicity. Circ Res. 1991;68:1280–93.
Grandi E, Dobrev D, Heijman J. Computational modeling: what does it tell us about atrial fibrillation therapy? Int J Cardiol. 2019;287:155–61.
Boyle PM, Zghaib T, Zahid S, Ali RL, Deng D, Franceschi WH, et al. Computationally guided personalized targeted ablation of persistent atrial fibrillation. Nat Biomed Eng. 2019;3:870–9.
Priebe L, Beuckelmann DJ. Simulation study of cellular electric properties in heart failure. Circ Res. 1998;82:1206–23.
ten Tusscher KH, Panfilov AV. Alternans and spiral breakup in a human ventricular tissue model. Am J Physiol Heart Circ Physiol. 2006;291:H1088–100.
Ten Tusscher KH, Bernus O, Hren R, Panfilov AV. Comparison of electrophysiological models for human ventricular cells and tissues. Prog Biophys Mol Biol. 2006;90:326–45.
ten Tusscher KH, Noble D, Noble PJ, Panfilov AV. A model for human ventricular tissue. Am J Physiol Heart Circ Physiol. 2004;286:H1573–89.
Iyer V, Mazhari R, Winslow RL. A computational model of the human left-ventricular epicardial myocyte. Biophys J. 2004;87:1507–25.
Keldermann RH, ten Tusscher KH, Nash MP, Hren R, Taggart P, Panfilov AV. Effect of heterogeneous APD restitution on VF organization in a model of the human ventricles. Am J Physiol Heart Circ Physiol. 2008;294:H764–74.
Ten Tusscher KH, Hren R, Panfilov AV. Organization of ventricular fibrillation in the human heart. Circ Res. 2007;100:e87–101.
Dutta S, Minchole A, Zacur E, Quinn TA, Taggart P, Rodriguez B. Early afterdepolarizations promote transmural reentry in ischemic human ventricles with reduced repolarization reserve. Prog Biophys Mol Biol. 2016;120:236–48.
Kazbanov IV, Clayton RH, Nash MP, Bradley CP, Paterson DJ, Hayward MP, et al. Effect of global cardiac ischemia on human ventricular fibrillation: insights from a multi-scale mechanistic model of the human heart. PLoS Comput Biol. 2014;10:e1003891.
Van Nieuwenhuyse E, Seemann G, Panfilov AV, Vandersickel N. Effects of early afterdepolarizations on excitation patterns in an accurate model of the human ventricles. PLoS One. 2017;12:e0188867.
Bayer JD, Lalani GG, Vigmond EJ, Narayan SM, Trayanova NA. Mechanisms linking electrical alternans and clinical ventricular arrhythmia in human heart failure. Heart Rhythm. 2016;13:1922–31.
Vandersickel N, de Boer TP, Vos MA, Panfilov AV. Perpetuation of torsade de pointes in heterogeneous hearts: competing foci or re-entry? J Physiol. 2016;594:6865–78.
Bacharova L, Mateasik A, Krause R, Prinzen FW, Auricchio A, Potse M. The effect of reduced intercellular coupling on electrocardiographic signs of left ventricular hypertrophy. J Electrocardiol. 2011;44:571–6.
Keller DU, Weiss DL, Dossel O, Seemann G. Influence of I(Ks) heterogeneities on the genesis of the T-wave: a computational evaluation. IEEE Trans Biomed Eng. 2012;59:311–22.
Nguyen UC, Potse M, Regoli F, Caputo ML, Conte G, Murzilli R, et al. An in-silico analysis of the effect of heart position and orientation on the ECG morphology and vectorcardiogram parameters in patients with heart failure and intraventricular conduction defects. J Electrocardiol. 2015;48:617–25.
Sadrieh A, Domanski L, Pitt-Francis J, Mann SA, Hodkinson EC, Ng CA, et al. Multiscale cardiac modelling reveals the origins of notched T waves in long QT syndrome type 2. Nat Commun. 2014;5:5069.
Chen X, Hu Y, Fetics BJ, Berger RD, Trayanova NA. Unstable QT interval dynamics precedes ventricular tachycardia onset in patients with acute myocardial infarction: a novel approach to detect instability in QT interval dynamics from clinical ECG. Circ Arrhythm Electrophysiol. 2011;4:858–66.
Deng D, Arevalo HJ, Prakosa A, Callans DJ, Trayanova NA. A feasibility study of arrhythmia risk prediction in patients with myocardial infarction and preserved ejection fraction. Europace. 2016;18:iv60–6.
Arevalo HJ, Vadakkumpadan F, Guallar E, Jebb A, Malamas P, Wu KC, et al. Arrhythmia risk stratification of patients after myocardial infarction using personalized heart models. Nat Commun. 2016;7:11437.
Ashikaga H, Arevalo H, Vadakkumpadan F, Blake RC 3rd, Bayer JD, Nazarian S, et al. Feasibility of image-based simulation to estimate ablation target in human ventricular arrhythmia. Heart Rhythm. 2013;10:1109–16.
Rantner LJ, Vadakkumpadan F, Spevak PJ, Crosson JE, Trayanova NA. Placement of implantable cardioverter-defibrillators in paediatric and congenital heart defect patients: a pipeline for model generation and simulation prediction of optimal configurations. J Physiol. 2013;591:4321–34.
Tusscher KH, Panfilov AV. Modelling of the ventricular conduction system. Prog Biophys Mol Biol. 2008;96:152–70.
Sampson KJ, Iyer V, Marks AR, Kass RS. A computational model of Purkinje fibre single cell electrophysiology: implications for the long QT syndrome. J Physiol. 2010;588:2643–55.
Iyer V, Sampson KJ, Kass RS. Modeling tissue- and mutation-specific electrophysiological effects in the long QT syndrome: role of the Purkinje fiber. PLoS One. 2014;9:e97720.
Liu BR, Cherry EM. Image-based structural modeling of the cardiac Purkinje network. Biomed Res Int. 2015;2015:621034.
Trovato C, Passini E, Nagy N, Varro A, Abi-Gerges N, Severi S, et al. Human Purkinje in silico model enables mechanistic investigations into automaticity and pro-arrhythmic abnormalities. J Mol Cell Cardiol. 2020;142:24–38.
Zeigler AC, Richardson WJ, Holmes JW, Saucerman JJ. Computational modeling of cardiac fibroblasts and fibrosis. J Mol Cell Cardiol. 2016;93:73–83.
Seemann G, Hoper C, Sachse FB, Dossel O, Holden AV, Zhang H. Heterogeneous three-dimensional anatomical and electrophysiological model of human atria. Philos Trans A Math Phys Eng Sci. 2006;364:1465–81.
Chandler NJ, Greener ID, Tellez JO, Inada S, Musa H, Molenaar P, et al. Molecular architecture of the human sinus node: insights into the function of the cardiac pacemaker. Circulation. 2009;119:1562–75.
Fabbri A, Fantini M, Wilders R, Severi S. Computational analysis of the human sinus node action potential: model development and effects of mutations. J Physiol. 2017;595:2365–96.
Trayanova NA, Boyle PM, Nikolov PP. Personalized imaging and modeling strategies for arrhythmia prevention and therapy. Curr Opin Biomed Eng. 2018;5:21–8.
Prakosa A, Arevalo HJ, Deng D, Boyle PM, Nikolov PP, Ashikaga H, et al. Personalized virtual-heart technology for guiding the ablation of infarct-related ventricular tachycardia. Nat Biomed Eng. 2018;2:732–40.
Feeny AK, Chung MK, Madabhushi A, Attia ZI, Cikes M, Firouznia M, et al. Artificial intelligence and machine learning in arrhythmias and cardiac electrophysiology. Circ Arrhythm Electrophysiol. 2020;13:e007952.
Funding
The study was supported in part by American Heart Association (AHA) Beginning Grant-in-Aid 14BGIA18870087 and National Institutes of Health (NIH) R56 HL138392 (XZ), NIH R01 HL085727, NIH R01 HL085844, and NIH R01 HL137228; Research Award from the Rosenfeld Heart Foundation; VA Merit Review Grants I01 BX000576 and I01 CX001490 (NC), Postdoctoral Fellowship from NIH T32 Training Grant in Basic & Translational Cardiovascular Science (NIH T32 HL086350) and NIH F32 HL149288 Postdoctoral Research Fellowship (PNT); and California Institute of Regenerative Medicine (CIRM) Basic Biology Grant (DISC2-10120) (DKL). NC is the holder of the Roger Tatarian Endowed Professorship in Cardiovascular Medicine and a part-time staff physician at VA Northern California Health Care System, Mather, CA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the author
Conflict of Interest
Dr. Lieu reports personal fees from Novoheart Ltd., outside the submitted work.
The other authors declare that they have no conflict of interestrs.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Regenerative Medicine
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, XD., Thai, P.N., Lieu, D.K. et al. Model Systems for Addressing Mechanism of Arrhythmogenesis in Cardiac Repair. Curr Cardiol Rep 23, 72 (2021). https://doi.org/10.1007/s11886-021-01498-z
Accepted:
Published:
DOI: https://doi.org/10.1007/s11886-021-01498-z