Abstract
The pandemic of Severe Acute Respiratory Syndrome Coronavirus-2 has affected millions of people worldwide with common symptoms of fever, cough, and respiratory complications. The pandemic has posed a huge challenge to emergency health services due to unavailability of potent therapeutic drugs. The proteins associated with the viral pathogenesis has been identified as suitable targets for drug design and warrants effective drug discovery to abate COVID-19. The papain-like protease (PLpro), nucleocapsid (N), main protease (Mpro) and non-structural protein (nsp12) of SARS-CoV-2, key component of processing of viral polyproteins, transcription, assembly and replication. On this streak, present study evaluated the interaction of ligand 2,4-diacetylphloroglucinol (DAPG) with viral proteins using molecular docking with (i) AutoDock 4.2.6 and (ii) AutoDock Vina followed by molecular dynamic simulation studies of protein-ligand complex configuration. The analysis revealed that PLpro (3E9S) and N (4J3K) protein corresponds to the highest docking score and therefore, selected for molecular dynamics simulation study (100 ns). The study comprised analysis of parameters: (i) RMSD and RMSF, (ii) radius of gyration- which indicated interaction of protein entities with ligand supported steadiness of the complex, (iii) Coulombic and Lennard–Jones interactions, which played a significant role in complex stability. DAPG showed a good number of H-bonds with PLpro and MM-PBSA binding energy when compared to the N protein. This study showed DAPG as a potential bioactive molecule to act as an inhibitor for the PLpro thereby, DAPG can be used as potential inhibitor against SARS-CoV-2 and is potential drug candidate against COVID-19.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Coronavirus Disease 2019 (COVID-19) pandemic disease is caused by the novel coronavirus SARS-CoV-2, a genetic homolog of the Severe Acute Respiratory Syndrome (SARS) coronavirus. The virus is transmitted from human-to-human contact through droplet nuclei during coughing or sneezing. The viral infection has an incubation period of around 5.2 days from the first symptom to 14 days with high incidence to death (Shang et al. 2020). The disease spread rapidly across the globe currently beyond 207 million confirmed infections despite record early approved vaccination drives (WHO 2021).
In the past, the zoonotic human Coronaviruses (CoVs) have caused not only SARS that spread across the globe with mortality rate of 15%, but also MERS-CoV (Middle East Respiratory Syndrome Coronavirus) which recorded a mortality rate of 35% (Chan et al. 2013). The CoVs are grouped into order Nidovirales; family- Coronavirideae and grouped into four genera: α-CoVs, β-CoVs, γ-CoVs, and ɗ-CoVs. The SARS-CoV-2 phylogenetically clusters to the β genus, equipped with organisational proteins: (i) spike (S), (ii) envelope (E), (iii) membrane (Mpro), (iv) nucleocapsid (N), (v) papain like protease (PLpro), (vi) RNA dependent RNA polymerase (RdRp), and (vii) RNA binding N terminal domain (NTD) which orchestrate COVID-19 pathogenesis and viral life cycle (Gil et al. 2020). These structural and functional proteins are recognized as suitable drugs against the virus, albeit the genes encoding these proteins are undergoing frequent mutations (Amamuddy et al. 2020) leading to formation of new variants (WHO 2021). During virus infection, spike protein determines the host cell specificity through its glycoprotein and receptor Angiotensin Converting Enzyme-2 (ACE-2) present on epithelial cells of lungs, kidneys, and heart cellular system (Zumla et al. 2016; Kulkarni et al. 2020).
The SARS-CoV-2 genome specifically encodes for two polyproteins, viz. Ppla (450 kD) and Pplb (750 kD), and both are processed with the help of chymotrypsin-fold proteinase (33 kD) to diverse useful proteins viz. (i) spike, (ii) membrane, (iii) envelop, (iv) nucleoprotein, (v) replicase, and (vi) polymerase. The proteinase, called main protease (Mpro) is the most suitable target for drug design (Luan et al. 2020; Bello 2020; Sepay et al. 2021). The Mpro is a dimer, comprising two protomers, containing three domains, domains I (residues 8 − 101) and II (residues 102 − 184) that contain β-barrels and domain III (residues 201− 306) which consists mainly of the α-helices. Drug discovery requires the dimeric state of SARS-CoV-2 Mpro for good results because monomer of Mpro is mostly inactive (Bello 2020; Mengist et al. 2021). Saquinavir and diastereomers of nelfinavir have been identified as a potent inhibitor of dimeric SARS-CoV-2 Mpro (Bello et al. 2020; Sargolzaei 2021).
Presently, various potential broad-spectrum antiviral drugs have been suggested for therapeutic and prophylactic purpose. However, few promising clinically approved beneficial drug/s, specific medicine, inhibitors (especially, targeting proteases), or treatment against SARS-CoV-2 is available. At present, only 8 candidate vaccines are approved, 11 for early or limited use, 127 vaccines are under various stages of human clinical trials and 99 preclinical candidates are under investigation. Despite the global herculean efforts, none of the vaccines warrants to secretary IgA antibody production or protection against the new variants. Several efforts to target structural and functional proteins to: (i) prevent the viral RNA synthesis, (ii) inhibit virus replication, (iii) block the virus binding to ACE-2, and (iv) arrest the virus’s self-assembly process have been reported. Three principal approaches for developing new drugs has emerged (Zumla et al. 2016), namely, (i) testing of known broad-spectrum antiviral compounds (Francés-Monerris et al. 2020), (ii) screening for various drug molecules with therapeutic potential against coronavirus using current molecular databases (Wang 2020), and (iii) advance new targeted drugs with the help of genomic data and pathological characteristics of diverse CoVs (Spreafico et al. 2020). These strategies may prove to be effective, but requires long time span, cost intensive and suffers from high failure rates. Hence, search for potential lead molecules from the marketed drugs appears as the most preferred alternative to manage SARS-CoV-2 infection.
Lonafarnib, tegobuvir, olysio, filibuvir, cepharanthine, theaflavin-3-3`-digallate, epigallocatechin gallate, epicatechingallate, gallocatechin-3-gallate polyphenols, hydroxychloroquine, chloroquine, remdesivir, thiopurine analogs, desacetylgedunin and several phenolic compounds act as protease inhibitors (Ruan et al. 2020; Shivanika et al. 2020; Ghosh et al. 2020; Baildya et al. 2020; Mirza et al. 2020; Baildya et al. 2021a, 2021b; Sabet et al. 2021). Various antiviral compounds (including FDA approved) as well as experimental drugs have been examined to protect against COVID-19 disease using in silico approach. For this, software tools viz. (i) molecular docking, (ii) molecular dynamic simulation, (iii) target point determination, and (iv) chemical stability appears first line of effective strategy for rapid screening in the emergency coronavirus pandemic situation. In particular, molecular docking rapidly identifies potential new drugs and evaluate preliminary efficacy (Grinter and Zou 2014). In short span, numerous in silico studies has recorded more than 900 antiviral drugs against protein targets involved in viral pathogenesis (mainly proteases), but targeted only a single viral protein entity (Joshi et al. 2020). Preferentially, search for robust inhibitors from natural lead compounds and, chemical inhibitors with capacity to target multiple proteins capable to arrest cross-infection appears promising choice as against single-entity viral targets (Baildya et al. 2021a, 2021b).
Among the natural compounds, 2,4 -diacetylphloroglucinol (DAPG) is a bioactive metabolite with low-molecular weight, non-volatile phenolic polyketide, with myriad environmental and clinical role. DAPG is reported to be produced by plants, algae, and especially, soil microbes (plant growth promoting rhizobacteria Pseudomonas spp.) making it amenable to industrial fermentative production. DAPG has been reported for its antagonistic activity against bacteria, fungi, helminths, protozoa, cancerous cells, and mycobacteria (Kankariya et al. 2019). Notably, DAPG has been reported to display strong activity against enveloped RNA and DNA viruses, such as (i) vesicular stomatitis virus (VSV), an enveloped RNA virus, and (ii) herpes simplex virus (HSV-I), an enveloped DNA virus, but unable to affect non-enveloped RNA virus (polio-I) (Tada et al. 1990).
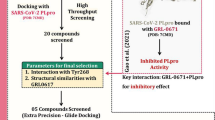
Biological production of DAPG is usually slow with very low yield, but DAPG can be chemically synthesized, a production route amenable for industrial scale-up (Zakrzewski et al. 2007). Thus, DAPG appears as bespoke bioentity with proven activity against enveloped viruses, fungal and bacterial pathogens which can be economically produced in purified form, tailor made to function, industrial-scale ready to make available in short span to control the COVID-19 pandemic as well as cross infection. Based on this premise, DAPG was selected for virtual screening against papain-like protease (PLpro), nucleocapsid (N), main protease (Mpro) and non-structural protein (nsp12) of SAR-CoV-2. The PDB: 3E9S – papain like protease has role in cleavage and maturation of viral polyproteins, assembly of the replicase-transcriptase complex, interruption of host responses, transmission and virulence of the virus (Osipiuk et al. 2021; Sabet et al. 2021). PDB: 4J3K and PDB: 1SSK- nucleocapsid (N) – RNA-binding protein dangerous for viral genome packaging and virus particle release (Zeng et al. 2020; Cubuk et al. 2021). PDB: 6LU7- main protease (Mpro) plays a critical role in viral replication and maturation (Joshi et al. 2020; Mengist et al. 2021). PDB: 6NUR- non-structural protein (nsp12) - essential subunit of RdRp (RNA-dependent RNA polymerase) and participate in RNA package and virus particle release (Ruan et al. 2020; Zeng et al. 2020). In the present study, Molecular docking study has been performed and higher binding energy was observed, which is being reported for the first time. Also we report the molecular dynamics simulation to assess the stability of proteins (of two ligand protein complexes which exhibited strong stability) with the DAPG ligand by estimating diverse parameters like SASA (solvent accessible surface area), RMSF (root mean square fluctuation), RMSD (root mean square deviation), and Rg (radius of gyration analysis). Prediction of Activity Spectra for Substances (PASS) and Bioactivity score analysis also depicted the drug-like nature of ligand and complexes. Thus further validation of DAPG and PLpro of SARS-CoV-2 through wet lab and clinical studies is warranted.
Materials and methods
Molecular docking analysis
AutoDock 4.2.6
The crystal structures of SARS-CoV-2 proteins were downloaded from RCSB Protein Data Bank (PDB) database as follows: (i) PDB ID: 3E9S (papain-like protease) (Ratia et al. 2008), (ii) PDB ID: 4J3K (nucleocapsid N protein) (Chen et al. 2013), (iii) PDB ID: 6LU7 (SARS-CoV-2 main protease) (Jin et al. 2020), (iv) PDB ID: 1SSK (nucleocapsid protein) (Huang et al. 2004), and (v) PDB ID: 6NUR (SARS-Coronavirus NSP12) (Kirchdoerfer and Ward 2019). The structure of the DAPG was retrieved from PubChem database. All the molecules from PDB and PubChem databases were converted to *.pdbqt format using Autodock tool and used for initial docking studies. The molecular docking technique examined for the folding pattern and molecular interactions between proteins and DAPG ligand molecule. Molecular docking was performed by AutoDock 4.2.6 program, using the implemented empirical free energy function and the Lamarckian Genetic Algorithm (LGA) for ligand tethering of proteins (Morris et al. 2009; Joshi et al. 2020; Shivanika et al. 2020). The interactions of complex protein-ligand conformation including hydrogen bonds and bond lengths were analyzed using Pymol software, UCSF Chimera, Molegro Molecular Viewer and Accelrys DS Visualizer software. Further validation through additional docking software was performed using AutoDock Vina.
AutoDock Vina
The docking of viral proteins with the DAPG was performed with the aid of AutoDock Vina, which is now widely used as a successor of AutoDock Tools (Ghosh et al. 2020). The binding affinities of DAPG–viral proteins were determined and analyzed using the same software. Execution of AutoDock Vina is faster than AutoDock 4.2.6, which reduces the size of the conformational space, allowing it to be searched reliably and reduces the computational effort in predictions of binding pockets. AutoDock Vina was used to perform docking simulations, generating conformations of ligand in complex with the receptor, which were finally ranked on the basis of binding energy (Siddiqui et al. 2020).
Molecular dynamics (MD) simulation study
MD simulation (100 ns) was achieved with the minimum energy conformer of two proteins and DAPG (PLpro-DAPG and N-DAPG) complex using Groningen Machine for Chemical Simulations (GROMACS) version 2020.2 (Krupanidhi et al. 2020). For the simulation, the protein topology was generated using the pdb2gmx tool of GROMACS. The ligand topology was generated using ANTECHAMBER from Amber Tools 16. The amber ff99SB-ILDN force field was chosen with the TIP3P water model. Amber ff99sb-ILDN (Lindorff-Larsen et al. 2010) is designed for protein, improved version by refitting the amino acid side chain torsion potentials of the AMBER99SB force field for four residues: isoleucine, leucine, aspartic acid, and asparagine (Aliev et al. 2014). Also, it has been used in several protein dynamics and found to give better results (Fan et al. 2021). The ligand topology generated using ANTECHAMBER from AmberTools 16 wherein we have used amber ff99sb charge model. The complex was placed in the box and was defined as a cubic unit cell with 1 nm. For PLpro-DAPG and N-DAPG complex, the system was soluted by adding 44,395 and 9655 water molecules and neutralized by adding 3 and 5 sodium ions, respectively. The energy minimization of each system was performed using the steepest descent method for force <10.0 kJ mol−1. Solvents and ions around the target proteins were equilibrated, maintaining temperature (300 K) and target pressure (1 bar). The produced trajectories were used to calculate root mean square deviation (RMSD), root mean square fluctuation (RMSF), radius of gyration (Rg), and hydrogen bond graphs by using numerous in-built scripts of GROMACS.
Binding free energy calculation
It was calculated by the Molecular Mechanics Poisson-Boltzmann Surface Area (MM-PBSA) method. For this purpose, binding free energy was calculated by the ‘g_mmpbsa’, a GROMACS tool and the tree-step calculation was performed to calculate interaction energy between enzyme and ligand (Kumari et al. 2014). The potential free energy in vacuum was calculated for polar solvation and non-polar solvation energy using SASA model. Python script MmPbSaStat.py, provided in the g_mmpbsa package, was used to calculate the binding energy components of protein-ligand binding. The binding free energy was calculated using,
where, △Gbind represents the binding free energy for complex (protein and ligand), Gcomplex, Gprotein and Gligand are the absolute free energy for complex, protein, and ligand, respectively.
Prediction of activity spectra for substances (PASS) analysis
It predicts biological activity spectrum of a compound under study based on its structure-activity relationship with a known chemical entity. PASS analysis was performed using various online tools as detailed viz. Lipinski’s rule of five. Lipinski’s rule of five is helpful in describing molecular properties of drug compounds required for estimation of important pharmacokinetic parameters (Siddiqui et al. 2020).
Bioactivity score prediction (BAS)
BAS values suggest a compound’s overall ability to be a potent drug candidate. Molinspiration is a web-based online tool which was used to predict the drug scores of the prospective DAPG with respect to several human receptors like GPCRs, ion channels, kinases, nuclear receptors, proteases and enzymes.
Pharmacokinetic (PK) parameters prediction
ADMET (A-Absorption, D-Distribution, M-Metabolism, E-Excretion and T-Toxicity) properties of DAPG were predicted using online Swiss ADME software. This software examines the essential pharmacokinetic properties of a compound like distribution viz. blood-brain barrier (BBB) and skin permeability (LogKp), and its metabolism in terms of it being a P glycoprotein (P-gp) substrate, Cytochrome P450 viz. CYP1A2, CYP2C19, CYP2C9, CYP2D6, CYP3A4 inhibitor as well as its lipophilicity for plasma membrane absorption (Siddiqui et al. 2020).
Results and discussion
Molecular docking
To study the binding interaction of the DAPG compound with the viral proteins (3E9S, 4J3K, 6LU7, 1SSK and 6NUR) molecular docking were performed with the help of AutoDock 4.2.6 program. Based on the binding energy of ligand-protein interactions with AutoDock 4.2.6 tool and ligand-protein complexes were selected for further validation through additional docking tool- AutoDock Vina. AutoDock 4.2.6 analyses revealed that the binding affinities of the DAPG with SARS-CoV-2 proteins which decreased in the order N (4J3K) > PLpro (3E9S) > Mpro (6LU7) > N (1SSK) > NSP12 (6NUR). Table 1 summarizes the binding energy parameters and interacting amino acid residues participating in binding pocket of proteins of SARS-CoV-2 with DAPG.
Results of molecular docking analysis using AutoDock 4.2.6 showed that DAPG has the best binding affinity with nucleocapsid (N) protein [−5.27 Kcal mol−1] followed by papain like protease (3E9S) [−5.20 Kcal mol−1]. On the other hand, DAPG exhibited the best binding affinity with papain like protease (3E9S) and main protease (6LU7) [−6.20; −5.97 Kcal mol−1 respectively] as analyzed by AutoDock Vina. Intrestingly, this study is in agreement with a previously published report where phytoconstituents from miracle herb Nigella sativa which showed similar kind of variation in binding energy with their selected protein targets (Siddiqui et al. 2020). Results propose that slight dissimilarities in binding affinity of the DAPG is because of the variances in the generation of grid boxes and determination of binding pockets on the target proteins by these software on account of slight differences in selection criteria. This has led to a difference in interacting amino acid residues in the binding pockets of proteins of SARS-CoV-2 as is evident from Table 1.
After exploring the molecular interactions, it was perceived that the bioactive molecule DAPG forms various interaction as in case of: (i) papain-like protease (3E9S), manifests hydrogen bonds formed with amino acid residues Glu168, Lys158, Leu163, Gln270, Asp165, and one Pi-Pi T shaped interaction with Gly164. DAPG also formed van der Waals (VdW) interactions with residues Tyr269, Tyr274 and Tyr265; (ii) the Nucleocapsid protein (4J3K) revealed hydrogen bonds with amino acid residues Val80, Ala77, Val74, and few residues formed other interactions like Ala85 (Pi-sigma), Ser138 (C-H), and Ala82 (Pi-alkyl), while residues Pro81, Pro75, Pro165, Ile76 and Tyr115 showed VdW interactions; (iii) the main protease (6 LU7) formed hydrogen bond with amino acid residues Glu166, His164, Arg188, Met165, His41 and other interaction with Met (Pi-alkyl) along with VdW interactions formed with residues Leu167, Gln192, Val186, Gln189, Thr190, Met49 and Asp187; (iv) Nucleocapsid protein (1SSK) residues, the Trp30, Phe88, Gly125, and Thr27 formed hydrogen bonds, while several residues such as Ser29, Phe31,Tyr87, Tyr89, Arg66 formed VdW interactions; (v) the NSP12 (6NUR), residues viz. Asp517, Gln524, Tyr521, Val495 formed hydrogen bonds, Leu498 formed Pi-alkyl interaction and VdW interactions formed with Ile494, Val493, Asn491, Tyr516, Asn496 stabilizing the binding of the ligand to the enzyme. Cumulatively, the hydrogen bonding interaction of DAPG ligand with viral proteins was found highly stable, while the prevalence of many hydrogen bonds indicated effective inhibition and DAPG can fit inside the pocket present in the viral protein’s domain (Fig. 1 and Fig. S1). The docking results obtained from AutoDock 4.2.6 and AutoDock Vina tools were visualized in Discovery Studio showing prominent interactions between various amino acid residues (Table 1) and Fig. S2 displayed the best docking poses of DAPG with proteins of SARS-CoV-2.
The results are in accordance with Udrea et al. 2020; Shivanika et al. 2020 and Sabet et al. 2021 who proposed that natural antiviral phytocompounds, natural compounds and phenolic acids could act as a lead for the COVID-19 treatment by using AutoDock 4.2.6. Thioridazine and sulphoridazine compounds are the class of Phenothiazines which displayed binding energy of −7.06 kcal mol−1 and − 8.72 kcal mol−1 (Udrea et al. 2020). Eight green tea polyphenols exhibited good binding affinity −7.1 to −9.0 kcal mol−1 (Ghosh et al. 2020). Also, Uercetin, Kaempherol, Nelfinavir and Lopinavir has demonstrated binding energy of −6.6, −6.4, −7.0 and − 6.4 kcal mol−1, respectively towards Mpro (Huynh et al. 2020). Similarly, the compounds ADM_13083841, LMG_15521745 and SYN_15517940 with binding energy score − 8.9, −8.7, −8.7 kcal mol−1, respectively along with 26 FDA-approved drugs revealed a docking score of −7 or higher (Alamri et al. 2020). On a similar streak, anti-malarial drugs, Chloroquine and Hydroxychloroquine, was recognized as an inhibitor towards SARS-CoV-2 PLpro (Baildya et al. 2020, 2021a, 2021b; Hussein and Elkhair 2021). Negative binding affinity indicated stronger protein-ligand complex (Kodchakorn et al. 2020). Similarly, Nilotinib, Saquinavir, Tipranavir, Lonafarnib, Tegobuvir, Olysio, Filibuvir, and Cepharanthine drugs are reported for the inhibition of NSP12 protein with binding energy of −7.7 to −8.6 kcal mol−1 (Ruan et al. 2020) by using AutoDock Vina. Interestingly, DAPG showed strong binding with all selected SARS-CoV-2 proteins and is previously reported as an antiviral compound for enveloped viruses (Tada et al. 1990), hence, it may be a potent inhibitor of the targeted key viral proteins used in this study.
Molecular dynamics (MD) simulation study
During the 100 ns MD simulations, in PLpro-DAPG complex, the RMSD values obtained between the ranges of 0 to 0.6 nm (Fig. 2), indicating that the protein fold fluctuated during the initial period of the simulation. After 40 ns, the structure started to find an equilibrated position, and just before 50 ns, the complex became dynamically stable. After 50 ns, the protein complex remained stable with 0.1 rms value, indicating that the complex established stability at 50 ns and remained stable for the rest of the span, i.e., the protein interacted with ligand quite well. In case of N-DAPG complex, the RMSD values remained between the ranges of 0 to 0.5 nm, which indicated that the protein complex fold was steady throughout the simulation. After 30 ns, the structure started to find an equilibrated position, and just before 50 ns, the complex became dynamically stable thereafter.
Thus, both PLpro-DAPG and N-DAPG complex remained stable with the average RMSD of 0.1 rms, suggesting that both the complexes established stability at 50 ns and 30 ns, respectively. Also, the complexes remained stable for the rest of span demonstrating faithful protein interaction with DAPG ligand. Although the RMSD of compound fluctuated in the initial period of the run, the complexes stabilized rapidly thereby, displayed negligible RMSD deviation. After 40 ns and 50 ns, for both PLpro-DAPG and N-DAPG complex changes in ligand was observed to be ˂0.1 nm indicating transformation of binding orientation over the simulation time. The results indicated that both the complex exhibited RMSD of <0.6 nm.
Further, to measure the fluctuation measure of atoms or residue in simulation, RMSF calculation on residue level was performed. The PLpro-DAPG complex residues at 50, 107, 192, 227, 228 and 281 showed main fluctuations but other residues showed variation in rms value of about 0.3 nm. In case of N-DAPG complex, residues at 70, 95, 98, 100, 101, 106, 123 and 158 showed minor fluctuations, between 95 to 107 showed major fluctuations, while other residues showed variation in rms value of <0.3 nm (Fig. 3).
During MD simulation, ligand steadiness and fluctuation were supported by several hydrogen bonds. In this study, both PLpro-DAPG and N-DAPG complex showed maximum of 4 to 5 hydrogen bonds formed during the simulation along with pairs within 0.35 nm (3.5 Å) which signifies the number of contacts formed within the cut-off value in between protein and ligand. It showed that a maximum of 13 and 11 contacts were formed by DAPG during simulation in PLpro-DAPG and N-DAPG complex, respectively. Most of the frames were found to have 3 to 6 contacts (Fig. 4), suggesting similar conformational stability and such network possibly played a significant role in strengthening the binding effect throughout the simulation period ultimately supporting the docking. Poor stability of ligand-binding is often identified from fewer number of hydrogen bonds, while flexibility alters protein shape, structure, and function. Proteins generally require significant degree of flexibility at physiological conditions to carry out native function. Under these circumstances, DAPG can certainly alter the flexibility of protein and decrease the enzymatic activity.
The radius of gyration (Rg) of a protein complex is a measure of its compactness and stably folded protein likely maintains a relatively steady value of Rg. However, Rg changes over time when protein unfolds (Alamri et al. 2020). In the present MD simulation run, Rg showed some fluctuations in the initial phase, which corroborated to the behaviour observed in the RMSD estimations for both complex, and stabilised under 2.4 nm and 1.5 nm, respectively. The Fig. 5 demonstrates the steadiness of PLpro-DAPG complex after 30 ns and N-DAPG complex after 50 ns.
Two types of short-range interaction energies were calculated: (i) LJ-SR (Lennard–Jones short-range), and (ii) Coul-SR (Coulombic short-range). The average Coul-SR interaction energy for PLpro-DAPG and N-DAPG complex showed to be −21.87 ± 1.9 and −18.26 ± 1.7 kJ mol−1, respectively [Fig. 6a], while the LJ-SR interaction energy was −125.12 ± 3.1 and −87.11 ± 4.4 kJ mol−1, respectively [Fig. 6b]. Considering the result of 100 ns simulation, contribution of Coul-SR potential in both complexes was greater than LJ-SR potential. The amount of LJ-SR and Coul-SR potential generally contribute to the interaction energy estimation.
(a) Coul-SR (Columbic short-range electrostatic) Interaction energy pattern between protein and ligand (i) PLpro-DAPG, and (ii) N-DAPG complex; and (b) LJ-SR (Lennard-Jones short-range electrostatic) Interaction energy pattern between protein and ligand, (iii) PLpro-DAPG, and (iv) N-DAPG complex estimated using GROMACS
Binding free energy calculation
To understand the interaction between proteins and DAPG moiety binding energy calculations by MM-PBSA method was performed (Table 2) and binding energies followed the order of PLpro > N. There were strong van der Waal and electrostatic interactions between PLpro and DAPG compared to N and DAPG.
MM-PBSA methodology has developed a popular method for estimated the free energy of a complex (Kollman et al. 2000; Kuhn et al. 2005). The DAPG ligand and protein interactions shown in the form of binding energy was considerably good because lesser the binding energy better is the protein-ligand interaction (Joshi et al. 2020). An aggregate sum of all energy estimates i.e., VdW energy, Electrostatic energy, Polar solvation energy and, SASA energy was calculated. Relatively, MM-PBSA study showed that PLpro-DAPG complex showed a least binding free energy of −96.65 kJ mol−1 and hydrogen bond formation supporting results of docking, hence, it can be claimed that DAPG can inhibit the viral protein PLpro, the Papain like protease. Similarly, N-DAPG complex showed slightly greater binding free energy of −62.58 kJ mol−1. Thus, the study demonstrates a promising stability of DAPG with the viral papain like protease (PLpro) over the simulated MD time of 100 ns. The entropy change upon binding was found to be negligible, and hence, it can be ignored if relative binding free energy is considered (Wichapong et al. 2016). In the recent past, it was found that naphthalene-based inhibitors have inhibitory effects towards PLpro of SARS-CoV-2, wherein MM-PBSA binding energy from simulations of the four different starting poses (A–D) was between −12.3 to −17.7 kcal mol−1. On this streak, a compound, Z93 revealed binding energy of −6.66 kcal mol−1 determined through MM-PBSA, the first chemical compound demonstrated against SARS-CoV-2 PLpro (Mirza et al. 2020). Similarly, Phenformin, Quercetin, and Ritonavir showed binding energy values of −56.6, −40.9, and −37.6 kcal mol−1 as calculated by MM-GBSA (Molecular Mechanics/Generalized Born Surface Area), while compounds ADM_13083841, LMG_15521745 and SYN_15517940 showed binding energy of 11.89, −10.84, and −12.96 kcal mol−1, respectively from the MM-PBSA which demonstrated specific inhibition towards PLpro (Alamri et al. 2020; Kandeel et al. 2020). Recently, Pitsillou et al. 2021 found that Hypericin, Rutin and Cyanidin-3-O-glucoside shows concentration-dependent inhibition of the PLpro in silico as well as in vitro enzymatic inhibition assay. The natural compound, desacetylgedunin extracted from Azadirachta indica (neem) was shown to have highest docking score with PLpro and induce a large structural change as evaluated by MD simulation (Baildya et al. 2021a, 2021b).
PASS analysis of DAPG using Lipinski’s rule of five
Lipinski’s rule of five evaluates drug likeliness of a compound having properties that would make it potential drug in humans. Table 3 shows the PASS analysis of DAPG in terms of their physicochemical properties by applying Lipinski’s rule of five. The oral activity of a drug compound is predicted by calculating the parameters like partition coefficient, polar surface area, number of hydrogen bond donors, number of hydrogen bond acceptors, and molecular weight. An orally active drug compound has no more than one violation (Khan et al. 2017). When compared with hydroxychloroquin, DAPG also showed Lipinski’s violation 0 (Siddiqui et al. 2020).
Bioactivity score (BAS) prediction
The predicted BAS of DAPG summarized in Table 4. Normally, a molecule having BAS >0.00 is most likely to possess considerable biological activities, while compounds having values between −0.50 and 0.00 are supposed to be moderately active and compounds having BAS < -0.50, are expected to be inactive. The results of the present study demonstrated that DAPG is biologically active molecule and capable of producing the physiological actions by multiple mechanisms after interacting with G Protein-Coupled Receptors (GPCRs) ligands, nuclear receptor ligands or by acting as inhibitors of proteases and other enzymes. DAPG displayed considerable activity as protease inhibitor as evident from their BAS score.
Pharmacokinetic (PK) parameters prediction
To assess the pharmacokinetic feasibility of DAPG as prospective drug candidate, its ADMET properties viz. absorption, distribution, metabolism, excretion and toxicity were calculated using online SwissADME software (Table 5).
Based on the calculated Log P value, DAPG found to be lipid soluble. Intriguingly, DAPG neither displayed blood-brain barrier (BBB) permeability nor predicted to act as permeability-glycoprotein (P-gp) substrate. The CYPs (Cytochromes P450) are involved in the biotransformation of xenobiotics. Compounds that inhibit the five classes of CYPs viz. CYP3A4, CYP1A2, CYP2C9, CYP2C19 and CYP2D6 would cause an increase in their plasma concentrations, thus contributing to improved bioavailability. In the present study, DAPG was not found to act as inhibitor of the class of CYPs. Kp is skin permeability, widely used to quantitatively describe the rate of chemical permeation through the outermost layer of the skin. DAPG showed negative Kp value which indicates less possibility of topical absorption of DAPG.
Overall, the concerted approach of molecular docking, MD simulation and interaction energy estimation have been proved to be useful for design and development of potent viral protein inhibitors. Mostly, inhibition of papain-like protease (PLpro) can potentially block continuation of the viral life cycle, viral replication and processing viral polyproteins during maturation. Hence, PLpro signifies an important antiviral target protease using DAPG as an inhibitory compound.
Consequently, this study primarily draws comprehensive merit from the natural bioactive compound DAPG as protease inhibitor to help design more effective inhibitor for the SARS-CoV-2 infection. Remarkably, DAPG has previously shown anti-bacterial, anti-fungal, anti-viral, anti-cancer, anti-helminthic and anti-protozoal activity as a natural antibiotic and therapeutic agent and hence, DAPG deserves to be designated as a multi-functional natural metabolite.
Further, the chemical derivation of DAPG functional groups is another potential area to extend its potency against the viral proteins using organic approaches. The derivatized-DAPG and viral protein interaction could be further enhanced, evaluated, and can help select even better synthetic lead drug targets, thereby help to design even more potent viral protein inhibitors, suggesting a pandora box for exploring even more effective antiviral drugs from natural sources with negligible side effects.
Conclusion
The present study computationally investigated DAPG compound for prediction of their potential inhibitory activities against papain-like protease (PLpro), nucleocapsid (N), main protease (Mpro) and non-structural protein (nsp12) of SARS-CoV-2. Analysis of favourable docked conformations of DAPG compound revealed the highest binding affinity with PLpro and N protein. MD simulation and evaluation of parameters like RMSD, RMSF and residue analysis of protein revealed that DAPG has a profound effect on the conformational alternation with these two proteins with noticeable highlighting effect especially on PLpro. This is probably the first report which explores the binding potential of DAPG against SARS-CoV-2 and in a multi-target approach. The interaction of DAPG could be further enhanced via functional group derivatizations and designing of novel compounds. Finally, DAPG could serve as potential lead molecule for multi-targeted drug development regime and future clinical studies.
Abbreviations
- COVID-19:
-
Coronavirus Disease 2019
- SARS:
-
Severe Acute Respiratory Syndrome
- CoVs:
-
Coronaviruses
- SARS-CoV-2:
-
Severe Acute Respiratory Syndrome Coronavirus-2
- PLpro:
-
papain-like protease
- N:
-
nucleocapsid
- Mpro:
-
main protease
- nsp12:
-
non-structural protein
- DAPG:
-
2, 4-diacetylphloroglucinol
- RMSD:
-
root mean square deviation
- RMSF:
-
root mean square fluctuation
- Rg:
-
radius of gyration
- MM-PBSA:
-
Molecular Mechanics Poisson-Boltzmann Surface Area
- ACE-2:
-
Angiotensin Converting Enzyme-2
- MD:
-
Molecular dynamics
- GROMACS:
-
Groningen Machine for Chemical Simulations
- LJ-SR:
-
Lennard–Jones short-range
- Coul-SR:
-
Coulombic short-range
- SASA:
-
solvent accessible surface area
- GPCR:
-
G Protein-Coupled Receptors
References
Alamri MA, ul Qamar MT, Mirza MU, Alqahtani SM, Froeyen M, Chen LL (2020) Discovery of human coronaviruses pan-papain-like protease inhibitors using computational approaches. J Pharm Anal, 10(6), 546-559. doi: https://doi.org/10.1016/j.jpha.2020.08.012
Aliev AE, Kulke M, Khaneja HS, Chudasama V, Sheppard TD, Lanigan RM (2014) Motional timescale predictions by molecular dynamics simulations: case study using proline and hydroxyproline sidechain dynamics. Proteins Struct Funct Bioinf 82(2):195–215. https://doi.org/10.1002/prot.24350
Baildya N, Ghosh NN, Chattopadhyay AP (2020) Inhibitory activity of hydroxychloroquine on COVID-19 main protease: an insight from MD-simulation studies. J Mol Struct 1219:128595. https://doi.org/10.1016/j.molstruc.2020.128595
Baildya N, Ghosh NN, Chattopadhyay AP (2021a) Inhibitory capacity of chloroquine against SARS-COV-2 by effective binding with angiotensin converting enzyme-2 receptor: an insight from molecular docking and MD-simulation studies. J Mol Struct 1230:129891. https://doi.org/10.1016/j.molstruc.2021.129891
Baildya N, Khan AA, Ghosh NN, Dutta T, Chattopadhyay AP (2021b) Screening of potential drug from Azadirachta Indica (Neem) extracts for SARS-CoV-2: an insight from molecular docking and MD-simulation studies. J Mol Struct 1227:129390. https://doi.org/10.1016/j.molstruc.2020.129390
Bello M (2020) Prediction of potential inhibitors of the dimeric SARS-CoV2 main proteinase through the MM/GBSA approach. J Mol Graph Model 101:107762. https://doi.org/10.1016/j.jmgm.2020.107762
Bello M, Martínez-Muñoz A, Balbuena-Rebolledo I (2020) Identification of saquinavir as a potent inhibitor of dimeric SARS-CoV2 main protease through MM/GBSA. J Mol Model 26(12):1–11. https://doi.org/10.1007/s00894-020-04600-4
Chan JF, Chan KH, Kao RY, To KK, Zheng BJ, Li CP, Li PT, Dai J, Mok FK, Chen H, Hayden FG, Yuen KY (2013) Broad-spectrum antivirals for the emerging Middle East respiratory syndrome coronavirus. J Inf Secur 67(6):606–616. https://doi.org/10.1016/j.jinf.2013.09.029
Chen IJ, Yuann JMP, Chang YM, Lin SY, Zhao J, Perlman S, Shen YY, Huang TH, Hou MH (2013) Crystal structure-based exploration of the important role of Arg106 in the RNA-binding domain of human coronavirus OC43 nucleocapsid protein. Biochim Biophys Acta, Proteins Proteomics 1834(6):1054–1062. https://doi.org/10.1016/j.bbapap.2013.03.003
Cubuk J, Alston JJ, Incicco JJ, Singh S, Stuchell-Brereton MD, Ward MD, Zimmerman MI, Vithani N, Griffith D, Wagoner JA, Bowman GR, Hall KB, Soranno A, Holehouse AS (2021) The SARS-CoV-2 nucleocapsid protein is dynamic, disordered, and phase separates with RNA. Nat Commun 12(1):1–17. https://doi.org/10.1038/s41467-021-21953-3
Fan W, Mar KB, Sari L, Gaszek IK, Cheng Q, Evers BM, Shelton JM, Carter MW, Siegwart DJ, Lin MM, Schoggins JW (2021) TRIM7 inhibits enterovirus replication and promotes emergence of a viral variant with increased pathogenicity. Cell. https://doi.org/10.1016/j.cell.2021.04.047
Francés-Monerris A, Hognon C, Miclot T, García-Iriepa C, Iriepa I, Terenzi A, Grandemange S, Barone G, Marazzi M, Monari A (2020) Molecular basis of SARS-CoV-2 infection and rational design of potential antiviral agents: modeling and simulation approaches. J Proteome Res 19(11):4291–4315. https://doi.org/10.1021/acs.jproteome.0c00779
Ghosh R, Chakraborty A, Biswas A, Chowdhuri S (2020) Evaluation of green tea polyphenols as novel corona virus (SARS CoV-2) main protease (Mpro) inhibitors–an in silico docking and molecular dynamics simulation study. J Biomol Struct Dyn:1–13. https://doi.org/10.1080/07391102.2020.1779818
Gil C, Ginex T, Maestro I, Nozal V, Barrado-Gil L, Cuesta-Geijo MÁ, Urquiza J, Ramírez D, Alonso C, Campillo NE, Martinez A (2020) COVID-19: drug targets and potential treatments. J Med Chem 63(21):12359–12386. https://doi.org/10.1021/acs.jmedchem.0c00606
Grinter SZ, Zou X (2014) Challenges, applications, and recent advances of protein-ligand docking in structure-based drug design. Molecules 19(7):10150–10176. https://doi.org/10.3390/molecules190710150
Huang Q, Yu L, Petros AM, Gunasekera A, Liu Z, Xu N, Hajduk P, Mack J, Fesik SW, Olejniczak ET (2004) Structure of the N-terminal RNA-binding domain of the SARS CoV nucleocapsid protein. Biochemistry 43(20):6059–6063. https://doi.org/10.1021/bi036155b
Hussein RK, Elkhair HM (2021) Molecular docking identification for the efficacy of some zinc complexes with chloroquine and hydroxychloroquine against main protease of COVID-19. J Mol Struct 1231:129979. https://doi.org/10.1016/j.molstruc.2021.129979
Huynh T, Wang H, Luan B (2020) In silico exploration of the molecular mechanism of clinically oriented drugs for possibly inhibiting SARS-CoV-2’s main protease. J Phys Chem Lett 11(11):4413–4420. https://doi.org/10.1021/acs.jpclett.0c00994
Jin Z, Du X, Xu Y, Deng Y, Liu M, Zhao Y, Zhang B, Li X, Zhang L, Peng C, Duan Y, Yu J, Wang L, Yang K, Liu F, Jiang R, Yang X, You T, Liu X et al (2020) Structure of M pro from SARS-CoV-2 and discovery of its inhibitors. Nature 582(7811):289–293. https://doi.org/10.1038/s41586-020-2223
Joshi RS, Jagdale SS, Bansode SB, Shankar SS, Tellis MB, Pandya VK, Chugh A, Giri AP, Kulkarni MJ (2020) Discovery of potential multi-target-directed ligands by targeting host-specific SARS-CoV-2 structurally conserved main protease. J Biomol Struct Dyn:1–16. https://doi.org/10.1080/07391102.2020.1760137
Kandeel M, Abdelrahman AH, Oh-Hashi K, Ibrahim A, Venugopala KN, Morsy MA, Ibrahim MA (2020) Repurposing of FDA-approved antivirals, antibiotics, anthelmintics, antioxidants, and cell protectives against SARS-CoV-2 papain-like protease. J Biomol Struct Dyn:1–8. https://doi.org/10.1080/07391102.2020.1784291
Kankariya RA, Chaudhari AB, Gavit PM, Dandi ND (2019) 2, 4-Diacetylphloroglucinol: a novel biotech bioactive compound for agriculture. In: Gupta VK, Prabha R (eds) Singh, DP. Microbial Interventions in Agriculture and Environment Springer, Singapore, pp 419–452. https://doi.org/10.1007/978-981-13-8391-5_16
Khan T, Dixit S, Ahmad R, Raza S, Azad I, Joshi S, Khan AR (2017) Molecular docking, PASS analysis, bioactivity score prediction, synthesis, characterization and biological activity evaluation of a functionalized 2-butanone thiosemicarbazone ligand and its complexes. J Chem Biol, 10(3), 91-104. http://DOI https://doi.org/10.1007/s12154-017-0167-y
Kirchdoerfer RN, Ward AB (2019) Structure of the SARS-CoV nsp12 polymerase bound to nsp7 and nsp8 co-factors. Nat Commun 10(1):1–9. https://doi.org/10.1038/s41467-019-10280-3
Kodchakorn K, Poovorawan Y, Suwannakarn K, Kongtawelert P (2020) Molecular modelling investigation for drugs and nutraceuticals against protease of SARS-CoV-2. J Mol Graph Model 101:107717. https://doi.org/10.1016/j.jmgm.2020.107717
Kollman PA, Massova I, Reyes C, Kuhn B, Huo S, Chong L, Lee M, Lee T, Duan Y, Wang W, Donini O, Cieplak P, Srinivasan J, Case D, Cheatham TE (2000) Calculating structures and free energies of complex molecules: combining molecular mechanics and continuum models. Acc Chem Res 33(12):889–897. https://doi.org/10.1021/ar000033j
Krupanidhi S, Peele KA, Venkateswarulu TC, Ayyagari VS, Bobby MN, Babu JD, Narayana V, Aishwarya G (2020) Screening of phytochemical compounds of Tinospora cordifolia for their inhibitory activity on SARS-CoV-2: an in-silico study. J Biomol Struct Dyn:1–5. https://doi.org/10.1080/07391102.2020.1787226
Kuhn B, Gerber P, Schulz-Gasch T, Stahl M (2005) Validation and use of the MM-PBSA approach for drug discovery. J Med Chem 48(12):4040–4048. https://doi.org/10.1021/jm049081q
Kulkarni SA, Nagarajan SK, Ramesh V, Palaniyandi V, Selvam SP, Madhavan T (2020) Computational evaluation of major components from plant essential oils as potent inhibitors of SARS-CoV-2 spike protein. J Mol Struct 1221:128823. https://doi.org/10.1016/j.molstruc.2020.128823
Kumari R, Kumar R, Consortium OSDD, Lynn A (2014) g_mmpbsa• a GROMACS tool for high-throughput MM-PBSA calculations. J Chem Inf Model 54(7):1951–1962. https://doi.org/10.1021/ci500020m
Lindorff-arsen K, Piana S, Palmo K, Maragakis P, Klepeis JL, Dror RO, Shaw DE (2010) Improved side-chain torsion potentials for the Amber ff99SB protein force field. Proteins: Struct Funct Bioinf 78(8):1950–1958. https://doi.org/10.1002/prot.22711
Luan B, Huynh T, Cheng X, Lan G, Wang HR (2020) Targeting proteases for treating COVID-19. J Proteome Res 19(11):4316–4326. https://doi.org/10.1021/acs.jproteome.0c00430
Mengist HM, Dilnessa T, Jin T (2021) Structural basis of potential inhibitors targeting SARS-CoV-2 main protease. Front Chem 9. https://doi.org/10.3389/fchem.2021.622898
Mirza MU, Ahmad S, Abdullah I, Froeyen M (2020) Identification of novel human USP2 inhibitor and its putative role in treatment of COVID-19 by inhibiting SARS-CoV-2 papain-like (PLpro) protease. Comput Biol Chem 89:107376. https://doi.org/10.1016/j.compbiolchem.2020.107376
Morris GM, Huey R, Lindstrom W, Sanner MF, Belew RK, Goodsell DS, Olson AJ (2009) AutoDock4 and AutoDockTools4: automated docking with selective receptor flexibility. J Comput Chem 30(16):2785–2791. https://doi.org/10.1002/jcc.21256
Osipiuk J, Azizi SA, Dvorkin S, Endres M, Jedrzejczak R, Jones KA, Kang S, Kathayat RS, Kim Y, Lisnyak GV, Maki SL, Nicolaescu V, Taylor CA, Tesar C, Zhang UA, Zhou Z, Randall G, Michalska K, Snyder SA et al (2021) Structure of papain-like protease from SARS-CoV-2 and its complexes with non-covalent inhibitors. Nat Commun 12(1):1–9. https://doi.org/10.1038/s41467-021-21060-3
Pitsillou E, Liang J, Ververis K, Hung A, Karagiannis TC (2021) Interaction of small molecules with the SARS-CoV-2 papain-like protease: in silico studies and in vitro validation of protease activity inhibition using an enzymatic inhibition assay. J Mol Graph Model 104:107851. https://doi.org/10.1016/j.jmgm.2021.107851
Ratia K, Pegan S, Takayama J, Sleeman K, Coughlin M, Baliji S, Chaudhuri R, Fu W, Prabhakar BS, Johnson ME, Baker SC, Ghosh AK, Mesecar AD (2008) A noncovalent class of papain-like protease/deubiquitinase inhibitors blocks SARS virus replication. Proc Natl Acad Sci U S A 105(42):16119–16124. https://doi.org/10.2210/pdb3e9s/pdb
Ruan Z, Liu C, Guo Y, He Z, Huang X, Jia X, Yang T (2020) SARS-CoV-2 and SARS-CoV: virtual screening of potential inhibitors targeting RNA-dependent RNA polymerase activity (NSP12). J Med Virol 93(1):389–400. https://doi.org/10.1002/jmv.26222
Sabet R, Sisakht M, Emami L, Sabahi Z (2021) Comparison of COVID-19 virus main protease inhibition activities of phenolic acids by molecular docking. Trends Pharmacol Sci 7(2):117–126. https://doi.org/10.30476/tips.2021.90386.1083
Sargolzaei M (2021) Effect of nelfinavir stereoisomers on coronavirus main protease: molecular docking, molecular dynamics simulation and MM/GBSA study. J Mol Graph Model 103:107803. https://doi.org/10.1016/j.jmgm.2020.107803
Sepay N, Sekar A, Halder UC, Alarifi A, Afzal M (2021) Anti-COVID-19 terpenoid from marine sources: a docking, admet and molecular dynamics study. J Mol Struct 1228:129433. https://doi.org/10.1016/j.molstruc.2020.129433
Shang Z, Chan SY, Liu WJ, Li P, Huang W (2020) Recent insights into emerging coronavirus: SARS-CoV-2. ACS Infect Dis. https://doi.org/10.1021/acsinfecdis.0c00646
Sheik Amamuddy O, Verkhivker GM, Tastan Bishop O (2020) Impact of early pandemic stage mutations on molecular dynamics of SARS-CoV-2 Mpro. J Chem Inf Model 60(10):5080–5102. https://doi.org/10.1021/acs.jcim.0c00634
Shin D, Mukherjee R, Grewe D, Bojkova D, Baek K, Bhattacharya A, Schulz L, Widera M, Mehdipour AR, Tascher G, Geurink PP, Wilhelm A, van der Heden van Noort GJ, Ovaa H, Müller S, Knobeloch KP, Rajalingam K, Schulman BA, Cinatl J et al (2020) Papain-like protease regulates SARS-CoV-2 viral spread and innate immunity. Nature 587(7835):657–662. https://doi.org/10.1038/s41586-020-2601-5
Shivanika C, Kumar D, Ragunathan V, Tiwari P, Sumitha A (2020) Molecular docking, validation, dynamics simulations, and pharmacokinetic prediction of natural compounds against the SARS-CoV-2 main-protease. J Biomol Struct Dyn 1. https://doi.org/10.1080/07391102.2020.1815584
Siddiqui S, Upadhyay S, Ahmad R, Gupta A, Srivastava A, Trivedi A, Husainb I, Ahmadc B, Ahamed MKhan MA (2020) Virtual screening of phytoconstituents from miracle herb nigella sativa targeting nucleocapsid protein and papain-like protease of SARS-CoV-2 for COVID-19 treatment. J Biomol Struct Dyn:1–21. https://doi.org/10.1080/07391102.2020.1852117
Spreafico R, Soriaga LB, Grosse J, Virgin HW, Telenti A (2020) Advances in genomics for drug development. Genes 11(8):942. https://doi.org/10.3390/genes11080942
Tada M, Takakuwa T, Nagai M, Yoshii T (1990) Antiviral and antimicrobial activity of 2, 4-Diacylphloroglucinols, 2-Acylcyclohexane-1, 3-diones and 2-Carboxamidocyclo-hexane-1, 3-diones. Agr Biol Chem 54(11):3061–3063. https://doi.org/10.1080/00021369.1990.10870402
Udrea AM, Avram S, Nistorescu S, Pascu ML, Romanitan MO (2020) Laser irradiated phenothiazines: new potential treatment for COVID-19 explored by molecular docking. J Photochem Photobiol B Biol 211:111997. https://doi.org/10.1016/j.jphotobiol.2020.111997
Wang J (2020) Fast identification of possible drug treatment of coronavirus disease-19 (COVID-19) through computational drug repurposing study. J Chem Inf Model 60(6):3277–3286. https://doi.org/10.1021/acs.jcim.0c00179
Wang M, Cao R, Zhang L, Yang X, Liu J, Xu M, Shi Z, Hu Z, Zhong W, Xiao G (2020) Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res 30(3):269–271. https://doi.org/10.1038/s41422-020-0282-0
Wichapong K, Alard JE, Ortega-Gomez A, Weber C, Hackeng TM, Soehnlein O, Nicolaes GA (2016) Structure-based design of peptidic inhibitors of the interaction between CC chemokine ligand 5 (CCL5) and human neutrophil peptides 1 (HNP1). J Med Chem 59(9):4289–4301. https://doi.org/10.1021/acs.jmedchem.5b01952
World Health Organization (2021) WHO coronavirus disease (COVID-19) dashboard https://covid19. who. int. Accessed 17 August, 2021
Zakrzewski J, Karpińska M, Maliński Z (2007) A large scale synthesis of a natural antibiotic, 2, 4-diacetylophloroglucinol (DAPG). Arch Pharm 340(2):103–106. https://doi.org/10.1002/ardp.200600139
Zeng W, Liu G, Ma H, Zhao D, Yang Y, Liu M, Mohammed A, Zhao C, Yang Y, Xie J, Ding C, Ma X, Weng J, Gao Y, He H, Jin T (2020) Biochemical characterization of SARS-CoV-2 nucleocapsid protein. Biochem Biophys Res Commun 527(3):618–623. https://doi.org/10.1016/j.bbrc.2020.04.136
Zumla A, Chan JF, Azhar EI, Hui DS, Yuen KY (2016) Coronaviruses—drug discovery and therapeutic options. Nat Rev Drug Discov 15(5):327–347. https://doi.org/10.1038/nrd.2015.37
Acknowledgments
Dr. Vaibhav Sable, MrBiologist, Pune for significant support in bioinformatic analysis and valuable suggestions.
Author information
Authors and Affiliations
Contributions
Raksha A. Kankariya: Conceptualization, Methodology, Software, Writing- Original draft preparation; Ambalal B. Chaudhari: Supervision, Writing- Reviewing and Editing; Navin D. Dandi: Conceptualization, Methodology, Software, Supervision, Writing- Original draft preparation, Writing- Reviewing and Editing; Funding acquisition: This work was supported by Department of Science and Technology, New Delhi, India [SR/WOS-A/LS-1209/2014(G); WoS-A scheme to RAK] and [SR/FST/LS-II/2017/101(C); DST-FIST]; University Grants Commission, New Delhi, India [F.4-23/2015/DRS-III (SAP-II); UGC-SAP-DRS (Phase-III)].
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Kankariya, R.A., Chaudhari, A.B. & Dandi, N.D. Inhibitory efficacy of 2, 4-diacetylphloroglucinol against SARS-COV-2 proteins: in silico study. Biologia 77, 815–828 (2022). https://doi.org/10.1007/s11756-021-00979-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-021-00979-4